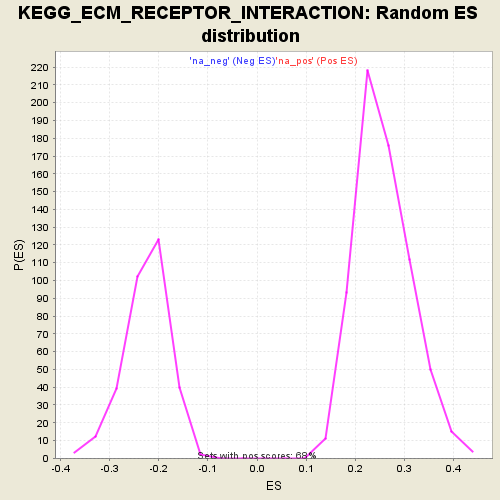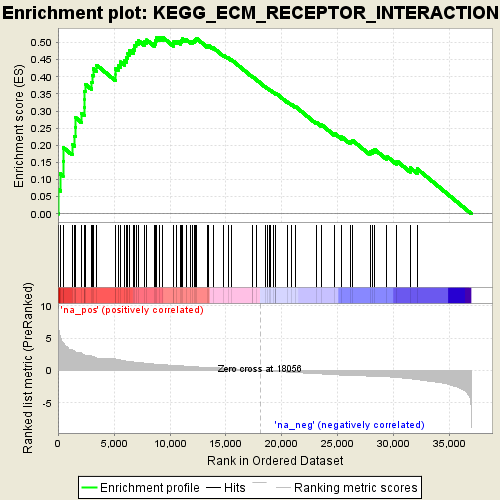
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | NOV0135_D11_log2foldchange_preranked |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_pos |
| GeneSet | KEGG_ECM_RECEPTOR_INTERACTION |
| Enrichment Score (ES) | 0.51490927 |
| Normalized Enrichment Score (NES) | 1.9983009 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.0059624705 |
| FWER p-Value | 0.079 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | ITGA11 | 23 | 7.335 | 0.0712 | Yes | ||
| 2 | CD44 | 202 | 5.144 | 0.1168 | Yes | ||
| 3 | COL3A1 | 444 | 4.306 | 0.1524 | Yes | ||
| 4 | COL5A3 | 469 | 4.243 | 0.1933 | Yes | ||
| 5 | FN1 | 1255 | 3.216 | 0.2035 | Yes | ||
| 6 | COL5A1 | 1472 | 3.013 | 0.2271 | Yes | ||
| 7 | COMP | 1555 | 2.891 | 0.2532 | Yes | ||
| 8 | ITGA10 | 1582 | 2.841 | 0.2803 | Yes | ||
| 9 | ITGB3 | 2077 | 2.717 | 0.2935 | Yes | ||
| 10 | COL1A2 | 2328 | 2.443 | 0.3106 | Yes | ||
| 11 | COL6A3 | 2337 | 2.421 | 0.3341 | Yes | ||
| 12 | SPP1 | 2343 | 2.416 | 0.3577 | Yes | ||
| 13 | COL6A2 | 2413 | 2.363 | 0.3789 | Yes | ||
| 14 | VTN | 3008 | 2.239 | 0.3847 | Yes | ||
| 15 | ITGA4 | 3094 | 2.181 | 0.4038 | Yes | ||
| 16 | COL1A1 | 3148 | 2.121 | 0.4231 | Yes | ||
| 17 | ITGA3 | 3450 | 1.946 | 0.4340 | Yes | ||
| 18 | ITGA1 | 5116 | 1.782 | 0.4063 | Yes | ||
| 19 | COL4A1 | 5119 | 1.780 | 0.4236 | Yes | ||
| 20 | ITGA5 | 5362 | 1.671 | 0.4334 | Yes | ||
| 21 | COL4A2 | 5574 | 1.601 | 0.4434 | Yes | ||
| 22 | LAMC2 | 5931 | 1.498 | 0.4484 | Yes | ||
| 23 | LAMC3 | 6145 | 1.432 | 0.4566 | Yes | ||
| 24 | COL6A1 | 6238 | 1.407 | 0.4679 | Yes | ||
| 25 | LAMB3 | 6408 | 1.364 | 0.4767 | Yes | ||
| 26 | THBS2 | 6744 | 1.310 | 0.4804 | Yes | ||
| 27 | VWF | 6846 | 1.287 | 0.4903 | Yes | ||
| 28 | ITGA9 | 6962 | 1.264 | 0.4995 | Yes | ||
| 29 | COL11A2 | 7169 | 1.218 | 0.5059 | Yes | ||
| 30 | GP5 | 7710 | 1.136 | 0.5024 | Yes | ||
| 31 | TNC | 7915 | 1.104 | 0.5076 | Yes | ||
| 32 | SV2C | 8603 | 0.990 | 0.4987 | Yes | ||
| 33 | SDC2 | 8720 | 0.972 | 0.5050 | Yes | ||
| 34 | LAMA4 | 8753 | 0.966 | 0.5136 | Yes | ||
| 35 | LAMC1 | 9059 | 0.924 | 0.5144 | Yes | ||
