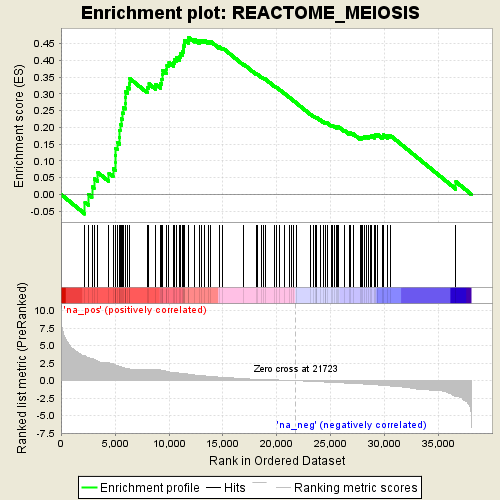
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | NOV0135_D3.1_log2foldchange_preranked |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_pos |
| GeneSet | REACTOME_MEIOSIS |
| Enrichment Score (ES) | 0.46846646 |
| Normalized Enrichment Score (NES) | 1.425397 |
| Nominal p-value | 0.0101936795 |
| FDR q-value | 1.0 |
| FWER p-Value | 1.0 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | HIST1H2BJ | 2214 | 3.527 | -0.0235 | Yes | ||
| 2 | HIST2H3D | 2565 | 3.240 | -0.0007 | Yes | ||
| 3 | HIST1H2BO | 2865 | 3.141 | 0.0224 | Yes | ||
| 4 | HIST1H2BG | 3055 | 3.054 | 0.0476 | Yes | ||
| 5 | PRDM9 | 3414 | 2.748 | 0.0653 | Yes | ||
| 6 | HIST1H2BH | 4437 | 2.532 | 0.0634 | Yes | ||
| 7 | HIST2H2BE | 4815 | 2.448 | 0.0776 | Yes | ||
| 8 | HIST1H2BB | 5026 | 2.275 | 0.0946 | Yes | ||
| 9 | HIST1H4H | 5032 | 2.273 | 0.1169 | Yes | ||
| 10 | HIST1H4L | 5077 | 2.234 | 0.1378 | Yes | ||
| 11 | HIST1H3H | 5208 | 2.159 | 0.1557 | Yes | ||
| 12 | HIST1H4B | 5425 | 2.072 | 0.1705 | Yes | ||
| 13 | HSPA2 | 5438 | 2.069 | 0.1906 | Yes | ||
| 14 | HIST1H3F | 5467 | 2.050 | 0.2102 | Yes | ||
| 15 | HIST1H4K | 5607 | 1.962 | 0.2259 | Yes | ||
| 16 | HIST1H3B | 5674 | 1.923 | 0.2431 | Yes | ||
| 17 | HIST1H2BK | 5754 | 1.875 | 0.2596 | Yes | ||
| 18 | HIST1H2AJ | 5930 | 1.794 | 0.2727 | Yes | ||
| 19 | HIST1H4E | 5945 | 1.787 | 0.2900 | Yes | ||
| 20 | HIST1H3C | 5975 | 1.764 | 0.3066 | Yes | ||
| 21 | HIST2H4A | 6170 | 1.706 | 0.3184 | Yes | ||
| 22 | HIST1H3I | 6313 | 1.667 | 0.3311 | Yes | ||
| 23 | HIST1H2BL | 6378 | 1.636 | 0.3456 | Yes | ||
| 24 | HIST1H2AE | 7989 | 1.594 | 0.3189 | Yes | ||
| 25 | HIST1H2BC | 8145 | 1.536 | 0.3300 | Yes | ||
| 26 | H3F3AP5 | 8762 | 1.525 | 0.3288 | Yes | ||
| 27 | SYNE1 | 9238 | 1.508 | 0.3312 | Yes | ||
| 28 | HIST1H3A | 9308 | 1.486 | 0.3441 | Yes | ||
| 29 | HIST1H2BF | 9362 | 1.467 | 0.3572 | Yes | ||
| 30 | HIST1H2BN | 9386 | 1.455 | 0.3709 | Yes | ||
| 31 | HIST1H2BD | 9801 | 1.321 | 0.3731 | Yes | ||
| 32 | HIST1H4I | 9809 | 1.319 | 0.3859 | Yes | ||
| 33 | HIST1H3J | 9997 | 1.254 | 0.3934 | Yes | ||
| 34 | HIST1H3G | 10451 | 1.154 | 0.3928 | Yes | ||
| 35 | HIST3H3 | 10516 | 1.145 | 0.4025 | Yes | ||
| 36 | SYCP1 | 10703 | 1.129 | 0.4087 | Yes | ||
| 37 | TERF2IP | 11013 | 1.063 | 0.4111 | Yes | ||
| 38 | HIST1H2BI | 11077 | 1.061 | 0.4199 | Yes | ||
| 39 | HIST4H4 | 11252 | 1.042 | 0.4256 | Yes | ||
| 40 | HIST1H2AC | 11324 | 1.021 | 0.4338 | Yes | ||
| 41 | LMNA | 11352 | 1.016 | 0.4431 | Yes | ||
| 42 | HIST1H2BE | 11402 | 1.005 | 0.4518 | Yes | ||
| 43 | HIST1H4D | 11463 | 0.994 | 0.4600 | Yes | ||
| 44 | H2AFZ | 11824 | 0.925 | 0.4597 | Yes | ||
| 45 | HIST1H4C | 11837 | 0.923 | 0.4685 | Yes | ||
| 46 | HIST1H4J | 12355 | 0.819 | 0.4629 | No | ||
