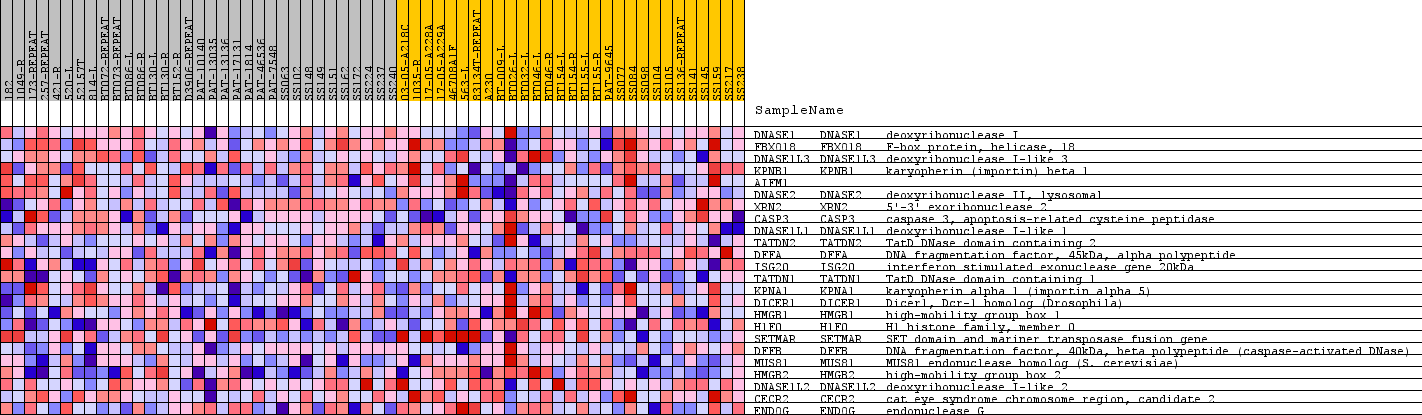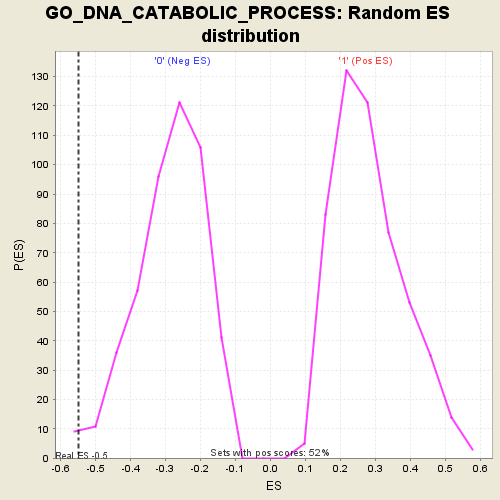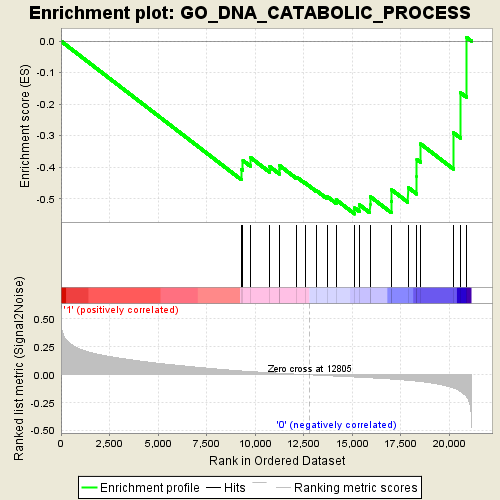
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | NOV0093_collapsed_to_symbols.NOV0093.cls #obese_versus_normal.NOV0093.cls #obese_versus_normal_repos |
| Phenotype | NOV0093.cls#obese_versus_normal_repos |
| Upregulated in class | 0 |
| GeneSet | GO_DNA_CATABOLIC_PROCESS |
| Enrichment Score (ES) | -0.54836845 |
| Normalized Enrichment Score (NES) | -1.9117553 |
| Nominal p-value | 0.008385744 |
| FDR q-value | 0.37730134 |
| FWER p-Value | 0.749 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | DNASE1 | DNASE1 Entrez, Source | deoxyribonuclease I | 9288 | 0.034 | -0.4067 | No |
| 2 | FBXO18 | FBXO18 Entrez, Source | F-box protein, helicase, 18 | 9371 | 0.033 | -0.3786 | No |
| 3 | DNASE1L3 | DNASE1L3 Entrez, Source | deoxyribonuclease I-like 3 | 9764 | 0.029 | -0.3694 | No |
| 4 | KPNB1 | KPNB1 Entrez, Source | karyopherin (importin) beta 1 | 10753 | 0.019 | -0.3981 | No |
| 5 | AIFM1 | 11253 | 0.014 | -0.4081 | No | ||
| 6 | DNASE2 | DNASE2 Entrez, Source | deoxyribonuclease II, lysosomal | 11255 | 0.014 | -0.3945 | No |
| 7 | XRN2 | XRN2 Entrez, Source | 5'-3' exoribonuclease 2 | 12140 | 0.006 | -0.4310 | No |
| 8 | CASP3 | CASP3 Entrez, Source | caspase 3, apoptosis-related cysteine peptidase | 12572 | 0.002 | -0.4495 | No |
| 9 | DNASE1L1 | DNASE1L1 Entrez, Source | deoxyribonuclease I-like 1 | 13155 | -0.003 | -0.4742 | No |
| 10 | TATDN2 | TATDN2 Entrez, Source | TatD DNase domain containing 2 | 13705 | -0.008 | -0.4926 | No |
