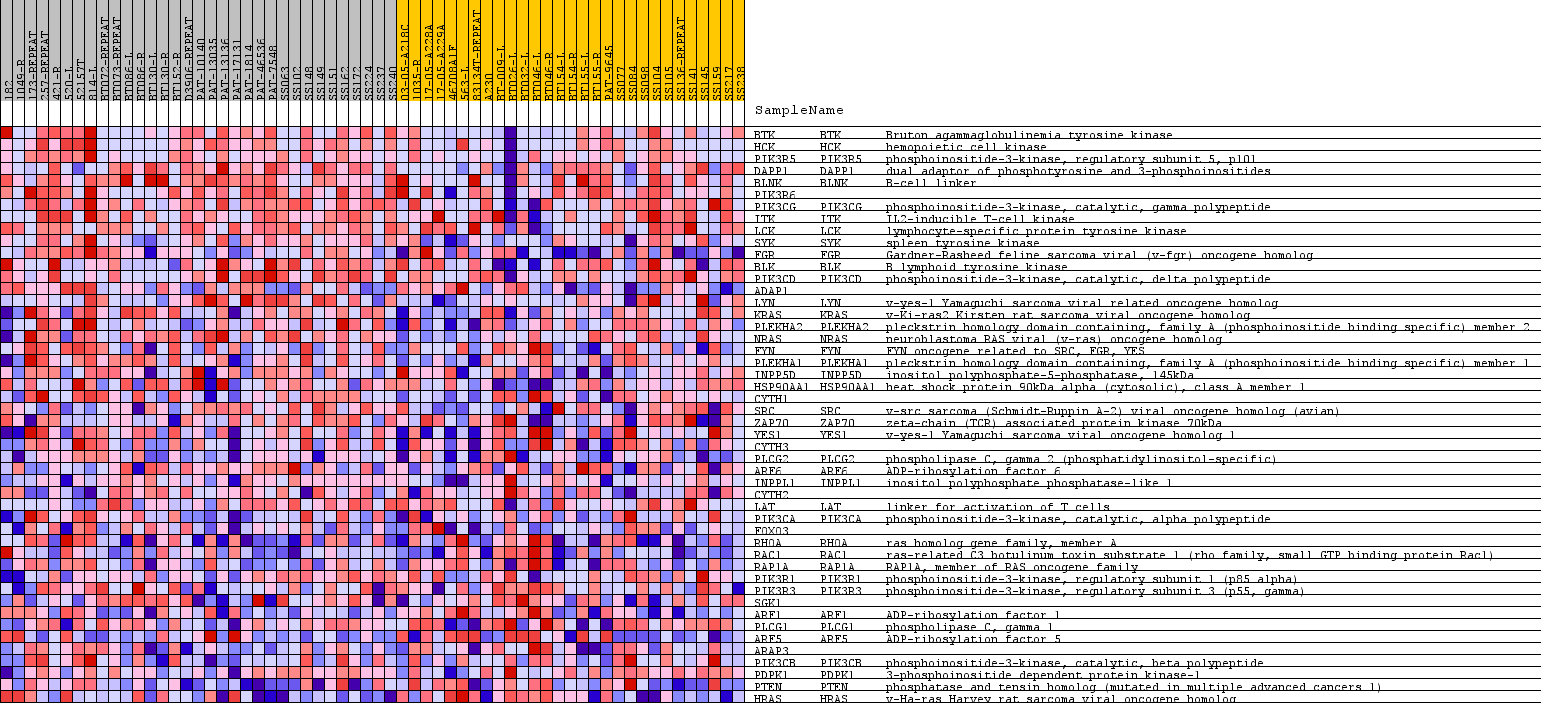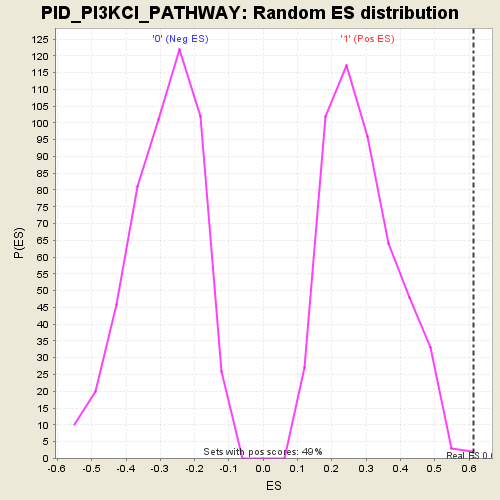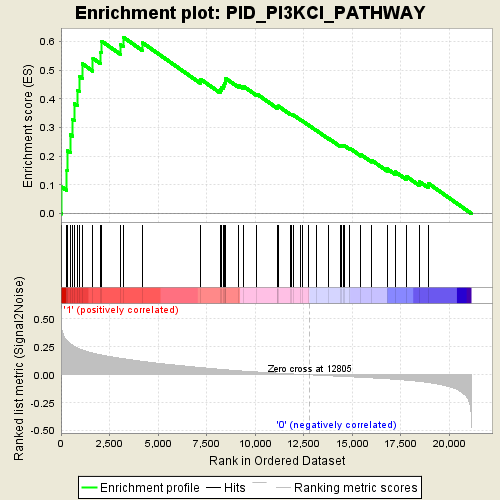
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | NOV0093_collapsed_to_symbols.NOV0093.cls #obese_versus_normal.NOV0093.cls #obese_versus_normal_repos |
| Phenotype | NOV0093.cls#obese_versus_normal_repos |
| Upregulated in class | 1 |
| GeneSet | PID_PI3KCI_PATHWAY |
| Enrichment Score (ES) | 0.61357117 |
| Normalized Enrichment Score (NES) | 2.12354 |
| Nominal p-value | 0.0020325202 |
| FDR q-value | 0.20192112 |
| FWER p-Value | 0.27 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | BTK | BTK Entrez, Source | Bruton agammaglobulinemia tyrosine kinase | 28 | 0.412 | 0.0931 | Yes |
| 2 | HCK | HCK Entrez, Source | hemopoietic cell kinase | 300 | 0.306 | 0.1503 | Yes |
| 3 | PIK3R5 | PIK3R5 Entrez, Source | phosphoinositide-3-kinase, regulatory subunit 5, p101 | 318 | 0.302 | 0.2188 | Yes |
| 4 | DAPP1 | DAPP1 Entrez, Source | dual adaptor of phosphotyrosine and 3-phosphoinositides | 474 | 0.276 | 0.2748 | Yes |
| 5 | BLNK | BLNK Entrez, Source | B-cell linker | 600 | 0.263 | 0.3290 | Yes |
| 6 | PIK3R6 | 687 | 0.253 | 0.3829 | Yes | ||
| 7 | PIK3CG | PIK3CG Entrez, Source | phosphoinositide-3-kinase, catalytic, gamma polypeptide | 853 | 0.238 | 0.4297 | Yes |
| 8 | ITK | ITK Entrez, Source | IL2-inducible T-cell kinase | 965 | 0.231 | 0.4773 | Yes |
| 9 | LCK | LCK Entrez, Source | lymphocyte-specific protein tyrosine kinase | 1096 | 0.222 | 0.5220 | Yes |
| 10 | SYK | SYK Entrez, Source | spleen tyrosine kinase | 1640 | 0.194 | 0.5407 | Yes |
| 11 | FGR | FGR Entrez, Source | Gardner-Rasheed feline sarcoma viral (v-fgr) oncogene homolog | 2034 | 0.178 | 0.5628 | Yes |
| 12 | BLK | BLK Entrez, Source | B lymphoid tyrosine kinase | 2079 | 0.177 | 0.6012 | Yes |
| 13 | PIK3CD | PIK3CD Entrez, Source | phosphoinositide-3-kinase, catalytic, delta polypeptide | 3055 | 0.148 | 0.5888 | Yes |
