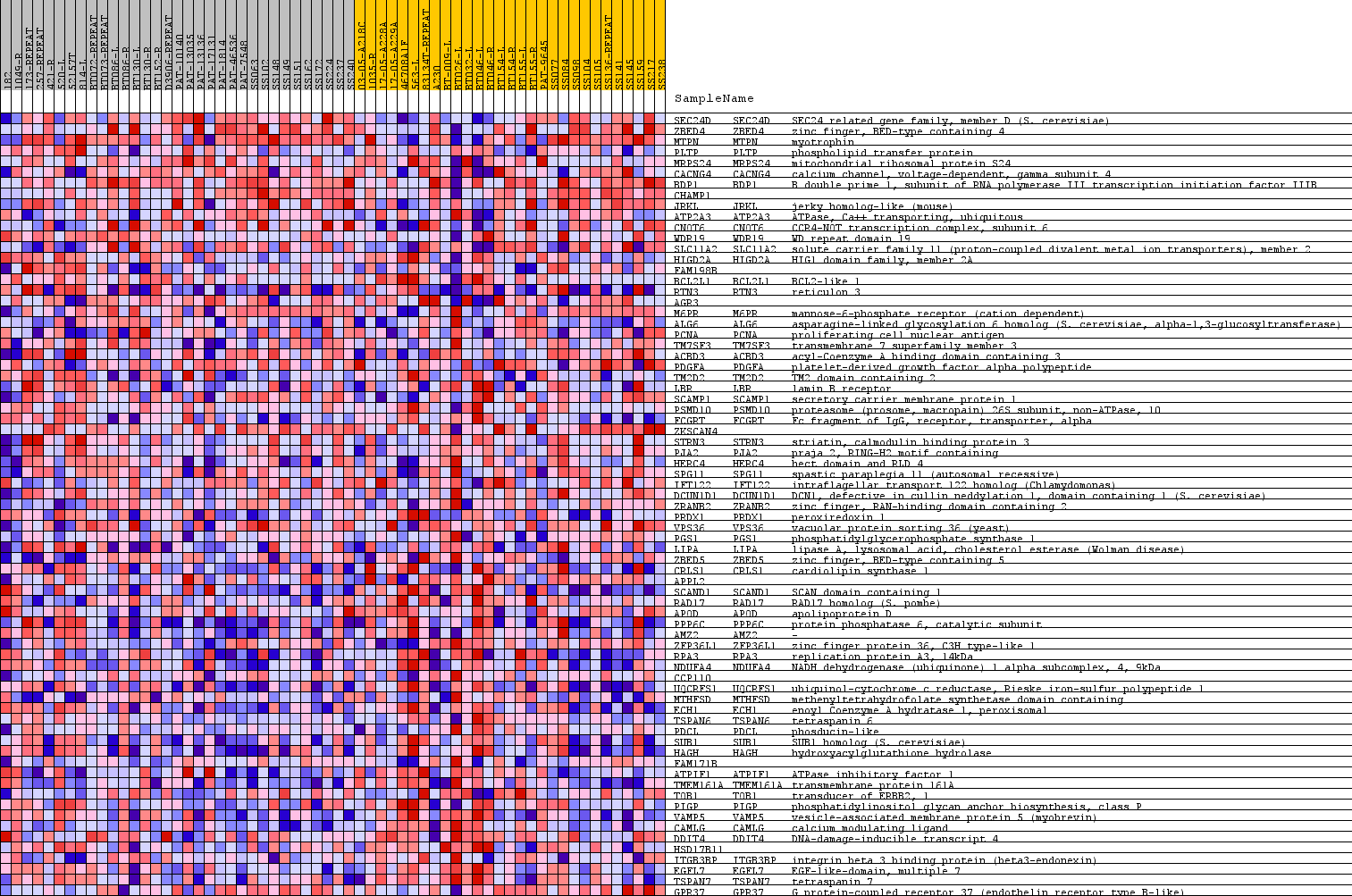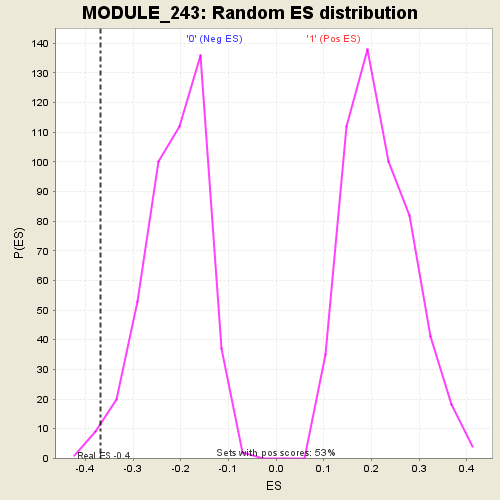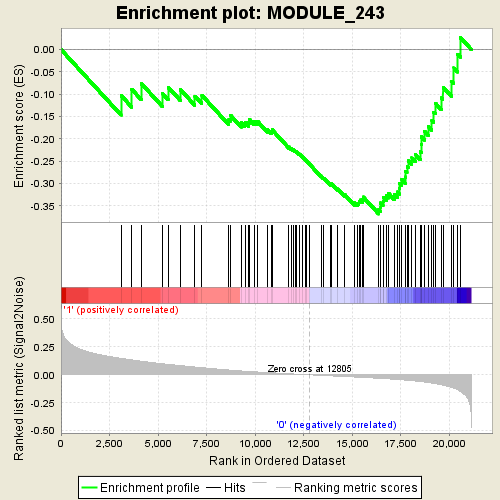
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | NOV0093_collapsed_to_symbols.NOV0093.cls #obese_versus_normal.NOV0093.cls #obese_versus_normal_repos |
| Phenotype | NOV0093.cls#obese_versus_normal_repos |
| Upregulated in class | 0 |
| GeneSet | MODULE_243 |
| Enrichment Score (ES) | -0.36852095 |
| Normalized Enrichment Score (NES) | -1.7412851 |
| Nominal p-value | 0.0148936175 |
| FDR q-value | 1.0 |
| FWER p-Value | 0.988 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | SEC24D | SEC24D Entrez, Source | SEC24 related gene family, member D (S. cerevisiae) | 3091 | 0.147 | -0.1029 | No |
| 2 | ZBED4 | ZBED4 Entrez, Source | zinc finger, BED-type containing 4 | 3644 | 0.133 | -0.0891 | No |
| 3 | MTPN | MTPN Entrez, Source | myotrophin | 4132 | 0.121 | -0.0760 | No |
| 4 | PLTP | PLTP Entrez, Source | phospholipid transfer protein | 5224 | 0.099 | -0.0983 | No |
| 5 | MRPS24 | MRPS24 Entrez, Source | mitochondrial ribosomal protein S24 | 5516 | 0.093 | -0.0842 | No |
| 6 | CACNG4 | CACNG4 Entrez, Source | calcium channel, voltage-dependent, gamma subunit 4 | 6143 | 0.082 | -0.0894 | No |
| 7 | BDP1 | BDP1 Entrez, Source | B double prime 1, subunit of RNA polymerase III transcription initiation factor IIIB | 6895 | 0.069 | -0.1044 | No |
| 8 | CHAMP1 | 7256 | 0.063 | -0.1025 | No | ||
| 9 | JRKL | JRKL Entrez, Source | jerky homolog-like (mouse) | 8647 | 0.042 | -0.1558 | No |
| 10 | ATP2A3 | ATP2A3 Entrez, Source | ATPase, Ca++ transporting, ubiquitous | 8742 | 0.041 | -0.1480 | No |
| 11 | CNOT6 | CNOT6 Entrez, Source | CCR4-NOT transcription complex, subunit 6 | 9283 | 0.034 | -0.1635 | No |
| 12 | WDR19 | WDR19 Entrez, Source | WD repeat domain 19 | 9476 | 0.032 | -0.1631 | No |
| 13 | SLC11A2 | SLC11A2 Entrez, Source | solute carrier family 11 (proton-coupled divalent metal ion transporters), member 2 | 9668 | 0.029 | -0.1634 | No |
