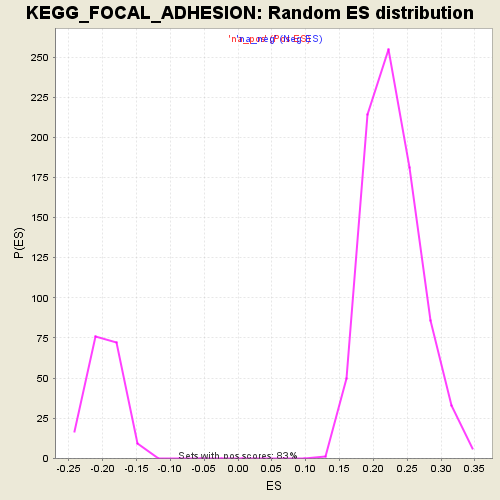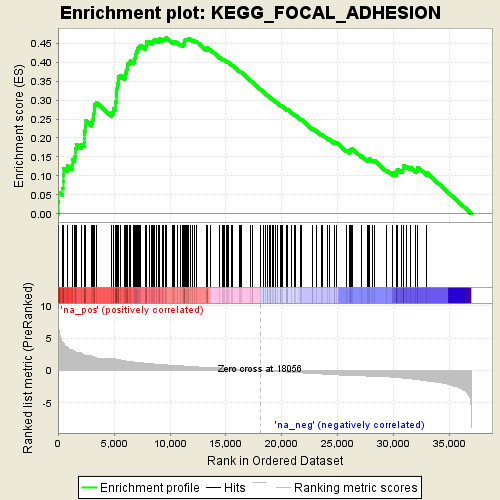
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | NOV0135_D11_log2foldchange_preranked |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_pos |
| GeneSet | KEGG_FOCAL_ADHESION |
| Enrichment Score (ES) | 0.46512848 |
| Normalized Enrichment Score (NES) | 2.0413134 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.0039823786 |
| FWER p-Value | 0.044 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | ITGA11 | 23 | 7.335 | 0.0314 | Yes | ||
| 2 | PIK3CG | 77 | 6.226 | 0.0572 | Yes | ||
| 3 | RAC2 | 415 | 4.352 | 0.0671 | Yes | ||
| 4 | COL3A1 | 444 | 4.306 | 0.0851 | Yes | ||
| 5 | COL5A3 | 469 | 4.243 | 0.1030 | Yes | ||
| 6 | IGF1 | 516 | 4.178 | 0.1200 | Yes | ||
| 7 | CAV1 | 808 | 3.647 | 0.1280 | Yes | ||
| 8 | FN1 | 1255 | 3.216 | 0.1300 | Yes | ||
| 9 | PARVG | 1269 | 3.190 | 0.1436 | Yes | ||
| 10 | COL5A1 | 1472 | 3.013 | 0.1512 | Yes | ||
| 11 | COMP | 1555 | 2.891 | 0.1616 | Yes | ||
| 12 | ITGA10 | 1582 | 2.841 | 0.1733 | Yes | ||
| 13 | CAV2 | 1678 | 2.742 | 0.1827 | Yes | ||
| 14 | ITGB3 | 2077 | 2.717 | 0.1838 | Yes | ||
| 15 | COL1A2 | 2328 | 2.443 | 0.1877 | Yes | ||
| 16 | COL6A3 | 2337 | 2.421 | 0.1980 | Yes | ||
| 17 | SPP1 | 2343 | 2.416 | 0.2085 | Yes | ||
| 18 | KDR | 2360 | 2.393 | 0.2185 | Yes | ||
| 19 | COL6A2 | 2413 | 2.363 | 0.2274 | Yes | ||
| 20 | MYL9 | 2439 | 2.323 | 0.2369 | Yes | ||
| 21 | FLNA | 2491 | 2.297 | 0.2455 | Yes | ||
| 22 | VTN | 3008 | 2.239 | 0.2413 | Yes | ||
| 23 | ITGA4 | 3094 | 2.181 | 0.2485 | Yes | ||
| 24 | COL1A1 | 3148 | 2.121 | 0.2563 | Yes | ||
| 25 | PDGFRB | 3203 | 2.095 | 0.2640 | Yes | ||
| 26 | CCND3 | 3214 | 2.085 | 0.2728 | Yes | ||
| 27 | PDGFB | 3261 | 2.051 | 0.2806 | Yes | ||
| 28 | FLT1 | 3266 | 2.049 | 0.2894 | Yes | ||
| 29 | ITGA3 | 3450 | 1.946 | 0.2929 | Yes | ||
| 30 | VEGFC | 4748 | 1.925 | 0.2660 | Yes | ||
| 31 | PDGFRA | 4926 | 1.860 | 0.2693 | Yes | ||
| 32 | BIRC3 | 4951 | 1.836 | 0.2767 | Yes | ||
| 33 | ZYX | 5109 | 1.784 | 0.2802 | Yes | ||
| 34 | ITGA1 | 5116 | 1.782 | 0.2879 | Yes | ||
| 35 | COL4A1 | 5119 | 1.780 | 0.2956 | Yes | ||
| 36 | BAD | 5181 | 1.749 | 0.3016 | Yes | ||
| 37 | ACTB | 5182 | 1.748 | 0.3092 | Yes | ||
| 38 | PRKCA | 5198 | 1.740 | 0.3164 | Yes | ||
| 39 | SHC1 | 5249 | 1.720 | 0.3226 | Yes | ||
| 40 | ACTG1 | 5260 | 1.715 | 0.3298 | Yes | ||
| 41 | PPP1CA | 5284 | 1.708 | 0.3366 | Yes | ||
| 42 | HGF | 5322 | 1.689 | 0.3430 | Yes | ||