| 14 | ADAP1 | 3226 | 0.143 | 0.6136 | Yes | ||
| 15 | LYN | LYN Entrez, Source | v-yes-1 Yamaguchi sarcoma viral related oncogene homolog | 4186 | 0.120 | 0.5957 | No |
| 16 | KRAS | KRAS Entrez, Source | v-Ki-ras2 Kirsten rat sarcoma viral oncogene homolog | 7196 | 0.064 | 0.4678 | No |
| 17 | PLEKHA2 | PLEKHA2 Entrez, Source | pleckstrin homology domain containing, family A (phosphoinositide binding specific) member 2 | 8198 | 0.049 | 0.4316 | No |
| 18 | NRAS | NRAS Entrez, Source | neuroblastoma RAS viral (v-ras) oncogene homolog | 8260 | 0.048 | 0.4397 | No |
| 19 | FYN | FYN Entrez, Source | FYN oncogene related to SRC, FGR, YES | 8363 | 0.046 | 0.4454 | No |
| 20 | PLEKHA1 | PLEKHA1 Entrez, Source | pleckstrin homology domain containing, family A (phosphoinositide binding specific) member 1 | 8398 | 0.046 | 0.4544 | No |
| 21 | INPP5D | INPP5D Entrez, Source | inositol polyphosphate-5-phosphatase, 145kDa | 8454 | 0.045 | 0.4621 | No |
| 22 | HSP90AA1 | HSP90AA1 Entrez, Source | heat shock protein 90kDa alpha (cytosolic), class A member 1 | 8482 | 0.045 | 0.4711 | No |
| 23 | CYTH1 | 9158 | 0.036 | 0.4473 | No | ||
| 24 | SRC | SRC Entrez, Source | v-src sarcoma (Schmidt-Ruppin A-2) viral oncogene homolog (avian) | 9410 | 0.033 | 0.4428 | No |
| 25 | ZAP70 | ZAP70 Entrez, Source | zeta-chain (TCR) associated protein kinase 70kDa | 10085 | 0.025 | 0.4166 | No |
| 26 | YES1 | YES1 Entrez, Source | v-yes-1 Yamaguchi sarcoma viral oncogene homolog 1 | 11136 | 0.015 | 0.3703 | No |
| 27 | CYTH3 | 11155 | 0.015 | 0.3729 | No | ||
| 28 | PLCG2 | PLCG2 Entrez, Source | phospholipase C, gamma 2 (phosphatidylinositol-specific) | 11182 | 0.015 | 0.3751 | No |
| 29 | ARF6 | ARF6 Entrez, Source | ADP-ribosylation factor 6 | 11818 | 0.009 | 0.3469 | No |
| 30 | INPPL1 | INPPL1 Entrez, Source | inositol polyphosphate phosphatase-like 1 | 11888 | 0.008 | 0.3455 | No |
