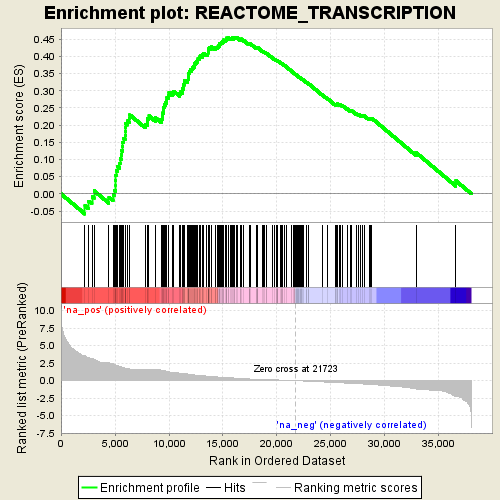
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | NOV0135_D3.1_log2foldchange_preranked |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_pos |
| GeneSet | REACTOME_TRANSCRIPTION |
| Enrichment Score (ES) | 0.4551508 |
| Normalized Enrichment Score (NES) | 1.4192697 |
| Nominal p-value | 0.0 |
| FDR q-value | 1.0 |
| FWER p-Value | 1.0 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | HIST1H2BJ | 2214 | 3.527 | -0.0341 | Yes | ||
| 2 | HIST2H3D | 2565 | 3.240 | -0.0209 | Yes | ||
| 3 | HIST1H2BO | 2865 | 3.141 | -0.0070 | Yes | ||
| 4 | HIST1H2BG | 3055 | 3.054 | 0.0092 | Yes | ||
| 5 | HIST1H2BH | 4437 | 2.532 | -0.0098 | Yes | ||
| 6 | HIST2H2BE | 4815 | 2.448 | -0.0028 | Yes | ||
| 7 | GTF2F1 | 4914 | 2.369 | 0.0110 | Yes | ||
| 8 | HIST1H2BB | 5026 | 2.275 | 0.0239 | Yes | ||
| 9 | HIST1H4H | 5032 | 2.273 | 0.0395 | Yes | ||
| 10 | HIST1H4L | 5077 | 2.234 | 0.0538 | Yes | ||
| 11 | TAF4B | 5100 | 2.226 | 0.0686 | Yes | ||
| 12 | HIST1H3H | 5208 | 2.159 | 0.0808 | Yes | ||
| 13 | HIST1H4B | 5425 | 2.072 | 0.0894 | Yes | ||
| 14 | HIST1H3F | 5467 | 2.050 | 0.1025 | Yes | ||
| 15 | POLR3K | 5561 | 1.996 | 0.1139 | Yes | ||
| 16 | HIST1H4K | 5607 | 1.962 | 0.1263 | Yes | ||
| 17 | HIST1H3B | 5674 | 1.923 | 0.1379 | Yes | ||
| 18 | TAF13 | 5705 | 1.899 | 0.1503 | Yes | ||
| 19 | HIST1H2BK | 5754 | 1.875 | 0.1620 | Yes | ||
| 20 | HIST1H2AJ | 5930 | 1.794 | 0.1698 | Yes | ||
| 21 | HIST1H4E | 5945 | 1.787 | 0.1818 | Yes | ||
| 22 | SNAPC2 | 5960 | 1.775 | 0.1937 | Yes | ||
| 23 | HIST1H3C | 5975 | 1.764 | 0.2056 | Yes | ||
| 24 | HIST2H4A | 6170 | 1.706 | 0.2123 | Yes | ||
| 25 | HIST1H3I | 6313 | 1.667 | 0.2201 | Yes | ||
| 26 | HIST1H2BL | 6378 | 1.636 | 0.2297 | Yes | ||
| 27 | U2AF1 | 7810 | 1.632 | 0.2032 | Yes | ||
| 28 | HIST1H2AE | 7989 | 1.594 | 0.2095 | Yes | ||
| 29 | GTF2A2 | 8029 | 1.577 | 0.2194 | Yes | ||
| 30 | HIST1H2BC | 8145 | 1.536 | 0.2271 | Yes | ||
| 31 | H3F3AP5 | 8762 | 1.525 | 0.2213 | Yes | ||
| 32 | HIST1H3A | 9308 | 1.486 | 0.2172 | Yes | ||
| 33 | HIST1H2BF | 9362 | 1.467 | 0.2260 | Yes | ||
| 34 | HIST1H2BN | 9386 | 1.455 | 0.2355 | Yes | ||
