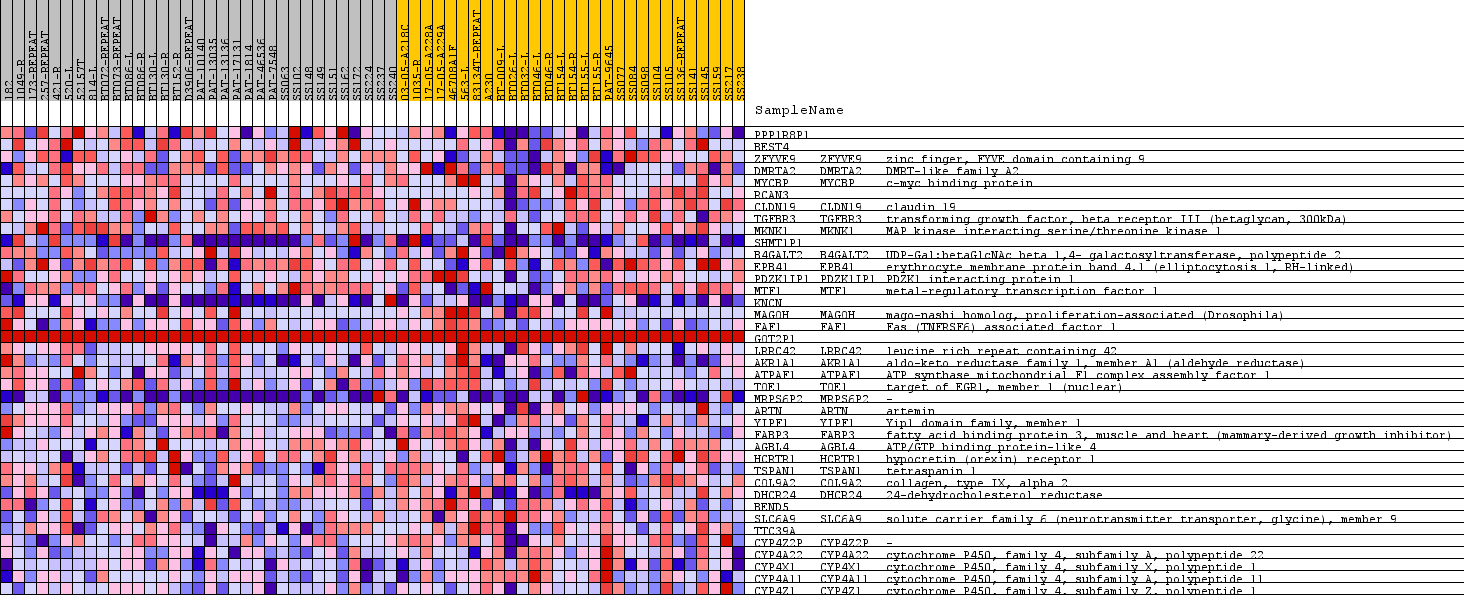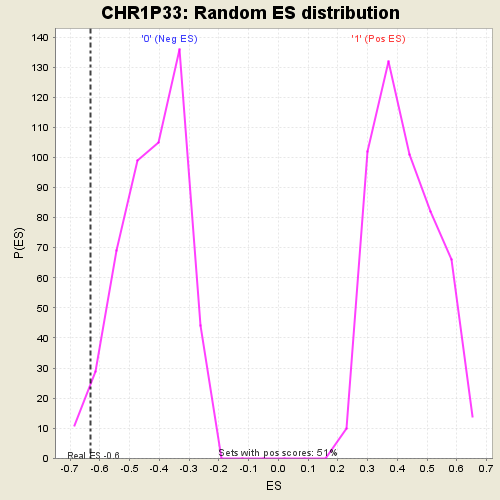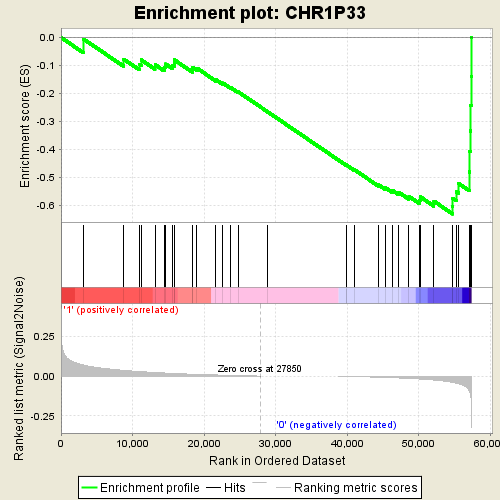
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | NOV0093_D0_fat_corrected_collapsed_to_symbols.NOV0093.cls #obese_versus_normal.NOV0093.cls #obese_versus_normal_repos |
| Phenotype | NOV0093.cls#obese_versus_normal_repos |
| Upregulated in class | 0 |
| GeneSet | CHR1P33 |
| Enrichment Score (ES) | -0.6314566 |
| Normalized Enrichment Score (NES) | -1.4962164 |
| Nominal p-value | 0.03448276 |
| FDR q-value | 1.0 |
| FWER p-Value | 1.0 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | PPP1R8P1 | 3167 | 0.069 | -0.0058 | No | ||
| 2 | BEST4 | 8779 | 0.036 | -0.0778 | No | ||
| 3 | ZFYVE9 | ZFYVE9 Entrez, Source | zinc finger, FYVE domain containing 9 | 10934 | 0.029 | -0.0949 | No |
| 4 | DMRTA2 | DMRTA2 Entrez, Source | DMRT-like family A2 | 11205 | 0.028 | -0.0797 | No |
| 5 | MYCBP | MYCBP Entrez, Source | c-myc binding protein | 13125 | 0.023 | -0.0968 | No |
| 6 | RCAN3 | 14429 | 0.020 | -0.1055 | No | ||
| 7 | CLDN19 | CLDN19 Entrez, Source | claudin 19 | 14599 | 0.019 | -0.0946 | No |
| 8 | TGFBR3 | TGFBR3 Entrez, Source | transforming growth factor, beta receptor III (betaglycan, 300kDa) | 15585 | 0.017 | -0.0995 | No |
| 9 | MKNK1 | MKNK1 Entrez, Source | MAP kinase interacting serine/threonine kinase 1 | 15798 | 0.017 | -0.0912 | No |
| 10 | SHMT1P1 | 15825 | 0.017 | -0.0797 | No | ||
| 11 | B4GALT2 | B4GALT2 Entrez, Source | UDP-Gal:betaGlcNAc beta 1,4- galactosyltransferase, polypeptide 2 | 18337 | 0.012 | -0.1149 | No |
| 12 | EPB41 | EPB41 Entrez, Source | erythrocyte membrane protein band 4.1 (elliptocytosis 1, RH-linked) | 18361 | 0.012 | -0.1066 | No |
| 13 | PDZK1IP1 | PDZK1IP1 Entrez, Source | PDZK1 interacting protein 1 | 18985 | 0.011 | -0.1095 | No |
| 14 | MTF1 | MTF1 Entrez, Source | metal-regulatory transcription factor 1 | 21647 | 0.007 | -0.1507 | No |
| 15 | KNCN | 22577 | 0.006 | -0.1626 | No | ||
| 16 | MAGOH | MAGOH Entrez, Source | mago-nashi homolog, proliferation-associated (Drosophila) | 23710 | 0.005 | -0.1788 | No |
| 17 | FAF1 | FAF1 Entrez, Source | Fas (TNFRSF6) associated factor 1 | 24778 | 0.004 | -0.1945 | No |
| 18 | GOT2P1 | 28904 | 0.000 | -0.2665 | No | ||