| 31 | CYTH2 | 11962 | 0.007 | 0.3437 | No | ||
| 32 | LAT | LAT Entrez, Source | linker for activation of T cells | 12332 | 0.004 | 0.3271 | No |
| 33 | PIK3CA | PIK3CA Entrez, Source | phosphoinositide-3-kinase, catalytic, alpha polypeptide | 12460 | 0.003 | 0.3217 | No |
| 34 | FOXO3 | 12759 | 0.000 | 0.3076 | No | ||
| 35 | RHOA | RHOA Entrez, Source | ras homolog gene family, member A | 13163 | -0.003 | 0.2892 | No |
| 36 | RAC1 | RAC1 Entrez, Source | ras-related C3 botulinum toxin substrate 1 (rho family, small GTP binding protein Rac1) | 13803 | -0.009 | 0.2610 | No |
| 37 | RAP1A | RAP1A Entrez, Source | RAP1A, member of RAS oncogene family | 14406 | -0.014 | 0.2356 | No |
| 38 | PIK3R1 | PIK3R1 Entrez, Source | phosphoinositide-3-kinase, regulatory subunit 1 (p85 alpha) | 14429 | -0.014 | 0.2378 | No |
| 39 | PIK3R3 | PIK3R3 Entrez, Source | phosphoinositide-3-kinase, regulatory subunit 3 (p55, gamma) | 14529 | -0.015 | 0.2364 | No |
| 40 | SGK1 | 14611 | -0.015 | 0.2361 | No | ||
| 41 | ARF1 | ARF1 Entrez, Source | ADP-ribosylation factor 1 | 14865 | -0.018 | 0.2282 | No |
| 42 | PLCG1 | PLCG1 Entrez, Source | phospholipase C, gamma 1 | 15449 | -0.023 | 0.2057 | No |
| 43 | ARF5 | ARF5 Entrez, Source | ADP-ribosylation factor 5 | 16001 | -0.028 | 0.1859 | No |
| 44 | ARAP3 | 16800 | -0.036 | 0.1563 | No | ||
| 45 | PIK3CB | PIK3CB Entrez, Source | phosphoinositide-3-kinase, catalytic, beta polypeptide | 17216 | -0.040 | 0.1458 | No |
| 46 | PDPK1 | PDPK1 Entrez, Source | 3-phosphoinositide dependent protein kinase-1 | 17787 | -0.048 | 0.1298 | No |
| 47 | PTEN | PTEN Entrez, Source | phosphatase and tensin homolog (mutated in multiple advanced cancers 1) | 18451 | -0.059 | 0.1118 | No |
| 48 | HRAS | HRAS Entrez, Source | v-Ha-ras Harvey rat sarcoma viral oncogene homolog | 18923 | -0.069 | 0.1053 | No |