| 14 | HIGD2A | HIGD2A Entrez, Source | HIG1 domain family, member 2A | 9705 | 0.029 | -0.1563 | No |
| 15 | FAM198B | 9971 | 0.026 | -0.1611 | No | ||
| 16 | BCL2L1 | BCL2L1 Entrez, Source | BCL2-like 1 | 10130 | 0.025 | -0.1611 | No |
| 17 | RTN3 | RTN3 Entrez, Source | reticulon 3 | 10635 | 0.020 | -0.1792 | No |
| 18 | AGR3 | 10839 | 0.018 | -0.1835 | No | ||
| 19 | M6PR | M6PR Entrez, Source | mannose-6-phosphate receptor (cation dependent) | 10867 | 0.018 | -0.1795 | No |
| 20 | ALG6 | ALG6 Entrez, Source | asparagine-linked glycosylation 6 homolog (S. cerevisiae, alpha-1,3-glucosyltransferase) | 11741 | 0.009 | -0.2182 | No |
| 21 | PCNA | PCNA Entrez, Source | proliferating cell nuclear antigen | 11876 | 0.008 | -0.2221 | No |
| 22 | TM7SF3 | TM7SF3 Entrez, Source | transmembrane 7 superfamily member 3 | 11964 | 0.007 | -0.2241 | No |
| 23 | ACBD3 | ACBD3 Entrez, Source | acyl-Coenzyme A binding domain containing 3 | 12074 | 0.006 | -0.2275 | No |
| 24 | PDGFA | PDGFA Entrez, Source | platelet-derived growth factor alpha polypeptide | 12138 | 0.006 | -0.2288 | No |
| 25 | TM2D2 | TM2D2 Entrez, Source | TM2 domain containing 2 | 12275 | 0.004 | -0.2340 | No |
| 26 | LBR | LBR Entrez, Source | lamin B receptor | 12419 | 0.003 | -0.2398 | No |
| 27 | SCAMP1 | SCAMP1 Entrez, Source | secretory carrier membrane protein 1 | 12616 | 0.002 | -0.2486 | No |
| 28 | PSMD10 | PSMD10 Entrez, Source | proteasome (prosome, macropain) 26S subunit, non-ATPase, 10 | 12635 | 0.001 | -0.2491 | No |
| 29 | FCGRT | FCGRT Entrez, Source | Fc fragment of IgG, receptor, transporter, alpha | 12778 | 0.000 | -0.2557 | No |
| 30 | ZKSCAN4 | 13391 | -0.005 | -0.2833 | No | ||
| 31 | STRN3 | STRN3 Entrez, Source | striatin, calmodulin binding protein 3 | 13534 | -0.006 | -0.2881 | No |
| 32 | PJA2 | PJA2 Entrez, Source | praja 2, RING-H2 motif containing | 13868 | -0.009 | -0.3011 | No |
| 33 | HERC4 | HERC4 Entrez, Source | hect domain and RLD 4 | 13922 | -0.010 | -0.3007 | No |
| 34 | SPG11 | SPG11 Entrez, Source | spastic paraplegia 11 (autosomal recessive) | 14244 | -0.013 | -0.3122 | No |
