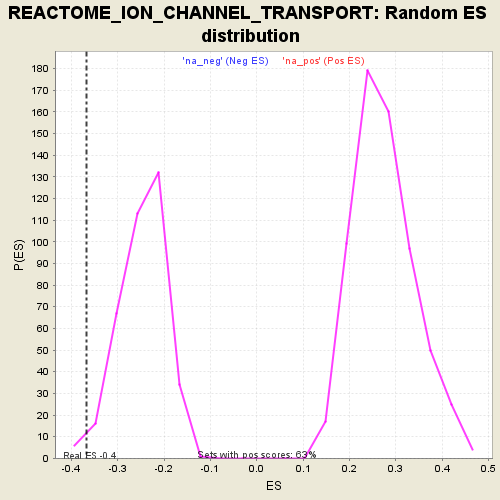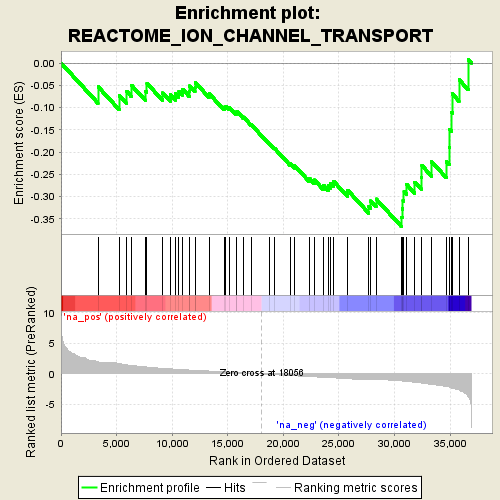
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | NOV0135_D11_log2foldchange_preranked |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_neg |
| GeneSet | REACTOME_ION_CHANNEL_TRANSPORT |
| Enrichment Score (ES) | -0.3664917 |
| Normalized Enrichment Score (NES) | -1.4931577 |
| Nominal p-value | 0.016260162 |
| FDR q-value | 1.0 |
| FWER p-Value | 1.0 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | HTR3B | 3324 | 2.022 | -0.0530 | No | ||
| 2 | ATP8B3 | 5250 | 1.719 | -0.0736 | No | ||
| 3 | GABRA1 | 5917 | 1.504 | -0.0640 | No | ||
| 4 | ATP2B3 | 6334 | 1.379 | -0.0499 | No | ||
| 5 | ATP10A | 7604 | 1.158 | -0.0631 | No | ||
| 6 | GABRA3 | 7726 | 1.134 | -0.0455 | No | ||
| 7 | ATP4B | 9141 | 0.912 | -0.0671 | No | ||
| 8 | ATP4A | 9845 | 0.839 | -0.0707 | No | ||
| 9 | HTR3A | 10303 | 0.784 | -0.0687 | No | ||
| 10 | ATP10D | 10603 | 0.753 | -0.0630 | No | ||
| 11 | ATP7B | 10932 | 0.723 | -0.0586 | No | ||
| 12 | GABRA5 | 11537 | 0.651 | -0.0630 | No | ||
| 13 | ATP1A1 | 11550 | 0.650 | -0.0514 | No | ||
| 14 | GLRA1 | 12051 | 0.594 | -0.0540 | No | ||
| 15 | ATP1B3 | 12086 | 0.593 | -0.0441 | No | ||
| 16 | ATP1B1 | 13313 | 0.479 | -0.0685 | No | ||
| 17 | ARHGEF9 | 14669 | 0.362 | -0.0986 | No | ||
| 18 | ATP2A3 | 14834 | 0.341 | -0.0968 | No | ||
| 19 | ATP2A2 | 15118 | 0.310 | -0.0987 | No | ||
| 20 | ATP9A | 15746 | 0.251 | -0.1111 | No | ||
| 21 | GABRB3 | 15821 | 0.245 | -0.1086 | No | ||
| 22 | ATP8A2 | 16405 | 0.185 | -0.1210 | No | ||
| 23 | ATP2A1 | 17125 | 0.104 | -0.1386 | No | ||
| 24 | ATP1A2 | 18788 | -0.077 | -0.1823 | No | ||
| 25 | ATP9B | 19181 | -0.112 | -0.1909 | No | ||
| 26 | ATP8B1 | 20616 | -0.253 | -0.2251 | No | ||
| 27 | ATP1A3 | 21010 | -0.293 | -0.2304 | No | ||
| 28 | GABRR2 | 22338 | -0.418 | -0.2587 | No | ||
| 29 | ATP8B4 | 22792 | -0.464 | -0.2624 | No | ||
| 30 | ATP2C1 | 23618 | -0.543 | -0.2748 | No | ||
| 31 | ATP2B2 | 24037 | -0.588 | -0.2754 | No | ||
| 32 | ATP11A | 24250 | -0.614 | -0.2698 | No | ||
| 33 | ATP7A | 24514 | -0.633 | -0.2653 | No | ||
| 34 | ATP1B2 | 25795 | -0.772 | -0.2859 | No | ||
| 35 | ATP1A4 | 27693 | -0.865 | -0.3214 | No | ||
| 36 | ATP8A1 | 27859 | -0.885 | -0.3096 | No | ||
| 37 | ATP11C | 28341 | -0.952 | -0.3052 | No | ||
| 38 | GABRA6 | 30603 | -1.133 | -0.3457 | Yes | ||
| 39 | GLRA2 | 30686 | -1.143 | -0.3269 | Yes | ||
| 40 | ATP11B | 30759 | -1.156 | -0.3076 | Yes | ||
| 41 | GABRB1 | 30850 | -1.174 | -0.2884 | Yes | ||
| 42 | GLRB | 31086 | -1.210 | -0.2725 | Yes | ||
| 43 | FXYD2 | 31830 | -1.358 | -0.2677 | Yes | ||
| 44 | GABRG3 | 32397 | -1.479 | -0.2559 | Yes | ||
| 45 | GABRA2 | 32465 | -1.495 | -0.2302 | Yes | ||
| 46 | GLRA3 | 33343 | -1.756 | -0.2217 | Yes | ||
| 47 | GABRB2 | 34668 | -1.991 | -0.2210 | Yes | ||
| 48 | ATP10B | 34935 | -2.155 | -0.1886 | Yes | ||
| 49 | GABRG2 | 34979 | -2.192 | -0.1495 | Yes | ||
| 50 | ATP2C2 | 35141 | -2.336 | -0.1109 | Yes | ||
| 51 | GABRR1 | 35193 | -2.350 | -0.0691 | Yes | ||
| 52 | GABRA4 | 35804 | -2.626 | -0.0374 | Yes | ||
| 53 | ATP12A | 36635 | -3.682 | 0.0078 | Yes |