| 35 | PABPN1 | 9463 | 1.434 | 0.2434 | Yes | ||
| 36 | GTF2B | 9484 | 1.425 | 0.2528 | Yes | ||
| 37 | POLR2A | 9581 | 1.397 | 0.2599 | Yes | ||
| 38 | RRN3 | 9699 | 1.356 | 0.2662 | Yes | ||
| 39 | HIST1H2BD | 9801 | 1.321 | 0.2727 | Yes | ||
| 40 | HIST1H4I | 9809 | 1.319 | 0.2816 | Yes | ||
| 41 | SNAPC3 | 9982 | 1.260 | 0.2858 | Yes | ||
| 42 | HIST1H3J | 9997 | 1.254 | 0.2941 | Yes | ||
| 43 | TFB2M | 10316 | 1.190 | 0.2940 | Yes | ||
| 44 | HIST1H3G | 10451 | 1.154 | 0.2984 | Yes | ||
| 45 | CCNT1 | 10998 | 1.068 | 0.2914 | Yes | ||
| 46 | HIST1H2BI | 11077 | 1.061 | 0.2967 | Yes | ||
| 47 | CSTF2 | 11232 | 1.044 | 0.2999 | Yes | ||
| 48 | HIST4H4 | 11252 | 1.042 | 0.3066 | Yes | ||
| 49 | HIST1H2AC | 11324 | 1.021 | 0.3118 | Yes | ||
| 50 | ZNF143 | 11362 | 1.014 | 0.3178 | Yes | ||
| 51 | HIST1H2BE | 11402 | 1.005 | 0.3238 | Yes | ||
| 52 | HIST1H4D | 11463 | 0.994 | 0.3291 | Yes | ||
| 53 | LZTS1 | 11693 | 0.950 | 0.3296 | Yes | ||
| 54 | CLP1 | 11760 | 0.937 | 0.3343 | Yes | ||
| 55 | GTF2E2 | 11787 | 0.932 | 0.3401 | Yes | ||
| 56 | H2AFZ | 11824 | 0.925 | 0.3456 | Yes | ||
| 57 | HIST1H4C | 11837 | 0.923 | 0.3516 | Yes | ||
| 58 | TAF5 | 11924 | 0.903 | 0.3556 | Yes | ||
| 59 | POLR2F | 11946 | 0.898 | 0.3613 | Yes | ||
| 60 | ZNF473 | 12103 | 0.871 | 0.3632 | Yes | ||
| 61 | CCNT2 | 12147 | 0.861 | 0.3680 | Yes | ||
| 62 | TAF9 | 12302 | 0.829 | 0.3697 | Yes | ||
| 63 | HIST1H4J | 12355 | 0.819 | 0.3740 | Yes | ||
| 64 | CTDP1 | 12367 | 0.817 | 0.3794 | Yes | ||
| 65 | MAGOH | 12461 | 0.800 | 0.3825 | Yes | ||
| 66 | POLR2C | 12582 | 0.783 | 0.3847 | Yes | ||
| 67 | TAF1A | 12626 | 0.775 | 0.3890 | Yes | ||
| 68 | SNAPC4 | 12685 | 0.765 | 0.3927 | Yes | ||
| 69 | HIST1H3E | 12795 | 0.747 | 0.3950 | Yes | ||
| 70 | CDK9 | 12861 | 0.743 | 0.3984 | Yes | ||
| 71 | TCEA1 | 12882 | 0.739 | 0.4030 | Yes | ||
| 72 | POLR1C | 13120 | 0.704 | 0.4017 | Yes | ||
| 73 | CDC40 | 13135 | 0.700 | 0.4061 | Yes | ||
| 74 | SSB | 13218 | 0.685 | 0.4087 | Yes | ||
| 75 | DHX38 | 13497 | 0.650 | 0.4059 | Yes | ||
| 76 | SNAPC1 | 13614 | 0.631 | 0.4072 | Yes | ||
| 77 | RNGTT | 13618 | 0.631 | 0.4115 | Yes | ||
| 78 | H2AFX | 13632 | 0.628 | 0.4155 | Yes | ||
