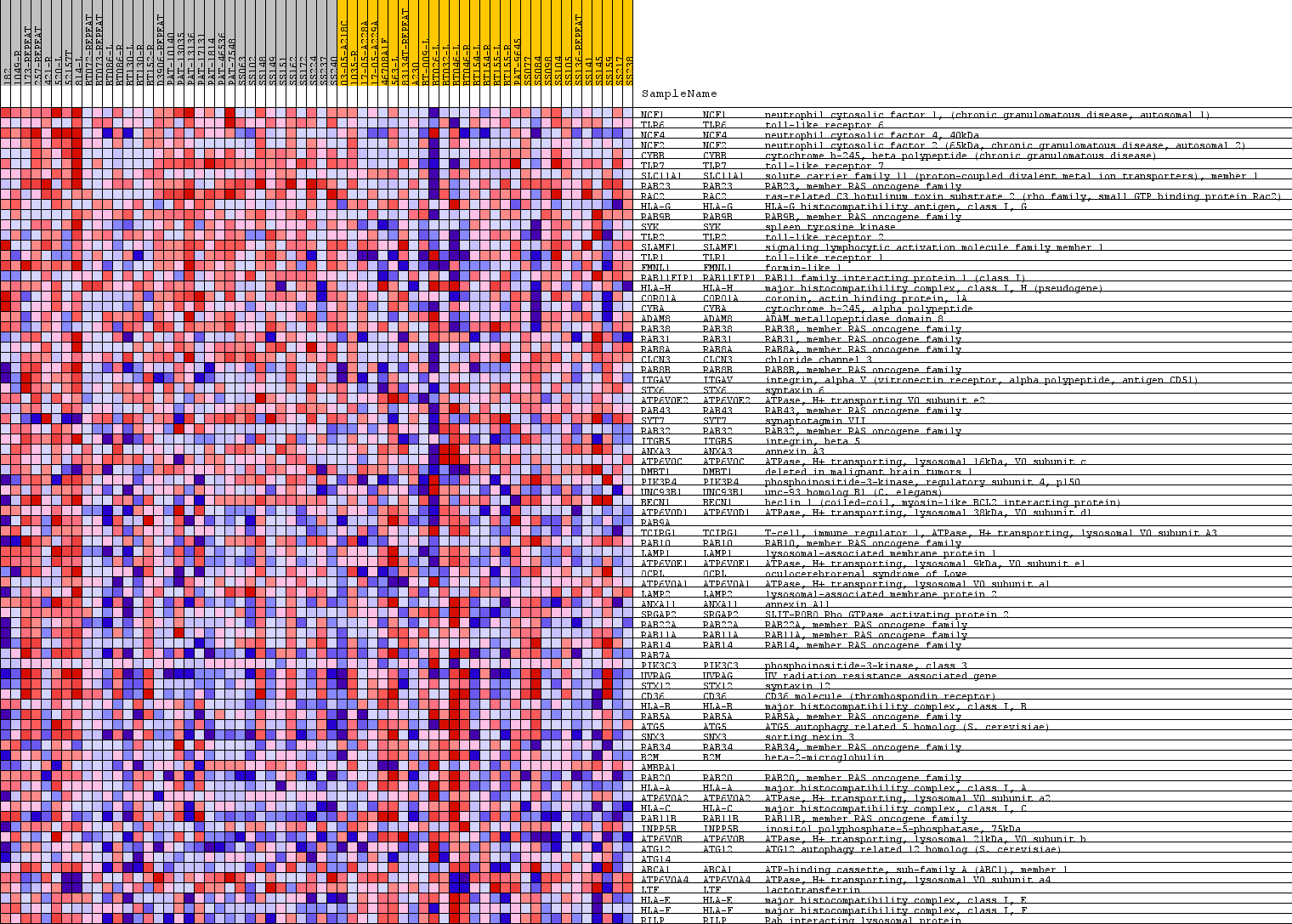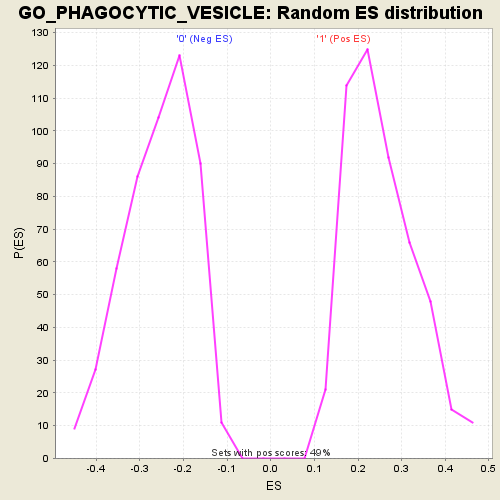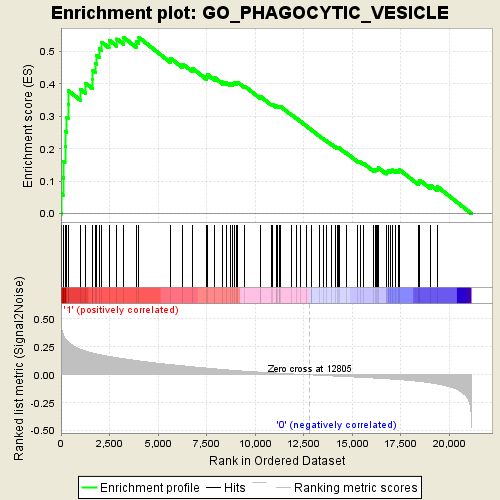
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | NOV0093_collapsed_to_symbols.NOV0093.cls #obese_versus_normal.NOV0093.cls #obese_versus_normal_repos |
| Phenotype | NOV0093.cls#obese_versus_normal_repos |
| Upregulated in class | 1 |
| GeneSet | GO_PHAGOCYTIC_VESICLE |
| Enrichment Score (ES) | 0.5427138 |
| Normalized Enrichment Score (NES) | 2.1402547 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.25282398 |
| FWER p-Value | 0.238 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | NCF1 | NCF1 Entrez, Source | neutrophil cytosolic factor 1, (chronic granulomatous disease, autosomal 1) | 21 | 0.420 | 0.0611 | Yes |
| 2 | TLR6 | TLR6 Entrez, Source | toll-like receptor 6 | 104 | 0.365 | 0.1112 | Yes |
| 3 | NCF4 | NCF4 Entrez, Source | neutrophil cytosolic factor 4, 40kDa | 139 | 0.348 | 0.1611 | Yes |
| 4 | NCF2 | NCF2 Entrez, Source | neutrophil cytosolic factor 2 (65kDa, chronic granulomatous disease, autosomal 2) | 203 | 0.327 | 0.2065 | Yes |
| 5 | CYBB | CYBB Entrez, Source | cytochrome b-245, beta polypeptide (chronic granulomatous disease) | 219 | 0.323 | 0.2536 | Yes |
| 6 | TLR7 | TLR7 Entrez, Source | toll-like receptor 7 | 292 | 0.306 | 0.2956 | Yes |
| 7 | SLC11A1 | SLC11A1 Entrez, Source | solute carrier family 11 (proton-coupled divalent metal ion transporters), member 1 | 360 | 0.294 | 0.3360 | Yes |
| 8 | RAB23 | RAB23 Entrez, Source | RAB23, member RAS oncogene family | 372 | 0.293 | 0.3788 | Yes |
| 9 | RAC2 | RAC2 Entrez, Source | ras-related C3 botulinum toxin substrate 2 (rho family, small GTP binding protein Rac2) | 1005 | 0.228 | 0.3826 | Yes |
| 10 | HLA-G | HLA-G Entrez, Source | HLA-G histocompatibility antigen, class I, G | 1253 | 0.212 | 0.4023 | Yes |
| 11 | RAB9B | RAB9B Entrez, Source | RAB9B, member RAS oncogene family | 1634 | 0.194 | 0.4129 | Yes |
| 12 | SYK | SYK Entrez, Source | spleen tyrosine kinase | 1640 | 0.194 | 0.4414 | Yes |
| 13 | TLR2 | TLR2 Entrez, Source | toll-like receptor 2 | 1778 | 0.188 | 0.4627 | Yes |
| 14 | SLAMF1 | SLAMF1 Entrez, Source | signaling lymphocytic activation molecule family member 1 | 1838 | 0.186 | 0.4874 | Yes |
