Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | NOV0093_D0_fat_corrected_collapsed_to_symbols.NOV0093.cls #obese_versus_normal.NOV0093.cls #obese_versus_normal_repos |
| Phenotype | NOV0093.cls#obese_versus_normal_repos |
| Upregulated in class | 0 |
| GeneSet | MODULE_318 |
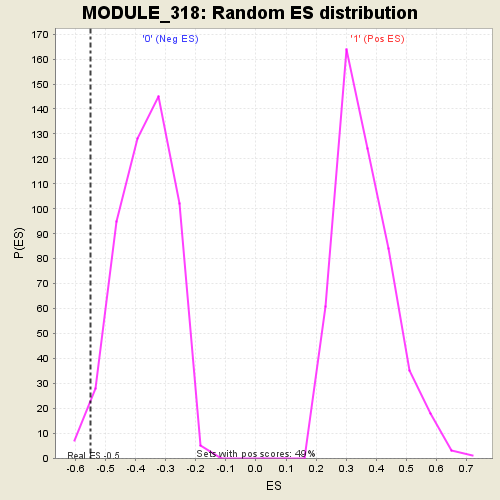
| Enrichment Score (ES) | -0.54888445 |
| Normalized Enrichment Score (NES) | -1.5027806 |
| Nominal p-value | 0.021568628 |
| FDR q-value | 1.0 |
| FWER p-Value | 0.999 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | HELZ | HELZ Entrez, Source | helicase with zinc finger | 12156 | 0.025 | -0.1587 | No |
| 2 | CHD4 | CHD4 Entrez, Source | chromodomain helicase DNA binding protein 4 | 12367 | 0.025 | -0.1100 | No |
| 3 | WRN | WRN Entrez, Source | Werner syndrome | 14165 | 0.020 | -0.0985 | No |
| 4 | GTF2F2 | GTF2F2 Entrez, Source | general transcription factor IIF, polypeptide 2, 30kDa | 18311 | 0.012 | -0.1450 | No |
| 5 | MCM6 | MCM6 Entrez, Source | MCM6 minichromosome maintenance deficient 6 (MIS5 homolog, S. pombe) (S. cerevisiae) | 20413 | 0.009 | -0.1626 | No |
| 6 | CHD3 | CHD3 Entrez, Source | chromodomain helicase DNA binding protein 3 | 21594 | 0.007 | -0.1676 | No |
| 7 | MCM3 | MCM3 Entrez, Source | MCM3 minichromosome maintenance deficient 3 (S. cerevisiae) | 22423 | 0.006 | -0.1686 | No |
| 8 | STAT2 | STAT2 Entrez, Source | signal transducer and activator of transcription 2, 113kDa | 22453 | 0.006 | -0.1557 | No |
| 9 | DHX9 | DHX9 Entrez, Source | DEAH (Asp-Glu-Ala-His) box polypeptide 9 | 22561 | 0.006 | -0.1445 | No |
| 10 | BLM | BLM Entrez, Source | Bloom syndrome | 22741 | 0.006 | -0.1351 | No |
| 11 | DDX5 | DDX5 Entrez, Source | DEAD (Asp-Glu-Ala-Asp) box polypeptide 5 | 23009 | 0.006 | -0.1277 | No |
| 12 | RAD51C | RAD51C Entrez, Source | RAD51 homolog C (S. cerevisiae) | 23051 | 0.006 | -0.1165 | No |
| 13 | DDX10 | DDX10 Entrez, Source | DEAD (Asp-Glu-Ala-Asp) box polypeptide 10 | 24547 | 0.004 | -0.1338 | No |
| 14 | RECQL4 | RECQL4 Entrez, Source | RecQ protein-like 4 | 26667 | 0.002 | -0.1672 | No |
| 15 | RAD51 | RAD51 Entrez, Source | RAD51 homolog (RecA homolog, E. coli) (S. cerevisiae) | 27179 | 0.001 | -0.1743 | No |
| 16 | XRCC5 | XRCC5 Entrez, Source | X-ray repair complementing defective repair in Chinese hamster cells 5 (double-strand-break rejoining; Ku autoantigen, 80kDa) | 27228 | 0.001 | -0.1734 | No |
| 17 | RAD51D | 39694 | -0.001 | -0.3882 | No | ||
| 18 | XRCC3 | XRCC3 Entrez, Source | X-ray repair complementing defective repair in Chinese hamster cells 3 | 42236 | -0.005 | -0.4228 | No |
| 19 | XRCC6 | XRCC6 Entrez, Source | X-ray repair complementing defective repair in Chinese hamster cells 6 (Ku autoantigen, 70kDa) | 44250 | -0.007 | -0.4430 | No |
| 20 | ERCC3 | ERCC3 Entrez, Source | excision repair cross-complementing rodent repair deficiency, complementation group 3 (xeroderma pigmentosum group B complementing) | 45168 | -0.008 | -0.4414 | No |
| 21 | ZNF254 | ZNF254 Entrez, Source | zinc finger protein 254 | 45358 | -0.009 | -0.4265 | No |
| 22 | FOS | FOS Entrez, Source | v-fos FBJ murine osteosarcoma viral oncogene homolog | 52375 | -0.026 | -0.4946 | Yes |
| 23 | RAD51B | 53406 | -0.030 | -0.4482 | Yes | ||
| 24 | JUNB | JUNB Entrez, Source | jun B proto-oncogene | 54479 | -0.037 | -0.3877 | Yes |
| 25 | CITED2 | CITED2 Entrez, Source | Cbp/p300-interacting transactivator, with Glu/Asp-rich carboxy-terminal domain, 2 | 55274 | -0.044 | -0.3074 | Yes |
| 26 | MYCN | MYCN Entrez, Source | v-myc myelocytomatosis viral related oncogene, neuroblastoma derived (avian) | 56695 | -0.071 | -0.1806 | Yes |
| 27 | DDX11 | DDX11 Entrez, Source | DEAD/H (Asp-Glu-Ala-Asp/His) box polypeptide 11 (CHL1-like helicase homolog, S. cerevisiae) | 57030 | -0.090 | 0.0056 | Yes |