| 11 | DFFA | DFFA Entrez, Source | DNA fragmentation factor, 45kDa, alpha polypeptide | 14175 | -0.012 | -0.5032 | No |
| 12 | ISG20 | ISG20 Entrez, Source | interferon stimulated exonuclease gene 20kDa | 15130 | -0.020 | -0.5291 | Yes |
| 13 | TATDN1 | TATDN1 Entrez, Source | TatD DNase domain containing 1 | 15366 | -0.022 | -0.5188 | Yes |
| 14 | KPNA1 | KPNA1 Entrez, Source | karyopherin alpha 1 (importin alpha 5) | 15916 | -0.027 | -0.5187 | Yes |
| 15 | DICER1 | DICER1 Entrez, Source | Dicer1, Dcr-1 homolog (Drosophila) | 15920 | -0.027 | -0.4927 | Yes |
| 16 | HMGB1 | HMGB1 Entrez, Source | high-mobility group box 1 | 17008 | -0.038 | -0.5074 | Yes |
| 17 | H1F0 | H1F0 Entrez, Source | H1 histone family, member 0 | 17030 | -0.038 | -0.4714 | Yes |
| 18 | SETMAR | SETMAR Entrez, Source | SET domain and mariner transposase fusion gene | 17874 | -0.049 | -0.4636 | Yes |
| 19 | DFFB | DFFB Entrez, Source | DNA fragmentation factor, 40kDa, beta polypeptide (caspase-activated DNase) | 18329 | -0.056 | -0.4303 | Yes |
| 20 | MUS81 | MUS81 Entrez, Source | MUS81 endonuclease homolog (S. cerevisiae) | 18334 | -0.056 | -0.3756 | Yes |
| 21 | HMGB2 | HMGB2 Entrez, Source | high-mobility group box 2 | 18526 | -0.060 | -0.3260 | Yes |
| 22 | DNASE1L2 | DNASE1L2 Entrez, Source | deoxyribonuclease I-like 2 | 20215 | -0.119 | -0.2905 | Yes |
| 23 | CECR2 | CECR2 Entrez, Source | cat eye syndrome chromosome region, candidate 2 | 20576 | -0.148 | -0.1632 | Yes |
| 24 | ENDOG | ENDOG Entrez, Source | endonuclease G | 20896 | -0.195 | 0.0117 | Yes |