| 15 | TLR1 | TLR1 Entrez, Source | toll-like receptor 1 | 1968 | 0.181 | 0.5081 | Yes |
| 16 | FMNL1 | FMNL1 Entrez, Source | formin-like 1 | 2087 | 0.176 | 0.5285 | Yes |
| 17 | RAB11FIP1 | RAB11FIP1 Entrez, Source | RAB11 family interacting protein 1 (class I) | 2472 | 0.164 | 0.5346 | Yes |
| 18 | HLA-H | HLA-H Entrez, Source | major histocompatibility complex, class I, H (pseudogene) | 2877 | 0.153 | 0.5380 | Yes |
| 19 | CORO1A | CORO1A Entrez, Source | coronin, actin binding protein, 1A | 3235 | 0.143 | 0.5422 | Yes |
| 20 | CYBA | CYBA Entrez, Source | cytochrome b-245, alpha polypeptide | 3890 | 0.127 | 0.5300 | Yes |
| 21 | ADAM8 | ADAM8 Entrez, Source | ADAM metallopeptidase domain 8 | 4010 | 0.124 | 0.5427 | Yes |
| 22 | RAB38 | RAB38 Entrez, Source | RAB38, member RAS oncogene family | 5644 | 0.091 | 0.4787 | No |
| 23 | RAB31 | RAB31 Entrez, Source | RAB31, member RAS oncogene family | 6268 | 0.080 | 0.4609 | No |
| 24 | RAB8A | RAB8A Entrez, Source | RAB8A, member RAS oncogene family | 6768 | 0.071 | 0.4478 | No |
| 25 | CLCN3 | CLCN3 Entrez, Source | chloride channel 3 | 7491 | 0.060 | 0.4223 | No |
| 26 | RAB8B | RAB8B Entrez, Source | RAB8B, member RAS oncogene family | 7530 | 0.059 | 0.4293 | No |
| 27 | ITGAV | ITGAV Entrez, Source | integrin, alpha V (vitronectin receptor, alpha polypeptide, antigen CD51) | 7925 | 0.053 | 0.4184 | No |
| 28 | STX6 | STX6 Entrez, Source | syntaxin 6 | 8338 | 0.047 | 0.4057 | No |
| 29 | ATP6V0E2 | ATP6V0E2 Entrez, Source | ATPase, H+ transporting V0 subunit e2 | 8497 | 0.045 | 0.4048 | No |
| 30 | RAB43 | RAB43 Entrez, Source | RAB43, member RAS oncogene family | 8734 | 0.041 | 0.3997 | No |
| 31 | SYT7 | SYT7 Entrez, Source | synaptotagmin VII | 8826 | 0.040 | 0.4013 | No |
| 32 | RAB32 | RAB32 Entrez, Source | RAB32, member RAS oncogene family | 8911 | 0.039 | 0.4030 | No |
| 33 | ITGB5 | ITGB5 Entrez, Source | integrin, beta 5 | 9045 | 0.037 | 0.4022 | No |
| 34 | ANXA3 | ANXA3 Entrez, Source | annexin A3 | 9095 | 0.036 | 0.4052 | No |
| 35 | ATP6V0C | ATP6V0C Entrez, Source | ATPase, H+ transporting, lysosomal 16kDa, V0 subunit c | 9466 | 0.032 | 0.3924 | No |
| 36 | DMBT1 | DMBT1 Entrez, Source | deleted in malignant brain tumors 1 | 10284 | 0.023 | 0.3570 | No |
| 37 | PIK3R4 | PIK3R4 Entrez, Source | phosphoinositide-3-kinase, regulatory subunit 4, p150 | 10298 | 0.023 | 0.3598 | No |
