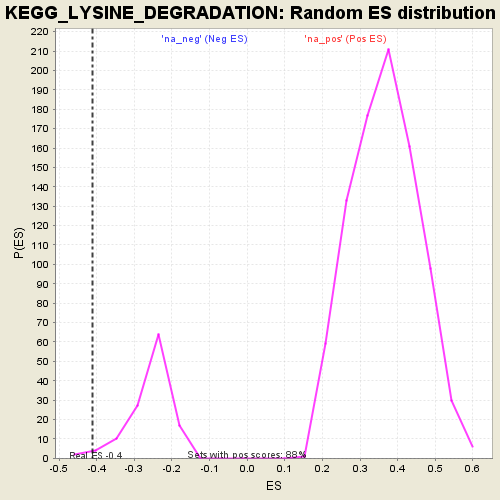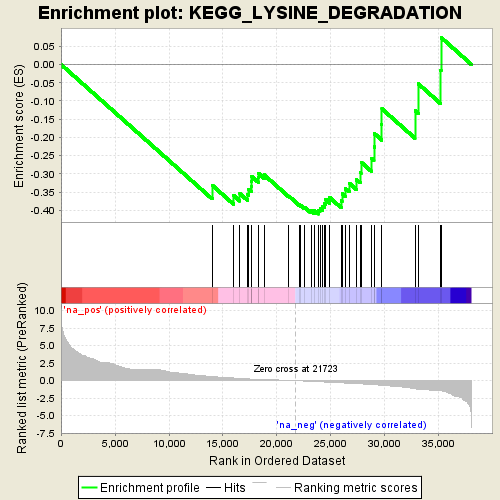
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | NOV0135_D3.1_log2foldchange_preranked |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_neg |
| GeneSet | KEGG_LYSINE_DEGRADATION |
| Enrichment Score (ES) | -0.40986627 |
| Normalized Enrichment Score (NES) | -1.595378 |
| Nominal p-value | 0.024193548 |
| FDR q-value | 0.5772841 |
| FWER p-Value | 0.888 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | SETD7 | 14002 | 0.582 | -0.3310 | No | ||
| 2 | ASH1L | 15971 | 0.376 | -0.3584 | No | ||
| 3 | SUV39H1 | 16573 | 0.315 | -0.3539 | No | ||
| 4 | DLST | 17265 | 0.255 | -0.3556 | No | ||
| 5 | ACAT2 | 17361 | 0.246 | -0.3422 | No | ||
| 6 | PLOD2 | 17614 | 0.225 | -0.3343 | No | ||
| 7 | SETD1A | 17623 | 0.224 | -0.3201 | No | ||
| 8 | PLOD1 | 17641 | 0.223 | -0.3061 | No | ||
| 9 | SETD2 | 18263 | 0.192 | -0.3101 | No | ||
| 10 | ALDH2 | 18279 | 0.190 | -0.2982 | No | ||
| 11 | PLOD3 | 18816 | 0.151 | -0.3025 | No | ||
| 12 | ACAT1 | 21095 | 0.031 | -0.3604 | No | ||
| 13 | OGDH | 22075 | -0.024 | -0.3846 | No | ||
| 14 | GCDH | 22229 | -0.036 | -0.3863 | No | ||
| 15 | ALDH9A1 | 22586 | -0.061 | -0.3918 | No | ||
| 16 | HADHA | 23167 | -0.102 | -0.4005 | No | ||
| 17 | ECHS1 | 23477 | -0.125 | -0.4005 | Yes | ||
| 18 | NSD1 | 23833 | -0.151 | -0.4001 | Yes | ||
| 19 | OGDHL | 23993 | -0.163 | -0.3938 | Yes | ||
| 20 | DOT1L | 24234 | -0.183 | -0.3883 | Yes | ||
| 21 | SETMAR | 24423 | -0.196 | -0.3806 | Yes | ||
| 22 | HADH | 24535 | -0.204 | -0.3703 | Yes | ||
| 23 | SUV39H2 | 24905 | -0.231 | -0.3651 | Yes | ||
| 24 | SETDB2 | 25964 | -0.313 | -0.3727 | Yes | ||
| 25 | TMLHE | 26036 | -0.317 | -0.3540 | Yes | ||
| 26 | ALDH7A1 | 26360 | -0.343 | -0.3404 | Yes | ||
| 27 | AASDHPPT | 26698 | -0.369 | -0.3254 | Yes | ||
| 28 | EHMT1 | 27344 | -0.422 | -0.3151 | Yes | ||
| 29 | AADAT | 27710 | -0.455 | -0.2952 | Yes | ||
| 30 | SETDB1 | 27797 | -0.462 | -0.2676 | Yes | ||
| 31 | EHMT2 | 28773 | -0.556 | -0.2574 | Yes | ||
| 32 | SETD1B | 28997 | -0.577 | -0.2259 | Yes | ||
| 33 | AASDH | 29027 | -0.582 | -0.1891 | Yes | ||
| 34 | ALDH3A2 | 29724 | -0.665 | -0.1644 | Yes | ||
| 35 | AASS | 29735 | -0.666 | -0.1216 | Yes | ||
| 36 | ALDH1B1 | 32797 | -1.170 | -0.1265 | Yes | ||
| 37 | PIPOX | 33089 | -1.263 | -0.0525 | Yes | ||
| 38 | EHHADH | 35145 | -1.402 | -0.0160 | Yes | ||
| 39 | BBOX1 | 35211 | -1.421 | 0.0742 | Yes |