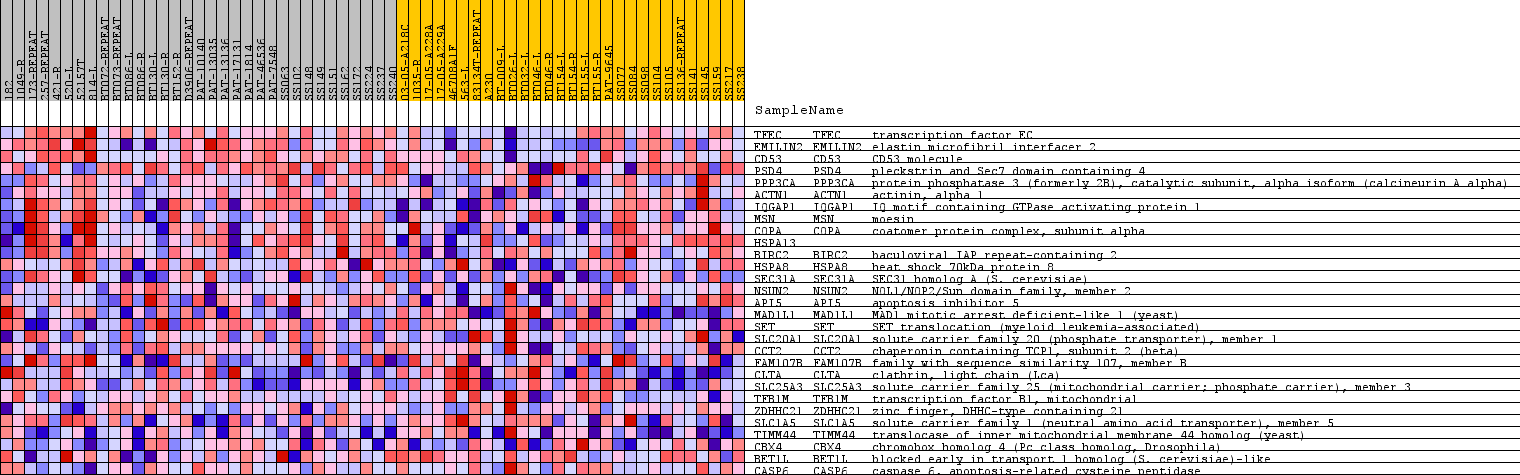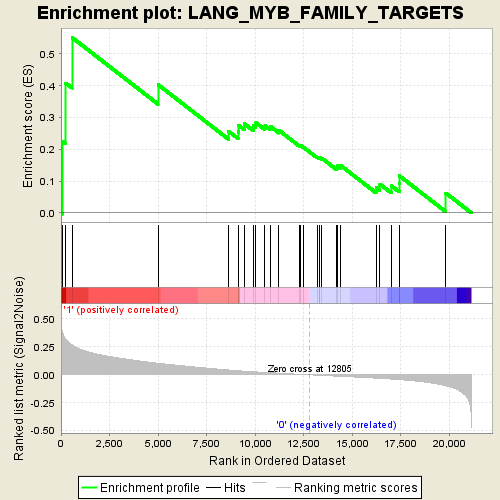
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | NOV0093_collapsed_to_symbols.NOV0093.cls #obese_versus_normal.NOV0093.cls #obese_versus_normal_repos |
| Phenotype | NOV0093.cls#obese_versus_normal_repos |
| Upregulated in class | 1 |
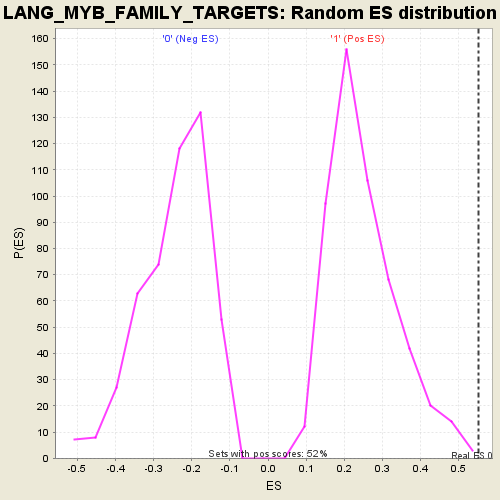
| GeneSet | LANG_MYB_FAMILY_TARGETS |
| Enrichment Score (ES) | 0.5518036 |
| Normalized Enrichment Score (NES) | 2.2201447 |
| Nominal p-value | 0.0019305019 |
| FDR q-value | 0.45204774 |
| FWER p-Value | 0.131 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | TFEC | TFEC Entrez, Source | transcription factor EC | 69 | 0.382 | 0.2249 | Yes |
| 2 | EMILIN2 | EMILIN2 Entrez, Source | elastin microfibril interfacer 2 | 232 | 0.320 | 0.4085 | Yes |
| 3 | CD53 | CD53 Entrez, Source | CD53 molecule | 565 | 0.266 | 0.5518 | Yes |
| 4 | PSD4 | PSD4 Entrez, Source | pleckstrin and Sec7 domain containing 4 | 4994 | 0.103 | 0.4037 | No |
| 5 | PPP3CA | PPP3CA Entrez, Source | protein phosphatase 3 (formerly 2B), catalytic subunit, alpha isoform (calcineurin A alpha) | 8632 | 0.043 | 0.2569 | No |
| 6 | ACTN1 | ACTN1 Entrez, Source | actinin, alpha 1 | 9122 | 0.036 | 0.2553 | No |
| 7 | IQGAP1 | IQGAP1 Entrez, Source | IQ motif containing GTPase activating protein 1 | 9153 | 0.036 | 0.2751 | No |
| 8 | MSN | MSN Entrez, Source | moesin | 9437 | 0.032 | 0.2810 | No |
| 9 | COPA | COPA Entrez, Source | coatomer protein complex, subunit alpha | 9913 | 0.027 | 0.2746 | No |
| 10 | HSPA13 | 10038 | 0.026 | 0.2840 | No | ||
| 11 | BIRC2 | BIRC2 Entrez, Source | baculoviral IAP repeat-containing 2 | 10505 | 0.021 | 0.2743 | No |
| 12 | HSPA8 | HSPA8 Entrez, Source | heat shock 70kDa protein 8 | 10769 | 0.018 | 0.2729 | No |
| 13 | SEC31A | SEC31A Entrez, Source | SEC31 homolog A (S. cerevisiae) | 11224 | 0.014 | 0.2599 | No |
