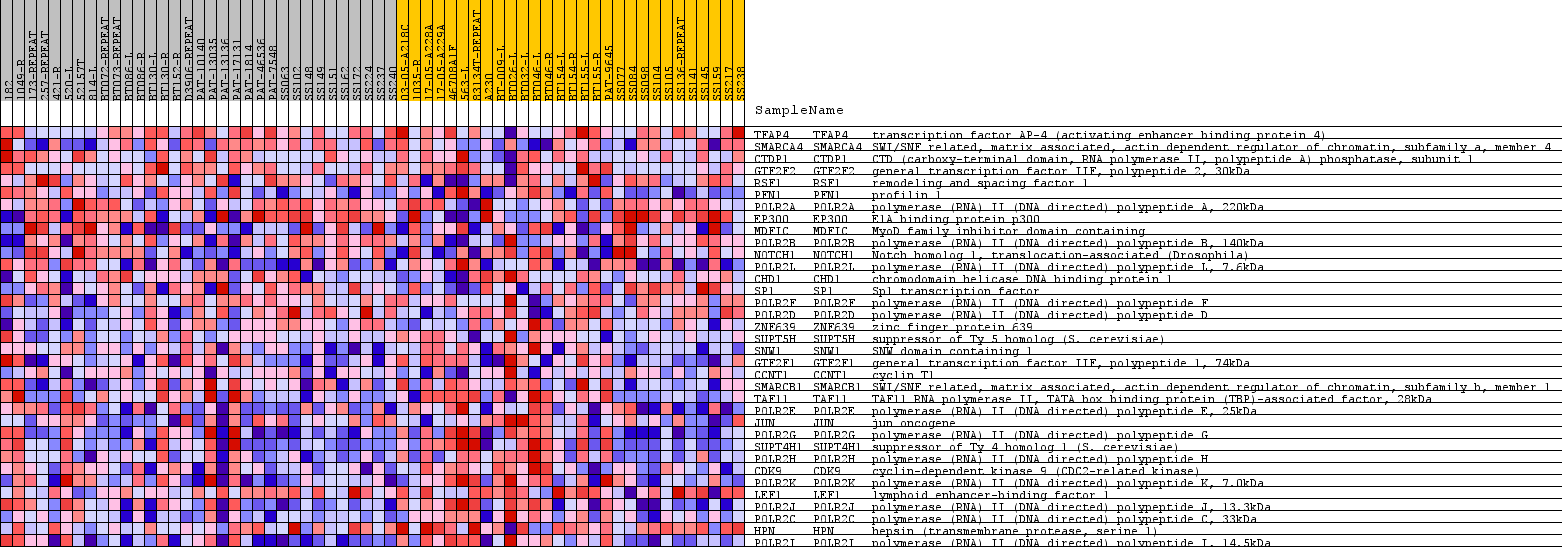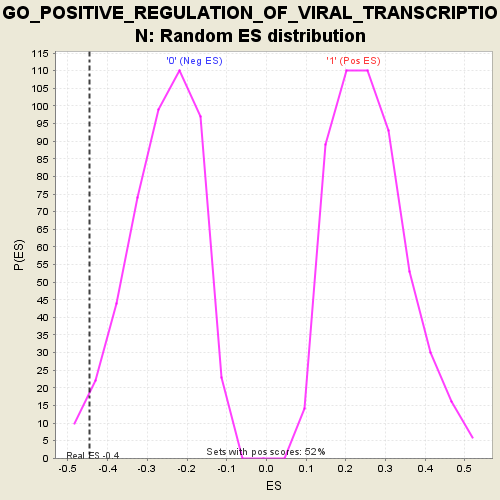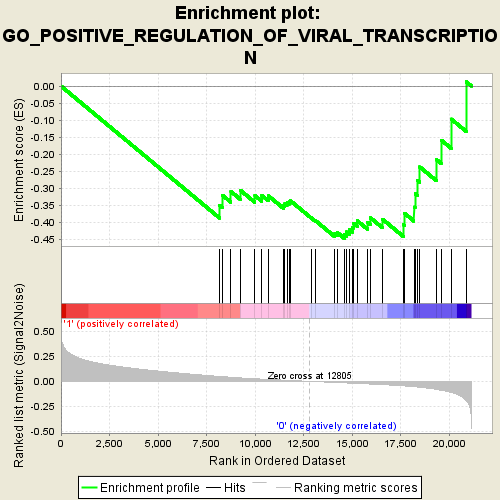
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | NOV0093_collapsed_to_symbols.NOV0093.cls #obese_versus_normal.NOV0093.cls #obese_versus_normal_repos |
| Phenotype | NOV0093.cls#obese_versus_normal_repos |
| Upregulated in class | 0 |
| GeneSet | GO_POSITIVE_REGULATION_OF_VIRAL_TRANSCRIPTION |
| Enrichment Score (ES) | -0.44696438 |
| Normalized Enrichment Score (NES) | -1.714965 |
| Nominal p-value | 0.025052192 |
| FDR q-value | 1.0 |
| FWER p-Value | 0.996 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | TFAP4 | TFAP4 Entrez, Source | transcription factor AP-4 (activating enhancer binding protein 4) | 8172 | 0.049 | -0.3493 | No |
| 2 | SMARCA4 | SMARCA4 Entrez, Source | SWI/SNF related, matrix associated, actin dependent regulator of chromatin, subfamily a, member 4 | 8315 | 0.047 | -0.3200 | No |
| 3 | CTDP1 | CTDP1 Entrez, Source | CTD (carboxy-terminal domain, RNA polymerase II, polypeptide A) phosphatase, subunit 1 | 8750 | 0.041 | -0.3090 | No |
| 4 | GTF2F2 | GTF2F2 Entrez, Source | general transcription factor IIF, polypeptide 2, 30kDa | 9231 | 0.035 | -0.3051 | No |
| 5 | RSF1 | RSF1 Entrez, Source | remodeling and spacing factor 1 | 9987 | 0.026 | -0.3209 | No |
| 6 | PFN1 | PFN1 Entrez, Source | profilin 1 | 10342 | 0.022 | -0.3204 | No |
| 7 | POLR2A | POLR2A Entrez, Source | polymerase (RNA) II (DNA directed) polypeptide A, 220kDa | 10684 | 0.019 | -0.3218 | No |
| 8 | EP300 | EP300 Entrez, Source | E1A binding protein p300 | 11472 | 0.012 | -0.3501 | No |
| 9 | MDFIC | MDFIC Entrez, Source | MyoD family inhibitor domain containing | 11533 | 0.011 | -0.3444 | No |
| 10 | POLR2B | POLR2B Entrez, Source | polymerase (RNA) II (DNA directed) polypeptide B, 140kDa | 11643 | 0.010 | -0.3416 | No |
| 11 | NOTCH1 | NOTCH1 Entrez, Source | Notch homolog 1, translocation-associated (Drosophila) | 11747 | 0.009 | -0.3393 | No |
| 12 | POLR2L | POLR2L Entrez, Source | polymerase (RNA) II (DNA directed) polypeptide L, 7.6kDa | 11823 | 0.009 | -0.3363 | No |
| 13 | CHD1 | CHD1 Entrez, Source | chromodomain helicase DNA binding protein 1 | 12888 | -0.001 | -0.3862 | No |
| 14 | SP1 | SP1 Entrez, Source | Sp1 transcription factor | 13122 | -0.003 | -0.3952 | No |
| 15 | POLR2F | POLR2F Entrez, Source | polymerase (RNA) II (DNA directed) polypeptide F | 14106 | -0.011 | -0.4330 | No |