| 43 | ITGA5 | 5362 | 1.671 | 0.3493 | Yes | ||
| 44 | BCAR1 | 5403 | 1.655 | 0.3554 | Yes | ||
| 45 | HRAS | 5413 | 1.652 | 0.3624 | Yes | ||
| 46 | COL4A2 | 5574 | 1.601 | 0.3650 | Yes | ||
| 47 | LAMC2 | 5931 | 1.498 | 0.3619 | Yes | ||
| 48 | MYLK3 | 5997 | 1.473 | 0.3665 | Yes | ||
| 49 | MYL7 | 6013 | 1.469 | 0.3726 | Yes | ||
| 50 | LAMC3 | 6145 | 1.432 | 0.3753 | Yes | ||
| 51 | MYLPF | 6154 | 1.430 | 0.3813 | Yes | ||
| 52 | VASP | 6169 | 1.426 | 0.3871 | Yes | ||
| 53 | TLN1 | 6213 | 1.413 | 0.3922 | Yes | ||
| 54 | COL6A1 | 6238 | 1.407 | 0.3976 | Yes | ||
| 55 | LAMB3 | 6408 | 1.364 | 0.3990 | Yes | ||
| 56 | ACTN1 | 6444 | 1.359 | 0.4040 | Yes | ||
| 57 | THBS2 | 6744 | 1.310 | 0.4016 | Yes | ||
| 58 | ELK1 | 6801 | 1.296 | 0.4057 | Yes | ||
| 59 | VWF | 6846 | 1.287 | 0.4102 | Yes | ||
| 60 | MYLK2 | 6883 | 1.280 | 0.4148 | Yes | ||
| 61 | CAV3 | 6905 | 1.275 | 0.4198 | Yes | ||
| 62 | ITGA9 | 6962 | 1.264 | 0.4238 | Yes | ||
| 63 | AKT1 | 6971 | 1.262 | 0.4291 | Yes | ||
| 64 | MAPK3 | 7075 | 1.240 | 0.4317 | Yes | ||
| 65 | RAC3 | 7089 | 1.236 | 0.4367 | Yes | ||
| 66 | COL11A2 | 7169 | 1.218 | 0.4399 | Yes | ||
| 67 | MYL10 | 7286 | 1.202 | 0.4420 | Yes | ||
| 68 | ACTN4 | 7372 | 1.199 | 0.4449 | Yes | ||
| 69 | PARVA | 7810 | 1.120 | 0.4379 | Yes | ||
| 70 | FLNB | 7819 | 1.118 | 0.4426 | Yes | ||
| 71 | JUN | 7866 | 1.109 | 0.4462 | Yes | ||
| 72 | RHOA | 7894 | 1.106 | 0.4503 | Yes | ||
| 73 | TNC | 7915 | 1.104 | 0.4546 | Yes | ||
| 74 | SHC3 | 8134 | 1.066 | 0.4533 | Yes | ||
| 75 | PIK3CD | 8342 | 1.031 | 0.4522 | Yes | ||
| 76 | PAK4 | 8430 | 1.018 | 0.4543 | Yes | ||
| 77 | PGF | 8500 | 1.007 | 0.4568 | Yes | ||
| 78 | GRB2 | 8587 | 0.992 | 0.4588 | Yes | ||
| 79 | LAMA4 | 8753 | 0.966 | 0.4585 | Yes | ||
| 80 | TLN2 | 8980 | 0.938 | 0.4564 | Yes | ||
| 81 | LAMC1 | 9059 | 0.924 | 0.4584 | Yes | ||
| 82 | CAPN2 | 9067 | 0.923 | 0.4622 | Yes | ||
| 83 | THBS1 | 9361 | 0.885 | 0.4581 | Yes | ||
| 84 | PIP5K1C | 9401 | 0.880 | 0.4609 | Yes | ||
| 85 | PXN | 9579 | 0.858 | 0.4598 | Yes | ||
| 86 | PARVB | 9590 | 0.857 | 0.4633 | Yes | ||
| 87 | RAPGEF1 | 9660 | 0.850 | 0.4651 | Yes | ||
| 88 | MYL12A | 10263 | 0.788 | 0.4522 | No | ||
| 89 | TNN | 10349 | 0.780 | 0.4533 | No | ||
| 90 | VEGFB | 10412 | 0.773 | 0.4550 | No | ||
| 91 | PDGFA | 10628 | 0.749 | 0.4524 | No | ||
| 92 | ITGB4 | 10912 | 0.726 | 0.4479 | No | ||
| 93 | ITGA2 | 11125 | 0.697 | 0.4451 | No | ||
| 94 | PIK3R2 | 11165 | 0.694 | 0.4471 | No | ||
| 95 | ERBB2 | 11236 | 0.688 | 0.4482 | No | ||