| 35 | IFT122 | IFT122 Entrez, Source | intraflagellar transport 122 homolog (Chlamydomonas) | 14602 | -0.015 | -0.3245 | No |
| 36 | DCUN1D1 | DCUN1D1 Entrez, Source | DCN1, defective in cullin neddylation 1, domain containing 1 (S. cerevisiae) | 15123 | -0.020 | -0.3433 | No |
| 37 | ZRANB2 | ZRANB2 Entrez, Source | zinc finger, RAN-binding domain containing 2 | 15273 | -0.021 | -0.3440 | No |
| 38 | PRDX1 | PRDX1 Entrez, Source | peroxiredoxin 1 | 15349 | -0.022 | -0.3410 | No |
| 39 | VPS36 | VPS36 Entrez, Source | vacuolar protein sorting 36 (yeast) | 15406 | -0.022 | -0.3370 | No |
| 40 | PGS1 | PGS1 Entrez, Source | phosphatidylglycerophosphate synthase 1 | 15533 | -0.023 | -0.3360 | No |
| 41 | LIPA | LIPA Entrez, Source | lipase A, lysosomal acid, cholesterol esterase (Wolman disease) | 15564 | -0.024 | -0.3303 | No |
| 42 | ZBED5 | ZBED5 Entrez, Source | zinc finger, BED-type containing 5 | 16370 | -0.031 | -0.3592 | Yes |
| 43 | CRLS1 | CRLS1 Entrez, Source | cardiolipin synthase 1 | 16442 | -0.032 | -0.3530 | Yes |
| 44 | APPL2 | 16449 | -0.032 | -0.3437 | Yes | ||
| 45 | SCAND1 | SCAND1 Entrez, Source | SCAN domain containing 1 | 16589 | -0.033 | -0.3403 | Yes |
| 46 | RAD17 | RAD17 Entrez, Source | RAD17 homolog (S. pombe) | 16604 | -0.033 | -0.3310 | Yes |
| 47 | APOD | APOD Entrez, Source | apolipoprotein D | 16752 | -0.035 | -0.3275 | Yes |
| 48 | PPP6C | PPP6C Entrez, Source | protein phosphatase 6, catalytic subunit | 16863 | -0.036 | -0.3218 | Yes |
| 49 | AMZ2 | AMZ2 Entrez, Source | - | 17176 | -0.040 | -0.3248 | Yes |
| 50 | ZFP36L1 | ZFP36L1 Entrez, Source | zinc finger protein 36, C3H type-like 1 | 17317 | -0.041 | -0.3191 | Yes |
| 51 | RPA3 | RPA3 Entrez, Source | replication protein A3, 14kDa | 17422 | -0.043 | -0.3112 | Yes |
| 52 | NDUFA4 | NDUFA4 Entrez, Source | NADH dehydrogenase (ubiquinone) 1 alpha subcomplex, 4, 9kDa | 17435 | -0.043 | -0.2990 | Yes |
| 53 | CCP110 | 17557 | -0.045 | -0.2914 | Yes | ||
