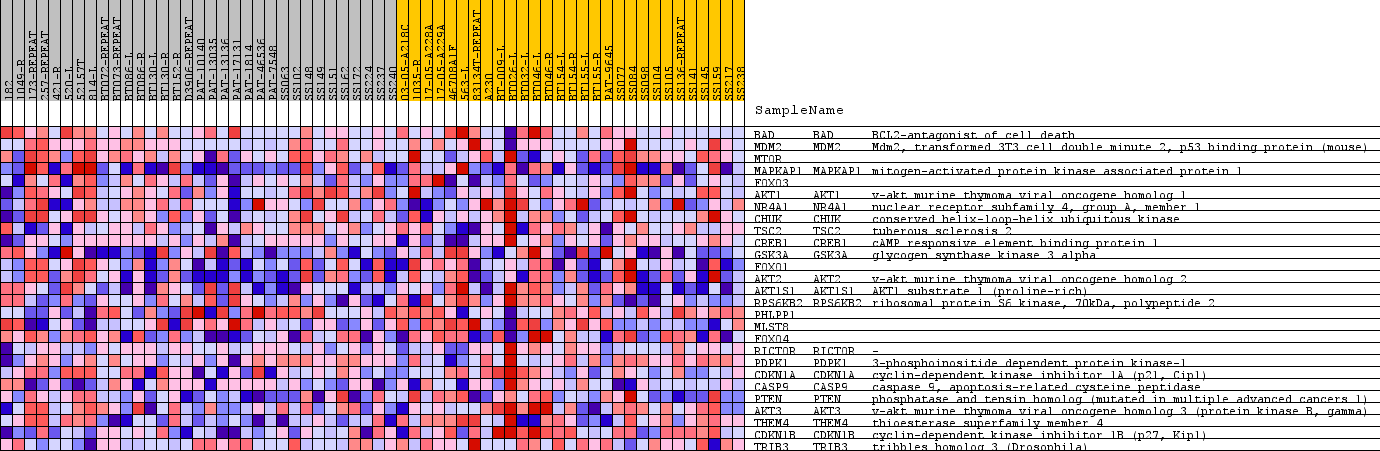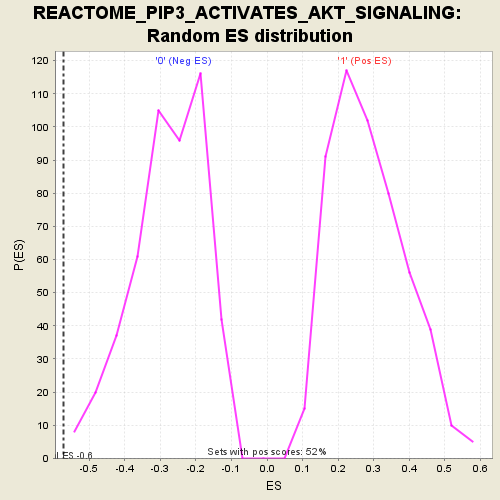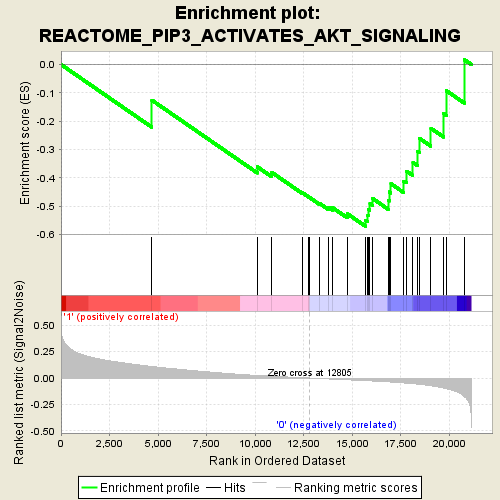
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | NOV0093_collapsed_to_symbols.NOV0093.cls #obese_versus_normal.NOV0093.cls #obese_versus_normal_repos |
| Phenotype | NOV0093.cls#obese_versus_normal_repos |
| Upregulated in class | 0 |
| GeneSet | REACTOME_PIP3_ACTIVATES_AKT_SIGNALING |
| Enrichment Score (ES) | -0.5713217 |
| Normalized Enrichment Score (NES) | -2.047696 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.8367604 |
| FWER p-Value | 0.397 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | BAD | BAD Entrez, Source | BCL2-antagonist of cell death | 4679 | 0.109 | -0.1259 | No |
| 2 | MDM2 | MDM2 Entrez, Source | Mdm2, transformed 3T3 cell double minute 2, p53 binding protein (mouse) | 10115 | 0.025 | -0.3614 | No |
| 3 | MTOR | 10857 | 0.018 | -0.3810 | No | ||
| 4 | MAPKAP1 | MAPKAP1 Entrez, Source | mitogen-activated protein kinase associated protein 1 | 12415 | 0.003 | -0.4520 | No |
| 5 | FOXO3 | 12759 | 0.000 | -0.4679 | No | ||
| 6 | AKT1 | AKT1 Entrez, Source | v-akt murine thymoma viral oncogene homolog 1 | 12791 | 0.000 | -0.4693 | No |
| 7 | NR4A1 | NR4A1 Entrez, Source | nuclear receptor subfamily 4, group A, member 1 | 13322 | -0.004 | -0.4906 | No |
| 8 | CHUK | CHUK Entrez, Source | conserved helix-loop-helix ubiquitous kinase | 13754 | -0.008 | -0.5036 | No |
| 9 | TSC2 | TSC2 Entrez, Source | tuberous sclerosis 2 | 13990 | -0.010 | -0.5057 | No |
| 10 | CREB1 | CREB1 Entrez, Source | cAMP responsive element binding protein 1 | 14742 | -0.017 | -0.5268 | No |
