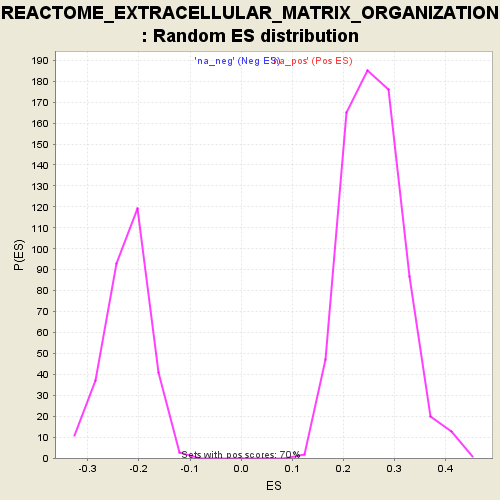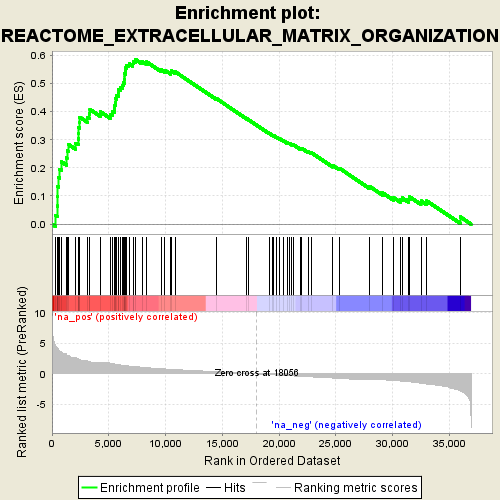
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | NOV0135_D11_log2foldchange_preranked |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_pos |
| GeneSet | REACTOME_EXTRACELLULAR_MATRIX_ORGANIZATION |
| Enrichment Score (ES) | 0.5853301 |
| Normalized Enrichment Score (NES) | 2.2496703 |
| Nominal p-value | 0.0 |
| FDR q-value | 4.2238442E-4 |
| FWER p-Value | 0.001 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | COL8A1 | 290 | 4.756 | 0.0323 | Yes | ||
| 2 | COL3A1 | 444 | 4.306 | 0.0645 | Yes | ||
| 3 | COL5A3 | 469 | 4.243 | 0.0996 | Yes | ||
| 4 | COL12A1 | 502 | 4.180 | 0.1340 | Yes | ||
| 5 | MMP7 | 591 | 4.023 | 0.1656 | Yes | ||
| 6 | MMP1 | 668 | 3.826 | 0.1958 | Yes | ||
| 7 | MMP9 | 834 | 3.549 | 0.2213 | Yes | ||
| 8 | MMP2 | 1281 | 3.155 | 0.2358 | Yes | ||
| 9 | CTRB1 | 1319 | 3.085 | 0.2609 | Yes | ||
| 10 | COL5A1 | 1472 | 3.013 | 0.2822 | Yes | ||
| 11 | TIMP1 | 2086 | 2.687 | 0.2882 | Yes | ||
| 12 | COL21A1 | 2309 | 2.471 | 0.3031 | Yes | ||
| 13 | COL1A2 | 2328 | 2.443 | 0.3232 | Yes | ||
| 14 | COL6A3 | 2337 | 2.421 | 0.3434 | Yes | ||
| 15 | COL6A2 | 2413 | 2.363 | 0.3613 | Yes | ||
| 16 | COL15A1 | 2422 | 2.359 | 0.3810 | Yes | ||
| 17 | COL1A1 | 3148 | 2.121 | 0.3792 | Yes | ||
| 18 | BMP1 | 3281 | 2.040 | 0.3929 | Yes | ||
| 19 | PPIB | 3336 | 2.021 | 0.4085 | Yes | ||
| 20 | MMP8 | 4243 | 1.927 | 0.4001 | Yes | ||
| 21 | COL4A1 | 5119 | 1.780 | 0.3914 | Yes | ||
| 22 | P4HB | 5339 | 1.687 | 0.3997 | Yes | ||
| 23 | MMP24 | 5468 | 1.634 | 0.4100 | Yes | ||
| 24 | MMP14 | 5536 | 1.622 | 0.4219 | Yes | ||
| 25 | COL4A2 | 5574 | 1.601 | 0.4344 | Yes | ||
| 26 | MMP25 | 5602 | 1.596 | 0.4471 | Yes | ||
| 27 | TIMP2 | 5708 | 1.553 | 0.4574 | Yes | ||
| 28 | PLOD1 | 5842 | 1.525 | 0.4667 | Yes | ||
| 29 | COL8A2 | 5853 | 1.522 | 0.4792 | Yes | ||
| 30 | CTRB2 | 6038 | 1.459 | 0.4866 | Yes | ||
| 31 | COL6A1 | 6238 | 1.407 | 0.4930 | Yes | ||
| 32 | PCOLCE | 6263 | 1.398 | 0.5042 | Yes | ||
| 33 | COL10A1 | 6360 | 1.372 | 0.5132 | Yes | ||
| 34 | ADAMTS14 | 6371 | 1.371 | 0.5245 | Yes | ||