| 47 | HIST1H3E | 12795 | 0.747 | 0.4587 | No | ||
| 48 | RAD51C | 13048 | 0.713 | 0.4591 | No | ||
| 49 | SYCP3 | 13269 | 0.678 | 0.4600 | No | ||
| 50 | H2AFX | 13632 | 0.628 | 0.4567 | No | ||
| 51 | TEX15 | 13824 | 0.603 | 0.4576 | No | ||
| 52 | HIST3H2BB | 14652 | 0.507 | 0.4408 | No | ||
| 53 | SMC1A | 14989 | 0.472 | 0.4366 | No | ||
| 54 | HIST2H2AC | 16919 | 0.286 | 0.3886 | No | ||
| 55 | H3F3A | 18071 | 0.204 | 0.3603 | No | ||
| 56 | H3F3AP6 | 18211 | 0.194 | 0.3585 | No | ||
| 57 | SMC3 | 18581 | 0.167 | 0.3504 | No | ||
| 58 | HIST1H2AB | 18760 | 0.155 | 0.3473 | No | ||
| 59 | RAD21 | 18910 | 0.144 | 0.3448 | No | ||
| 60 | HIST1H4A | 19810 | 0.110 | 0.3221 | No | ||
| 61 | H3F3B | 19965 | 0.105 | 0.3191 | No | ||
| 62 | SYNE2 | 20233 | 0.086 | 0.3129 | No | ||
| 63 | DIDO1 | 20733 | 0.054 | 0.3003 | No | ||
| 64 | TOP3A | 21157 | 0.026 | 0.2894 | No | ||
| 65 | TERF2 | 21313 | 0.017 | 0.2855 | No | ||
| 66 | TERF1 | 21559 | 0.012 | 0.2791 | No | ||
| 67 | UBE2I | 21801 | -0.005 | 0.2728 | No | ||
| 68 | STAG2 | 23141 | -0.100 | 0.2385 | No | ||
| 69 | MLH1 | 23416 | -0.121 | 0.2325 | No | ||
| 70 | CDK4 | 23557 | -0.131 | 0.2301 | No | ||
| 71 | ACD | 23643 | -0.137 | 0.2292 | No | ||
| 72 | TINF2 | 23698 | -0.141 | 0.2292 | No | ||
| 73 | POT1 | 24000 | -0.163 | 0.2229 | No | ||
| 74 | TEX12 | 24276 | -0.186 | 0.2174 | No | ||
| 75 | STAG1 | 24542 | -0.204 | 0.2125 | No | ||
| 76 | BRCA2 | 24543 | -0.204 | 0.2145 | No | ||
| 77 | HIST1H2BM | 24664 | -0.214 | 0.2134 | No | ||
| 78 | RAD51 | 25066 | -0.245 | 0.2053 | No | ||
| 79 | MSH5 | 25117 | -0.249 | 0.2064 | No | ||
| 80 | SYCP2 | 25318 | -0.265 | 0.2038 | No | ||
| 81 | RPA1 | 25537 | -0.281 | 0.2008 | No | ||
| 82 | REC8 | 25636 | -0.287 | 0.2010 | No | ||
| 83 | ATM | 25660 | -0.289 | 0.2033 | No | ||
| 84 | RPA3 | 26256 | -0.333 | 0.1909 | No | ||
| 85 | STAG3 | 26699 | -0.369 | 0.1829 | No | ||
| 86 | SMC1B | 26812 | -0.379 | 0.1837 | No | ||
| 87 | ATR | 27063 | -0.397 | 0.1810 | No | ||
| 88 | HIST1H3D | 27756 | -0.459 | 0.1673 | No | ||
| 89 | MLH3 | 27832 | -0.466 | 0.1699 | No | ||
| 90 | RPA2 | 27974 | -0.479 | 0.1709 | No | ||
| 91 | BLM | 28071 | -0.488 | 0.1732 | No | ||
| 92 | CDK2 | 28256 | -0.504 | 0.1734 | No | ||
| 93 | DMC1 | 28488 | -0.526 | 0.1725 | No | ||
| 94 | LMNB1 | 28635 | -0.541 | 0.1740 | No | ||
| 95 | RAD50 | 28779 | -0.556 | 0.1757 | No | ||
| 96 | NBN | 29059 | -0.583 | 0.1741 | No | ||
| 97 | MSH4 | 29090 | -0.587 | 0.1791 | No | ||
| 98 | SUN2 | 29359 | -0.617 | 0.1781 | No | ||
| 99 | RBBP8 | 29760 | -0.669 | 0.1742 | No | ||
| 100 | BRCA1 | 29881 | -0.685 | 0.1778 | No | ||
| 101 | FKBP6 | 30253 | -0.732 | 0.1753 | No | ||
| 102 | MND1 | 30495 | -0.764 | 0.1764 | No | ||
| 103 | HIST1H4F | 36591 | -2.244 | 0.0379 | No |