| 11 | GSK3A | GSK3A Entrez, Source | glycogen synthase kinase 3 alpha | 15684 | -0.025 | -0.5496 | Yes |
| 12 | FOXO1 | 15805 | -0.026 | -0.5326 | Yes | ||
| 13 | AKT2 | AKT2 Entrez, Source | v-akt murine thymoma viral oncogene homolog 2 | 15825 | -0.026 | -0.5106 | Yes |
| 14 | AKT1S1 | AKT1S1 Entrez, Source | AKT1 substrate 1 (proline-rich) | 15914 | -0.027 | -0.4913 | Yes |
| 15 | RPS6KB2 | RPS6KB2 Entrez, Source | ribosomal protein S6 kinase, 70kDa, polypeptide 2 | 16047 | -0.028 | -0.4730 | Yes |
| 16 | PHLPP1 | 16853 | -0.036 | -0.4795 | Yes | ||
| 17 | MLST8 | 16908 | -0.037 | -0.4499 | Yes | ||
| 18 | FOXO4 | 16996 | -0.038 | -0.4211 | Yes | ||
| 19 | RICTOR | RICTOR Entrez, Source | - | 17656 | -0.046 | -0.4121 | Yes |
| 20 | PDPK1 | PDPK1 Entrez, Source | 3-phosphoinositide dependent protein kinase-1 | 17787 | -0.048 | -0.3762 | Yes |
| 21 | CDKN1A | CDKN1A Entrez, Source | cyclin-dependent kinase inhibitor 1A (p21, Cip1) | 18129 | -0.053 | -0.3459 | Yes |
| 22 | CASP9 | CASP9 Entrez, Source | caspase 9, apoptosis-related cysteine peptidase | 18355 | -0.057 | -0.3067 | Yes |
| 23 | PTEN | PTEN Entrez, Source | phosphatase and tensin homolog (mutated in multiple advanced cancers 1) | 18451 | -0.059 | -0.2598 | Yes |
| 24 | AKT3 | AKT3 Entrez, Source | v-akt murine thymoma viral oncogene homolog 3 (protein kinase B, gamma) | 19056 | -0.072 | -0.2251 | Yes |
| 25 | THEM4 | THEM4 Entrez, Source | thioesterase superfamily member 4 | 19713 | -0.094 | -0.1734 | Yes |
| 26 | CDKN1B | CDKN1B Entrez, Source | cyclin-dependent kinase inhibitor 1B (p27, Kip1) | 19857 | -0.100 | -0.0927 | Yes |
| 27 | TRIB3 | TRIB3 Entrez, Source | tribbles homolog 3 (Drosophila) | 20776 | -0.175 | 0.0174 | Yes |