| 19 | LRRC42 | LRRC42 Entrez, Source | leucine rich repeat containing 42 | 39834 | -0.001 | -0.4562 | No |
| 20 | AKR1A1 | AKR1A1 Entrez, Source | aldo-keto reductase family 1, member A1 (aldehyde reductase) | 39849 | -0.001 | -0.4554 | No |
| 21 | ATPAF1 | ATPAF1 Entrez, Source | ATP synthase mitochondrial F1 complex assembly factor 1 | 41021 | -0.003 | -0.4735 | No |
| 22 | TOE1 | TOE1 Entrez, Source | target of EGR1, member 1 (nuclear) | 44399 | -0.007 | -0.5273 | No |
| 23 | MRPS6P2 | MRPS6P2 Entrez, Source | - | 45356 | -0.009 | -0.5378 | No |
| 24 | ARTN | ARTN Entrez, Source | artemin | 46356 | -0.010 | -0.5479 | No |
| 25 | YIPF1 | YIPF1 Entrez, Source | Yip1 domain family, member 1 | 47161 | -0.012 | -0.5536 | No |
| 26 | FABP3 | FABP3 Entrez, Source | fatty acid binding protein 3, muscle and heart (mammary-derived growth inhibitor) | 48617 | -0.015 | -0.5686 | No |
| 27 | AGBL4 | AGBL4 Entrez, Source | ATP/GTP binding protein-like 4 | 50111 | -0.018 | -0.5816 | No |
| 28 | HCRTR1 | HCRTR1 Entrez, Source | hypocretin (orexin) receptor 1 | 50211 | -0.018 | -0.5701 | No |
| 29 | TSPAN1 | TSPAN1 Entrez, Source | tetraspanin 1 | 52107 | -0.024 | -0.5856 | No |
| 30 | COL9A2 | COL9A2 Entrez, Source | collagen, type IX, alpha 2 | 54736 | -0.039 | -0.6031 | Yes |
| 31 | DHCR24 | DHCR24 Entrez, Source | 24-dehydrocholesterol reductase | 54759 | -0.040 | -0.5751 | Yes |
| 32 | BEND5 | 55301 | -0.045 | -0.5525 | Yes | ||
| 33 | SLC6A9 | SLC6A9 Entrez, Source | solute carrier family 6 (neurotransmitter transporter, glycine), member 9 | 55575 | -0.048 | -0.5227 | Yes |
| 34 | TTC39A | 57078 | -0.095 | -0.4809 | Yes | ||
| 35 | CYP4Z2P | CYP4Z2P Entrez, Source | - | 57133 | -0.102 | -0.4084 | Yes |
| 36 | CYP4A22 | CYP4A22 Entrez, Source | cytochrome P450, family 4, subfamily A, polypeptide 22 | 57166 | -0.105 | -0.3336 | Yes |
| 37 | CYP4X1 | CYP4X1 Entrez, Source | cytochrome P450, family 4, subfamily X, polypeptide 1 | 57269 | -0.130 | -0.2423 | Yes |
| 38 | CYP4A11 | CYP4A11 Entrez, Source | cytochrome P450, family 4, subfamily A, polypeptide 11 | 57302 | -0.146 | -0.1378 | Yes |
| 39 | CYP4Z1 | CYP4Z1 Entrez, Source | cytochrome P450, family 4, subfamily Z, polypeptide 1 | 57334 | -0.193 | 0.0003 | Yes |