| 38 | UNC93B1 | UNC93B1 Entrez, Source | unc-93 homolog B1 (C. elegans) | 10829 | 0.018 | 0.3372 | No |
| 39 | BECN1 | BECN1 Entrez, Source | beclin 1 (coiled-coil, myosin-like BCL2 interacting protein) | 10915 | 0.017 | 0.3357 | No |
| 40 | ATP6V0D1 | ATP6V0D1 Entrez, Source | ATPase, H+ transporting, lysosomal 38kDa, V0 subunit d1 | 11081 | 0.016 | 0.3302 | No |
| 41 | RAB9A | 11094 | 0.016 | 0.3319 | No | ||
| 42 | TCIRG1 | TCIRG1 Entrez, Source | T-cell, immune regulator 1, ATPase, H+ transporting, lysosomal V0 subunit A3 | 11175 | 0.015 | 0.3303 | No |
| 43 | RAB10 | RAB10 Entrez, Source | RAB10, member RAS oncogene family | 11254 | 0.014 | 0.3287 | No |
| 44 | LAMP1 | LAMP1 Entrez, Source | lysosomal-associated membrane protein 1 | 11257 | 0.014 | 0.3307 | No |
| 45 | ATP6V0E1 | ATP6V0E1 Entrez, Source | ATPase, H+ transporting, lysosomal 9kDa, V0 subunit e1 | 11305 | 0.013 | 0.3304 | No |
| 46 | OCRL | OCRL Entrez, Source | oculocerebrorenal syndrome of Lowe | 11873 | 0.008 | 0.3047 | No |
| 47 | ATP6V0A1 | ATP6V0A1 Entrez, Source | ATPase, H+ transporting, lysosomal V0 subunit a1 | 12150 | 0.005 | 0.2924 | No |
| 48 | LAMP2 | LAMP2 Entrez, Source | lysosomal-associated membrane protein 2 | 12355 | 0.004 | 0.2833 | No |
| 49 | ANXA11 | ANXA11 Entrez, Source | annexin A11 | 12633 | 0.001 | 0.2703 | No |
| 50 | SRGAP2 | SRGAP2 Entrez, Source | SLIT-ROBO Rho GTPase activating protein 2 | 12880 | -0.001 | 0.2588 | No |
| 51 | RAB22A | RAB22A Entrez, Source | RAB22A, member RAS oncogene family | 12919 | -0.001 | 0.2571 | No |
| 52 | RAB11A | RAB11A Entrez, Source | RAB11A, member RAS oncogene family | 13334 | -0.004 | 0.2381 | No |
| 53 | RAB14 | RAB14 Entrez, Source | RAB14, member RAS oncogene family | 13516 | -0.006 | 0.2304 | No |
| 54 | RAB7A | 13670 | -0.007 | 0.2243 | No | ||
| 55 | PIK3C3 | PIK3C3 Entrez, Source | phosphoinositide-3-kinase, class 3 | 13944 | -0.010 | 0.2128 | No |
| 56 | UVRAG | UVRAG Entrez, Source | UV radiation resistance associated gene | 14113 | -0.011 | 0.2065 | No |
| 57 | STX12 | STX12 Entrez, Source | syntaxin 12 | 14258 | -0.013 | 0.2015 | No |
| 58 | CD36 | CD36 Entrez, Source | CD36 molecule (thrombospondin receptor) | 14295 | -0.013 | 0.2017 | No |