| 79 | POLR3C | 13639 | 0.627 | 0.4197 | Yes | ||
| 80 | U2AF2 | 13698 | 0.619 | 0.4224 | Yes | ||
| 81 | TFAM | 13755 | 0.611 | 0.4252 | Yes | ||
| 82 | GTF2A1 | 13899 | 0.595 | 0.4255 | Yes | ||
| 83 | UPF3B | 13961 | 0.586 | 0.4280 | Yes | ||
| 84 | NFIB | 14274 | 0.551 | 0.4235 | Yes | ||
| 85 | GTF2F2 | 14320 | 0.546 | 0.4261 | Yes | ||
| 86 | POLR2L | 14463 | 0.529 | 0.4260 | Yes | ||
| 87 | POLR2B | 14538 | 0.521 | 0.4277 | Yes | ||
| 88 | MBD2 | 14566 | 0.517 | 0.4306 | Yes | ||
| 89 | HIST3H2BB | 14652 | 0.507 | 0.4318 | Yes | ||
| 90 | SNRPB | 14653 | 0.507 | 0.4353 | Yes | ||
| 91 | POLR3E | 14705 | 0.502 | 0.4375 | Yes | ||
| 92 | POLR2K | 14800 | 0.493 | 0.4384 | Yes | ||
| 93 | POLR2G | 14867 | 0.487 | 0.4400 | Yes | ||
| 94 | GTF2H2 | 14975 | 0.474 | 0.4405 | Yes | ||
| 95 | POLR3D | 14987 | 0.472 | 0.4435 | Yes | ||
| 96 | TAF1B | 15058 | 0.465 | 0.4448 | Yes | ||
| 97 | POLR3F | 15065 | 0.464 | 0.4479 | Yes | ||
| 98 | POLR3A | 15211 | 0.453 | 0.4472 | Yes | ||
| 99 | TBP | 15294 | 0.445 | 0.4481 | Yes | ||
| 100 | POLR2E | 15298 | 0.445 | 0.4511 | Yes | ||
| 101 | SUPT5H | 15319 | 0.442 | 0.4537 | Yes | ||
| 102 | SRSF4 | 15513 | 0.423 | 0.4515 | Yes | ||
| 103 | SNRPD3 | 15527 | 0.422 | 0.4541 | Yes | ||
| 104 | SRSF11 | 15686 | 0.406 | 0.4527 | Yes | ||
| 105 | MNAT1 | 15764 | 0.397 | 0.4534 | Yes | ||
| 106 | GTF2H4 | 15841 | 0.390 | 0.4541 | Yes | ||
| 107 | GTF2H1 | 16012 | 0.372 | 0.4522 | Yes | ||
| 108 | RNPS1 | 16043 | 0.369 | 0.4540 | Yes | ||
| 109 | GTF2H2B | 16094 | 0.363 | 0.4552 | Yes | ||
| 110 | NCBP2 | 16219 | 0.349 | 0.4543 | No | ||
| 111 | TAF12 | 16323 | 0.339 | 0.4539 | No | ||
| 112 | PCF11 | 16609 | 0.310 | 0.4485 | No | ||
| 113 | CSTF3 | 16652 | 0.306 | 0.4495 | No | ||
| 114 | SRSF5 | 16685 | 0.303 | 0.4508 | No | ||
| 115 | HIST2H2AC | 16919 | 0.286 | 0.4466 | No | ||
| 116 | PAPOLA | 17415 | 0.242 | 0.4352 | No | ||
| 117 | GTF3C4 | 17468 | 0.237 | 0.4355 | No | ||
| 118 | CCNH | 17502 | 0.234 | 0.4362 | No | ||
| 119 | POLR1A | 17545 | 0.230 | 0.4367 | No | ||
| 120 | H3F3A | 18071 | 0.204 | 0.4243 | No | ||
| 121 | POLR2J | 18098 | 0.201 | 0.4250 | No | ||
| 122 | POLRMT | 18192 | 0.196 | 0.4239 | No | ||