| 36 | THBS1 | 9361 | 0.885 | 0.5149 | Yes | ||
| 37 | SDC3 | 10316 | 0.783 | 0.4967 | No | ||
| 38 | TNN | 10349 | 0.780 | 0.5034 | No | ||
| 39 | AGRN | 10607 | 0.752 | 0.5038 | No | ||
| 40 | ITGB4 | 10912 | 0.726 | 0.5027 | No | ||
| 41 | DAG1 | 11016 | 0.712 | 0.5069 | No | ||
| 42 | ITGA2 | 11125 | 0.697 | 0.5108 | No | ||
| 43 | LAMA3 | 11453 | 0.662 | 0.5084 | No | ||
| 44 | LAMB2 | 11830 | 0.614 | 0.5042 | No | ||
| 45 | ITGB7 | 12042 | 0.594 | 0.5043 | No | ||
| 46 | THBS3 | 12182 | 0.579 | 0.5062 | No | ||
| 47 | HSPG2 | 12283 | 0.565 | 0.5090 | No | ||
| 48 | ITGB1 | 12370 | 0.555 | 0.5121 | No | ||
| 49 | ITGB8 | 13335 | 0.476 | 0.4906 | No | ||
| 50 | SDC4 | 13476 | 0.461 | 0.4913 | No | ||
| 51 | SDC1 | 13852 | 0.417 | 0.4852 | No | ||
| 52 | ITGA8 | 14808 | 0.345 | 0.4627 | No | ||
| 53 | ITGB5 | 15192 | 0.303 | 0.4552 | No | ||
| 54 | SV2B | 15492 | 0.276 | 0.4498 | No | ||
| 55 | COL11A1 | 17329 | 0.081 | 0.4008 | No | ||
| 56 | ITGA6 | 17332 | 0.081 | 0.4015 | No | ||
| 57 | GP6 | 17754 | 0.036 | 0.3904 | No | ||
| 58 | LAMA1 | 18555 | -0.051 | 0.3692 | No | ||
| 59 | ITGAV | 18713 | -0.069 | 0.3656 | No | ||
| 60 | LAMA5 | 18877 | -0.085 | 0.3620 | No | ||
| 61 | RELN | 18953 | -0.090 | 0.3609 | No | ||
| 62 | COL6A6 | 19212 | -0.116 | 0.3550 | No | ||
| 63 | SV2A | 19429 | -0.137 | 0.3505 | No | ||
| 64 | COL4A6 | 19436 | -0.138 | 0.3517 | No | ||
| 65 | CD36 | 19461 | -0.141 | 0.3524 | No | ||
| 66 | CHAD | 20501 | -0.243 | 0.3266 | No | ||
| 67 | COL4A4 | 20885 | -0.282 | 0.3190 | No | ||
| 68 | COL2A1 | 21231 | -0.315 | 0.3127 | No | ||
| 69 | ITGA7 | 23074 | -0.492 | 0.2675 | No | ||
| 70 | TNR | 23510 | -0.530 | 0.2609 | No | ||
| 71 | COL5A2 | 24734 | -0.655 | 0.2341 | No | ||
| 72 | GP1BA | 25332 | -0.719 | 0.2249 | No | ||
| 73 | LAMB1 | 26097 | -0.806 | 0.2121 | No | ||
| 74 | LAMB4 | 26273 | -0.826 | 0.2155 | No | ||
| 75 | HMMR | 27871 | -0.887 | 0.1808 | No | ||
| 76 | LAMA2 | 28079 | -0.914 | 0.1841 | No | ||
| 77 | ITGB6 | 28305 | -0.947 | 0.1873 | No | ||
| 78 | TNXB | 29385 | -0.973 | 0.1675 | No | ||
| 79 | THBS4 | 30275 | -1.084 | 0.1540 | No | ||
| 80 | ITGA2B | 31462 | -1.275 | 0.1343 | No | ||
| 81 | CD47 | 32088 | -1.414 | 0.1312 | No |