| 59 | HLA-B | HLA-B Entrez, Source | major histocompatibility complex, class I, B | 14319 | -0.013 | 0.2026 | No |
| 60 | RAB5A | RAB5A Entrez, Source | RAB5A, member RAS oncogene family | 14693 | -0.016 | 0.1872 | No |
| 61 | ATG5 | ATG5 Entrez, Source | ATG5 autophagy related 5 homolog (S. cerevisiae) | 15288 | -0.021 | 0.1622 | No |
| 62 | SNX3 | SNX3 Entrez, Source | sorting nexin 3 | 15418 | -0.022 | 0.1594 | No |
| 63 | RAB34 | RAB34 Entrez, Source | RAB34, member RAS oncogene family | 15578 | -0.024 | 0.1554 | No |
| 64 | B2M | B2M Entrez, Source | beta-2-microglobulin | 16102 | -0.029 | 0.1348 | No |
| 65 | AMBRA1 | 16177 | -0.029 | 0.1356 | No | ||
| 66 | RAB20 | RAB20 Entrez, Source | RAB20, member RAS oncogene family | 16261 | -0.030 | 0.1361 | No |
| 67 | HLA-A | HLA-A Entrez, Source | major histocompatibility complex, class I, A | 16326 | -0.031 | 0.1377 | No |
| 68 | ATP6V0A2 | ATP6V0A2 Entrez, Source | ATPase, H+ transporting, lysosomal V0 subunit a2 | 16346 | -0.031 | 0.1414 | No |
| 69 | HLA-C | HLA-C Entrez, Source | major histocompatibility complex, class I, C | 16751 | -0.035 | 0.1274 | No |
| 70 | RAB11B | RAB11B Entrez, Source | RAB11B, member RAS oncogene family | 16787 | -0.035 | 0.1310 | No |
| 71 | INPP5B | INPP5B Entrez, Source | inositol polyphosphate-5-phosphatase, 75kDa | 16859 | -0.036 | 0.1329 | No |
| 72 | ATP6V0B | ATP6V0B Entrez, Source | ATPase, H+ transporting, lysosomal 21kDa, V0 subunit b | 16966 | -0.037 | 0.1334 | No |
| 73 | ATG12 | ATG12 Entrez, Source | ATG12 autophagy related 12 homolog (S. cerevisiae) | 17057 | -0.038 | 0.1348 | No |
| 74 | ATG14 | 17231 | -0.040 | 0.1326 | No | ||
| 75 | ABCA1 | ABCA1 Entrez, Source | ATP-binding cassette, sub-family A (ABC1), member 1 | 17369 | -0.042 | 0.1323 | No |
| 76 | ATP6V0A4 | ATP6V0A4 Entrez, Source | ATPase, H+ transporting, lysosomal V0 subunit a4 | 17440 | -0.043 | 0.1353 | No |
| 77 | LTF | LTF Entrez, Source | lactotransferrin | 18416 | -0.058 | 0.0977 | No |
| 78 | HLA-E | HLA-E Entrez, Source | major histocompatibility complex, class I, E | 18485 | -0.059 | 0.1032 | No |
| 79 | HLA-F | HLA-F Entrez, Source | major histocompatibility complex, class I, F | 19036 | -0.072 | 0.0877 | No |
| 80 | RILP | RILP Entrez, Source | Rab interacting lysosomal protein | 19403 | -0.083 | 0.0826 | No |