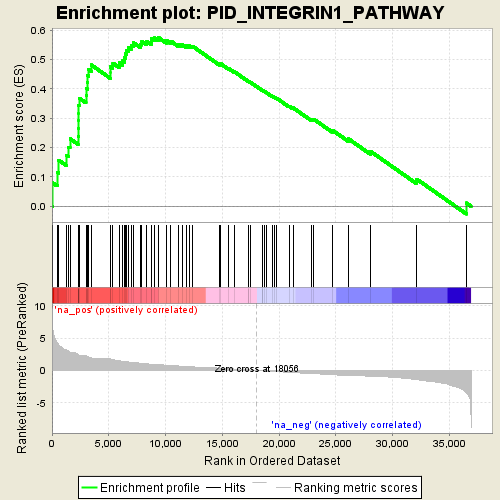
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | NOV0135_D11_log2foldchange_preranked |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_pos |
| GeneSet | PID_INTEGRIN1_PATHWAY |
| Enrichment Score (ES) | 0.5747513 |
| Normalized Enrichment Score (NES) | 2.1380823 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.0011010364 |
| FWER p-Value | 0.007 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | ITGA11 | 23 | 7.335 | 0.0803 | Yes | ||
| 2 | COL3A1 | 444 | 4.306 | 0.1164 | Yes | ||
| 3 | TGM2 | 596 | 3.996 | 0.1564 | Yes | ||
| 4 | FN1 | 1255 | 3.216 | 0.1740 | Yes | ||
| 5 | COL5A1 | 1472 | 3.013 | 0.2014 | Yes | ||
| 6 | ITGA10 | 1582 | 2.841 | 0.2298 | Yes | ||
| 7 | TGFBI | 2290 | 2.481 | 0.2380 | Yes | ||
| 8 | COL1A2 | 2328 | 2.443 | 0.2639 | Yes | ||
| 9 | COL6A3 | 2337 | 2.421 | 0.2904 | Yes | ||
| 10 | SPP1 | 2343 | 2.416 | 0.3170 | Yes | ||
| 11 | PLAU | 2348 | 2.409 | 0.3434 | Yes | ||
| 12 | COL6A2 | 2413 | 2.363 | 0.3678 | Yes | ||
| 13 | F13A1 | 2991 | 2.252 | 0.3769 | Yes | ||
| 14 | VTN | 3008 | 2.239 | 0.4012 | Yes | ||
| 15 | ITGA4 | 3094 | 2.181 | 0.4230 | Yes | ||
| 16 | COL1A1 | 3148 | 2.121 | 0.4449 | Yes | ||
| 17 | COL18A1 | 3225 | 2.080 | 0.4658 | Yes | ||
| 18 | ITGA3 | 3450 | 1.946 | 0.4812 | Yes | ||
| 19 | ITGA1 | 5116 | 1.782 | 0.4557 | Yes | ||
| 20 | COL4A1 | 5119 | 1.780 | 0.4753 | Yes | ||
| 21 | ITGA5 | 5362 | 1.671 | 0.4872 | Yes | ||
| 22 | LAMC2 | 5931 | 1.498 | 0.4883 | Yes | ||
| 23 | COL6A1 | 6238 | 1.407 | 0.4955 | Yes | ||
| 24 | LAMB3 | 6408 | 1.364 | 0.5060 | Yes | ||
| 25 | IGSF8 | 6483 | 1.348 | 0.5188 | Yes | ||
| 26 | CD81 | 6569 | 1.325 | 0.5311 | Yes | ||
| 27 | THBS2 | 6744 | 1.310 | 0.5409 | Yes | ||
| 28 | ITGA9 | 6962 | 1.264 | 0.5489 | Yes | ||
| 29 | COL11A2 | 7169 | 1.218 | 0.5568 | Yes | ||
| 30 | CSPG4 | 7786 | 1.124 | 0.5525 | Yes | ||
| 31 | TNC | 7915 | 1.104 | 0.5612 | Yes | ||
| 32 | COL4A3 | 8325 | 1.035 | 0.5615 | Yes | ||
| 33 | LAMA4 | 8753 | 0.966 | 0.5606 | Yes | ||
| 34 | NID1 | 8767 | 0.964 | 0.5709 | Yes | ||
| 35 | LAMC1 | 9059 | 0.924 | 0.5732 | Yes | ||
| 36 | THBS1 | 9361 | 0.885 | 0.5748 | Yes | ||
| 37 | MDK | 10064 | 0.813 | 0.5647 | No | ||
| 38 | FBN1 | 10452 | 0.770 | 0.5627 | No | ||
| 39 | ITGA2 | 11125 | 0.697 | 0.5521 | No | ||
| 40 | LAMA3 | 11453 | 0.662 | 0.5506 | No | ||
| 41 | LAMB2 | 11830 | 0.614 | 0.5471 | No | ||
| 42 | JAM2 | 12104 | 0.591 | 0.5462 | No | ||
| 43 | ITGB1 | 12370 | 0.555 | 0.5452 | No | ||
| 44 | PLAUR | 14791 | 0.347 | 0.4833 | No | ||
| 45 | ITGA8 | 14808 | 0.345 | 0.4867 | No | ||
| 46 | FGB | 15581 | 0.266 | 0.4687 | No | ||
| 47 | FGG | 16048 | 0.221 | 0.4585 | No | ||
| 48 | COL11A1 | 17329 | 0.081 | 0.4247 | No | ||
| 49 | ITGA6 | 17332 | 0.081 | 0.4255 | No | ||
| 50 | VCAM1 | 17475 | 0.066 | 0.4224 | No | ||
| 51 | FGA | 18548 | -0.050 | 0.3938 | No | ||
| 52 | LAMA1 | 18555 | -0.051 | 0.3942 | No | ||
| 53 | ITGAV | 18713 | -0.069 | 0.3907 | No | ||
| 54 | LAMA5 | 18877 | -0.085 | 0.3873 | No | ||
| 55 | COL4A6 | 19436 | -0.138 | 0.3736 | No | ||
| 56 | CD14 | 19614 | -0.159 | 0.3706 | No | ||
| 57 | COL7A1 | 19797 | -0.178 | 0.3676 | No | ||
| 58 | COL4A4 | 20885 | -0.282 | 0.3412 | No | ||
| 59 | COL2A1 | 21231 | -0.315 | 0.3353 | No | ||
| 60 | COL4A5 | 22847 | -0.469 | 0.2967 | No | ||
| 61 | ITGA7 | 23074 | -0.492 | 0.2960 | No | ||
| 62 | COL5A2 | 24734 | -0.655 | 0.2582 | No | ||
| 63 | LAMB1 | 26097 | -0.806 | 0.2301 | No | ||
| 64 | LAMA2 | 28079 | -0.914 | 0.1865 | No | ||
| 65 | VEGFA | 32146 | -1.428 | 0.0919 | No | ||
| 66 | NPNT | 36513 | -3.410 | 0.0111 | No |