| 54 | UQCRFS1 | UQCRFS1 Entrez, Source | ubiquinol-cytochrome c reductase, Rieske iron-sulfur polypeptide 1 | 17729 | -0.047 | -0.2855 | Yes |
| 55 | MTHFSD | MTHFSD Entrez, Source | methenyltetrahydrofolate synthetase domain containing | 17757 | -0.047 | -0.2726 | Yes |
| 56 | ECH1 | ECH1 Entrez, Source | enoyl Coenzyme A hydratase 1, peroxisomal | 17824 | -0.048 | -0.2612 | Yes |
| 57 | TSPAN6 | TSPAN6 Entrez, Source | tetraspanin 6 | 17884 | -0.049 | -0.2493 | Yes |
| 58 | PDCL | PDCL Entrez, Source | phosducin-like | 18076 | -0.052 | -0.2428 | Yes |
| 59 | SUB1 | SUB1 Entrez, Source | SUB1 homolog (S. cerevisiae) | 18261 | -0.055 | -0.2350 | Yes |
| 60 | HAGH | HAGH Entrez, Source | hydroxyacylglutathione hydrolase | 18510 | -0.060 | -0.2289 | Yes |
| 61 | FAM171B | 18556 | -0.061 | -0.2129 | Yes | ||
| 62 | ATPIF1 | ATPIF1 Entrez, Source | ATPase inhibitory factor 1 | 18570 | -0.061 | -0.1952 | Yes |
| 63 | TMEM161A | TMEM161A Entrez, Source | transmembrane protein 161A | 18733 | -0.065 | -0.1836 | Yes |
| 64 | TOB1 | TOB1 Entrez, Source | transducer of ERBB2, 1 | 18917 | -0.069 | -0.1718 | Yes |
| 65 | PIGP | PIGP Entrez, Source | phosphatidylinositol glycan anchor biosynthesis, class P | 19105 | -0.074 | -0.1586 | Yes |
| 66 | VAMP5 | VAMP5 Entrez, Source | vesicle-associated membrane protein 5 (myobrevin) | 19202 | -0.077 | -0.1402 | Yes |
| 67 | CAMLG | CAMLG Entrez, Source | calcium modulating ligand | 19289 | -0.080 | -0.1205 | Yes |
| 68 | DDIT4 | DDIT4 Entrez, Source | DNA-damage-inducible transcript 4 | 19597 | -0.090 | -0.1081 | Yes |
| 69 | HSD17B11 | 19677 | -0.093 | -0.0841 | Yes | ||
| 70 | ITGB3BP | ITGB3BP Entrez, Source | integrin beta 3 binding protein (beta3-endonexin) | 20122 | -0.113 | -0.0714 | Yes |
| 71 | EGFL7 | EGFL7 Entrez, Source | EGF-like-domain, multiple 7 | 20241 | -0.120 | -0.0411 | Yes |
| 72 | TSPAN7 | TSPAN7 Entrez, Source | tetraspanin 7 | 20436 | -0.133 | -0.0106 | Yes |
| 73 | GPR37 | GPR37 Entrez, Source | G protein-coupled receptor 37 (endothelin receptor type B-like) | 20570 | -0.148 | 0.0272 | Yes |