| 96 | PIK3R3 | 11277 | 0.683 | 0.4501 | No | ||
| 97 | FLT4 | 11287 | 0.682 | 0.4528 | No | ||
| 98 | ILK | 11325 | 0.678 | 0.4548 | No | ||
| 99 | VCL | 11326 | 0.678 | 0.4578 | No | ||
| 100 | DIAPH1 | 11372 | 0.671 | 0.4595 | No | ||
| 101 | LAMA3 | 11453 | 0.662 | 0.4602 | No | ||
| 102 | EGFR | 11542 | 0.651 | 0.4606 | No | ||
| 103 | FLNC | 11634 | 0.641 | 0.4610 | No | ||
| 104 | AKT2 | 11660 | 0.638 | 0.4631 | No | ||
| 105 | LAMB2 | 11830 | 0.614 | 0.4612 | No | ||
| 106 | ITGB7 | 12042 | 0.594 | 0.4580 | No | ||
| 107 | THBS3 | 12182 | 0.579 | 0.4567 | No | ||
| 108 | ITGB1 | 12370 | 0.555 | 0.4541 | No | ||
| 109 | VAV2 | 13229 | 0.489 | 0.4329 | No | ||
| 110 | SRC | 13273 | 0.485 | 0.4338 | No | ||
| 111 | ITGB8 | 13335 | 0.476 | 0.4342 | No | ||
| 112 | RAF1 | 13339 | 0.476 | 0.4362 | No | ||
| 113 | RASGRF1 | 13347 | 0.475 | 0.4381 | No | ||
| 114 | CDC42 | 13591 | 0.448 | 0.4335 | No | ||
| 115 | PDGFC | 14435 | 0.386 | 0.4122 | No | ||
| 116 | RAC1 | 14658 | 0.363 | 0.4077 | No | ||
| 117 | ITGA8 | 14808 | 0.345 | 0.4052 | No | ||
| 118 | FYN | 14842 | 0.340 | 0.4058 | No | ||
| 119 | PRKCB | 15061 | 0.317 | 0.4012 | No | ||
| 120 | MYL5 | 15130 | 0.309 | 0.4007 | No | ||
| 121 | ITGB5 | 15192 | 0.303 | 0.4004 | No | ||
| 122 | PDPK1 | 15482 | 0.277 | 0.3937 | No | ||
| 123 | CRKL | 15584 | 0.266 | 0.3921 | No | ||
| 124 | MAP2K1 | 16237 | 0.199 | 0.3753 | No | ||
| 125 | SHC2 | 16286 | 0.193 | 0.3748 | No | ||
| 126 | MYL12B | 16368 | 0.189 | 0.3734 | No | ||
| 127 | IGF1R | 17207 | 0.095 | 0.3510 | No | ||
| 128 | COL11A1 | 17329 | 0.081 | 0.3481 | No | ||
| 129 | ITGA6 | 17332 | 0.081 | 0.3484 | No | ||
| 130 | CTNNB1 | 18080 | -0.004 | 0.3280 | No | ||
| 131 | CCND2 | 18346 | -0.028 | 0.3209 | No | ||
| 132 | MAPK1 | 18358 | -0.029 | 0.3208 | No | ||
| 133 | LAMA1 | 18555 | -0.051 | 0.3157 | No | ||
| 134 | ITGAV | 18713 | -0.069 | 0.3117 | No | ||
| 135 | PAK1 | 18748 | -0.073 | 0.3111 | No | ||
| 136 | LAMA5 | 18877 | -0.085 | 0.3080 | No | ||
| 137 | PIK3R5 | 18946 | -0.090 | 0.3065 | No | ||
| 138 | RELN | 18953 | -0.090 | 0.3067 | No | ||
| 139 | RAP1A | 19126 | -0.107 | 0.3025 | No | ||
| 140 | COL6A6 | 19212 | -0.116 | 0.3007 | No | ||
| 141 | COL4A6 | 19436 | -0.138 | 0.2952 | No | ||
| 142 | CCND1 | 19463 | -0.141 | 0.2952 | No | ||
| 143 | DOCK1 | 19647 | -0.162 | 0.2909 | No | ||
| 144 | CRK | 19873 | -0.186 | 0.2856 | No | ||
| 145 | PAK2 | 19880 | -0.186 | 0.2862 | No | ||
| 146 | ARHGAP35 | 19964 | -0.191 | 0.2848 | No | ||