| 14 | NSUN2 | NSUN2 Entrez, Source | NOL1/NOP2/Sun domain family, member 2 | 12271 | 0.004 | 0.2130 | No |
| 15 | API5 | API5 Entrez, Source | apoptosis inhibitor 5 | 12334 | 0.004 | 0.2124 | No |
| 16 | MAD1L1 | MAD1L1 Entrez, Source | MAD1 mitotic arrest deficient-like 1 (yeast) | 12494 | 0.003 | 0.2064 | No |
| 17 | SET | SET Entrez, Source | SET translocation (myeloid leukemia-associated) | 13194 | -0.003 | 0.1753 | No |
| 18 | SLC20A1 | SLC20A1 Entrez, Source | solute carrier family 20 (phosphate transporter), member 1 | 13292 | -0.004 | 0.1731 | No |
| 19 | CCT2 | CCT2 Entrez, Source | chaperonin containing TCP1, subunit 2 (beta) | 13315 | -0.004 | 0.1746 | No |
| 20 | FAM107B | FAM107B Entrez, Source | family with sequence similarity 107, member B | 13436 | -0.005 | 0.1722 | No |
| 21 | CLTA | CLTA Entrez, Source | clathrin, light chain (Lca) | 14198 | -0.012 | 0.1434 | No |
| 22 | SLC25A3 | SLC25A3 Entrez, Source | solute carrier family 25 (mitochondrial carrier; phosphate carrier), member 3 | 14260 | -0.013 | 0.1480 | No |
| 23 | TFB1M | TFB1M Entrez, Source | transcription factor B1, mitochondrial | 14403 | -0.014 | 0.1495 | No |
| 24 | ZDHHC21 | ZDHHC21 Entrez, Source | zinc finger, DHHC-type containing 21 | 16230 | -0.030 | 0.0809 | No |
| 25 | SLC1A5 | SLC1A5 Entrez, Source | solute carrier family 1 (neutral amino acid transporter), member 5 | 16425 | -0.032 | 0.0908 | No |
| 26 | TIMM44 | TIMM44 Entrez, Source | translocase of inner mitochondrial membrane 44 homolog (yeast) | 17010 | -0.038 | 0.0856 | No |
| 27 | CBX4 | CBX4 Entrez, Source | chromobox homolog 4 (Pc class homolog, Drosophila) | 17425 | -0.043 | 0.0915 | No |
| 28 | BET1L | BET1L Entrez, Source | blocked early in transport 1 homolog (S. cerevisiae)-like | 17433 | -0.043 | 0.1168 | No |
| 29 | CASP6 | CASP6 Entrez, Source | caspase 6, apoptosis-related cysteine peptidase | 19827 | -0.099 | 0.0624 | No |