| 35 | PLOD3 | 6373 | 1.370 | 0.5360 | Yes | ||
| 36 | MMP10 | 6462 | 1.356 | 0.5450 | Yes | ||
| 37 | KLKB1 | 6490 | 1.347 | 0.5557 | Yes | ||
| 38 | SERPINH1 | 6552 | 1.329 | 0.5652 | Yes | ||
| 39 | COL23A1 | 6778 | 1.303 | 0.5701 | Yes | ||
| 40 | COL13A1 | 7131 | 1.227 | 0.5709 | Yes | ||
| 41 | COL11A2 | 7169 | 1.218 | 0.5802 | Yes | ||
| 42 | TLL2 | 7355 | 1.201 | 0.5853 | Yes | ||
| 43 | FURIN | 7929 | 1.101 | 0.5791 | No | ||
| 44 | COL4A3 | 8325 | 1.035 | 0.5771 | No | ||
| 45 | COL16A1 | 9608 | 0.855 | 0.5495 | No | ||
| 46 | MMP11 | 9925 | 0.829 | 0.5479 | No | ||
| 47 | MMP17 | 10464 | 0.769 | 0.5398 | No | ||
| 48 | MMP15 | 10497 | 0.765 | 0.5454 | No | ||
| 49 | CRTAP | 10843 | 0.734 | 0.5422 | No | ||
| 50 | COL25A1 | 14525 | 0.378 | 0.4455 | No | ||
| 51 | COL27A1 | 17169 | 0.099 | 0.3746 | No | ||
| 52 | COL11A1 | 17329 | 0.081 | 0.3710 | No | ||
| 53 | COL24A1 | 19120 | -0.106 | 0.3233 | No | ||
| 54 | COL4A6 | 19436 | -0.138 | 0.3159 | No | ||
| 55 | COL9A3 | 19536 | -0.150 | 0.3145 | No | ||
| 56 | COL7A1 | 19797 | -0.178 | 0.3089 | No | ||
| 57 | PLOD2 | 20084 | -0.202 | 0.3029 | No | ||
| 58 | COL17A1 | 20406 | -0.234 | 0.2961 | No | ||
| 59 | TLL1 | 20732 | -0.266 | 0.2896 | No | ||
| 60 | COL4A4 | 20885 | -0.282 | 0.2878 | No | ||
| 61 | ADAMTS2 | 21126 | -0.302 | 0.2839 | No | ||
| 62 | COL2A1 | 21231 | -0.315 | 0.2837 | No | ||
| 63 | CMA1 | 21862 | -0.383 | 0.2698 | No | ||
| 64 | ADAMTS3 | 22022 | -0.400 | 0.2689 | No | ||
| 65 | COL9A1 | 22587 | -0.443 | 0.2573 | No | ||
| 66 | COL4A5 | 22847 | -0.469 | 0.2542 | No | ||
| 67 | COL5A2 | 24734 | -0.655 | 0.2086 | No | ||
| 68 | COL14A1 | 25319 | -0.719 | 0.1988 | No | ||
| 69 | COL9A2 | 27990 | -0.903 | 0.1339 | No | ||
| 70 | ELANE | 29137 | -0.952 | 0.1109 | No | ||
| 71 | MMP3 | 30126 | -1.066 | 0.0931 | No | ||
| 72 | COL28A1 | 30689 | -1.145 | 0.0875 | No | ||
| 73 | COL22A1 | 30847 | -1.174 | 0.0931 | No | ||
| 74 | COL19A1 | 31443 | -1.271 | 0.0877 | No | ||
| 75 | PCOLCE2 | 31504 | -1.284 | 0.0969 | No | ||
| 76 | PRSS1 | 32513 | -1.512 | 0.0823 | No | ||
| 77 | MMP16 | 33024 | -1.637 | 0.0823 | No | ||
| 78 | PLG | 35971 | -2.782 | 0.0258 | No |