| 147 | MAPK9 | 20044 | -0.199 | 0.2835 | No | ||
| 148 | ACTN2 | 20441 | -0.238 | 0.2738 | No | ||
| 149 | MYLK | 20442 | -0.238 | 0.2748 | No | ||
| 150 | VAV1 | 20472 | -0.240 | 0.2751 | No | ||
| 151 | CHAD | 20501 | -0.243 | 0.2754 | No | ||
| 152 | COL4A4 | 20885 | -0.282 | 0.2662 | No | ||
| 153 | RAP1B | 21154 | -0.306 | 0.2602 | No | ||
| 154 | COL2A1 | 21231 | -0.315 | 0.2595 | No | ||
| 155 | PTEN | 21651 | -0.360 | 0.2497 | No | ||
| 156 | SHC4 | 21764 | -0.373 | 0.2483 | No | ||
| 157 | PPP1CC | 22752 | -0.460 | 0.2234 | No | ||
| 158 | PAK3 | 22753 | -0.460 | 0.2254 | No | ||
| 159 | ITGA7 | 23074 | -0.492 | 0.2188 | No | ||
| 160 | TNR | 23510 | -0.530 | 0.2093 | No | ||
| 161 | PTK2 | 23590 | -0.540 | 0.2095 | No | ||
| 162 | VAV3 | 24073 | -0.592 | 0.1990 | No | ||
| 163 | GSK3B | 24264 | -0.615 | 0.1965 | No | ||
| 164 | PIK3CB | 24695 | -0.651 | 0.1877 | No | ||
| 165 | COL5A2 | 24734 | -0.655 | 0.1895 | No | ||
| 166 | BIRC2 | 24892 | -0.671 | 0.1881 | No | ||
| 167 | PPP1CB | 25785 | -0.771 | 0.1672 | No | ||
| 168 | SOS2 | 26045 | -0.800 | 0.1637 | No | ||
| 169 | PDGFD | 26089 | -0.806 | 0.1660 | No | ||
| 170 | LAMB1 | 26097 | -0.806 | 0.1694 | No | ||
| 171 | PIK3R1 | 26229 | -0.821 | 0.1694 | No | ||
| 172 | LAMB4 | 26273 | -0.826 | 0.1718 | No | ||
| 173 | MYL2 | 27108 | -0.838 | 0.1528 | No | ||
| 174 | PIK3CA | 27643 | -0.860 | 0.1420 | No | ||
| 175 | XIAP | 27722 | -0.868 | 0.1437 | No | ||
| 176 | ROCK2 | 27828 | -0.882 | 0.1447 | No | ||
| 177 | LAMA2 | 28079 | -0.914 | 0.1419 | No | ||
| 178 | ITGB6 | 28305 | -0.947 | 0.1399 | No | ||
| 179 | TNXB | 29385 | -0.973 | 0.1147 | No | ||
| 180 | AKT3 | 29865 | -1.035 | 0.1062 | No | ||
| 181 | PPP1R12A | 29888 | -1.038 | 0.1102 | No | ||
| 182 | MET | 30219 | -1.073 | 0.1059 | No | ||
| 183 | BCL2 | 30238 | -1.076 | 0.1101 | No | ||
| 184 | THBS4 | 30275 | -1.084 | 0.1138 | No | ||
| 185 | MAPK10 | 30362 | -1.099 | 0.1163 | No | ||
| 186 | MAPK8 | 30659 | -1.138 | 0.1132 | No | ||
| 187 | PAK6 | 30817 | -1.168 | 0.1141 | No | ||
| 188 | ROCK1 | 30820 | -1.169 | 0.1191 | No | ||
| 189 | PRKCG | 30838 | -1.173 | 0.1238 | No | ||
| 190 | BRAF | 30894 | -1.182 | 0.1274 | No | ||
| 191 | SOS1 | 31146 | -1.221 | 0.1260 | No | ||
| 192 | ITGA2B | 31462 | -1.275 | 0.1229 | No | ||
| 193 | EGF | 31967 | -1.388 | 0.1153 | No | ||
| 194 | ACTN3 | 32138 | -1.426 | 0.1169 | No | ||
| 195 | VEGFA | 32146 | -1.428 | 0.1230 | No | ||
| 196 | ARHGAP5 | 32953 | -1.615 | 0.1081 | No |