| 16 | POLR2D | POLR2D Entrez, Source | polymerase (RNA) II (DNA directed) polypeptide D | 14239 | -0.012 | -0.4297 | No |
| 17 | ZNF639 | ZNF639 Entrez, Source | zinc finger protein 639 | 14605 | -0.015 | -0.4352 | Yes |
| 18 | SUPT5H | SUPT5H Entrez, Source | suppressor of Ty 5 homolog (S. cerevisiae) | 14708 | -0.016 | -0.4275 | Yes |
| 19 | SNW1 | SNW1 Entrez, Source | SNW domain containing 1 | 14850 | -0.018 | -0.4207 | Yes |
| 20 | GTF2F1 | GTF2F1 Entrez, Source | general transcription factor IIF, polypeptide 1, 74kDa | 15006 | -0.019 | -0.4137 | Yes |
| 21 | CCNT1 | CCNT1 Entrez, Source | cyclin T1 | 15062 | -0.019 | -0.4015 | Yes |
| 22 | SMARCB1 | SMARCB1 Entrez, Source | SWI/SNF related, matrix associated, actin dependent regulator of chromatin, subfamily b, member 1 | 15258 | -0.021 | -0.3946 | Yes |
| 23 | TAF11 | TAF11 Entrez, Source | TAF11 RNA polymerase II, TATA box binding protein (TBP)-associated factor, 28kDa | 15799 | -0.026 | -0.4003 | Yes |
| 24 | POLR2E | POLR2E Entrez, Source | polymerase (RNA) II (DNA directed) polypeptide E, 25kDa | 15942 | -0.027 | -0.3862 | Yes |
| 25 | JUN | JUN Entrez, Source | jun oncogene | 16551 | -0.033 | -0.3897 | Yes |
| 26 | POLR2G | POLR2G Entrez, Source | polymerase (RNA) II (DNA directed) polypeptide G | 17652 | -0.046 | -0.4066 | Yes |
| 27 | SUPT4H1 | SUPT4H1 Entrez, Source | suppressor of Ty 4 homolog 1 (S. cerevisiae) | 17700 | -0.047 | -0.3730 | Yes |
| 28 | POLR2H | POLR2H Entrez, Source | polymerase (RNA) II (DNA directed) polypeptide H | 18183 | -0.054 | -0.3544 | Yes |
| 29 | CDK9 | CDK9 Entrez, Source | cyclin-dependent kinase 9 (CDC2-related kinase) | 18241 | -0.055 | -0.3150 | Yes |
| 30 | POLR2K | POLR2K Entrez, Source | polymerase (RNA) II (DNA directed) polypeptide K, 7.0kDa | 18360 | -0.057 | -0.2768 | Yes |
| 31 | LEF1 | LEF1 Entrez, Source | lymphoid enhancer-binding factor 1 | 18483 | -0.059 | -0.2369 | Yes |
| 32 | POLR2J | POLR2J Entrez, Source | polymerase (RNA) II (DNA directed) polypeptide J, 13.3kDa | 19332 | -0.081 | -0.2150 | Yes |
| 33 | POLR2C | POLR2C Entrez, Source | polymerase (RNA) II (DNA directed) polypeptide C, 33kDa | 19583 | -0.090 | -0.1580 | Yes |
| 34 | HPN | HPN Entrez, Source | hepsin (transmembrane protease, serine 1) | 20099 | -0.112 | -0.0964 | Yes |
| 35 | POLR2I | POLR2I Entrez, Source | polymerase (RNA) II (DNA directed) polypeptide I, 14.5kDa | 20873 | -0.190 | 0.0128 | Yes |