| 123 | POLR3H | 18199 | 0.195 | 0.4251 | No | ||
| 124 | H3F3AP6 | 18211 | 0.194 | 0.4261 | No | ||
| 125 | ELL | 18700 | 0.159 | 0.4143 | No | ||
| 126 | HIST1H2AB | 18760 | 0.155 | 0.4138 | No | ||
| 127 | POLR2D | 18862 | 0.147 | 0.4122 | No | ||
| 128 | TAF11 | 19049 | 0.133 | 0.4082 | No | ||
| 129 | SUPT4H1 | 19077 | 0.130 | 0.4084 | No | ||
| 130 | BRF2 | 19566 | 0.115 | 0.3963 | No | ||
| 131 | HIST1H4A | 19810 | 0.110 | 0.3906 | No | ||
| 132 | SRSF1 | 19942 | 0.106 | 0.3879 | No | ||
| 133 | H3F3B | 19965 | 0.105 | 0.3880 | No | ||
| 134 | TAF10 | 20013 | 0.103 | 0.3875 | No | ||
| 135 | LSM10 | 20057 | 0.099 | 0.3870 | No | ||
| 136 | POLR2I | 20092 | 0.097 | 0.3868 | No | ||
| 137 | TAF4 | 20288 | 0.083 | 0.3822 | No | ||
| 138 | GTF3C5 | 20425 | 0.074 | 0.3792 | No | ||
| 139 | SNRPF | 20434 | 0.073 | 0.3794 | No | ||
| 140 | SNRPG | 20483 | 0.070 | 0.3787 | No | ||
| 141 | SRSF7 | 20530 | 0.066 | 0.3779 | No | ||
| 142 | TAF1C | 20659 | 0.059 | 0.3749 | No | ||
| 143 | SRRM1 | 20926 | 0.042 | 0.3682 | No | ||
| 144 | SLBP | 21350 | 0.015 | 0.3571 | No | ||
| 145 | GTF2H3 | 21535 | 0.014 | 0.3523 | No | ||
| 146 | GTF3C3 | 21592 | 0.010 | 0.3509 | No | ||
| 147 | POLR1D | 21594 | 0.010 | 0.3510 | No | ||
| 148 | NCBP1 | 21677 | 0.004 | 0.3488 | No | ||
| 149 | SRSF3 | 21795 | -0.005 | 0.3458 | No | ||
| 150 | RBM8A | 21805 | -0.005 | 0.3456 | No | ||
| 151 | GTF2E1 | 21895 | -0.012 | 0.3433 | No | ||
| 152 | CBX3 | 21949 | -0.016 | 0.3420 | No | ||
| 153 | TAF1 | 21973 | -0.018 | 0.3415 | No | ||
| 154 | SNAPC5 | 21988 | -0.019 | 0.3413 | No | ||
| 155 | SNRPE | 22094 | -0.026 | 0.3387 | No | ||
| 156 | POLR3B | 22164 | -0.031 | 0.3371 | No | ||
| 157 | SSRP1 | 22219 | -0.035 | 0.3359 | No | ||
| 158 | CDK7 | 22286 | -0.041 | 0.3345 | No | ||
| 159 | ERCC3 | 22319 | -0.042 | 0.3339 | No | ||
| 160 | TAF6 | 22343 | -0.045 | 0.3336 | No | ||
| 161 | SRSF9 | 22494 | -0.055 | 0.3300 | No | ||
| 162 | CPSF1 | 22724 | -0.071 | 0.3245 | No | ||
| 163 | ERCC6 | 22771 | -0.074 | 0.3238 | No | ||
| 164 | NFX1 | 22917 | -0.084 | 0.3205 | No | ||
| 165 | SRSF6 | 22965 | -0.088 | 0.3199 | No | ||
| 166 | MAPK3 | 24183 | -0.177 | 0.2890 | No | ||
| 167 | HIST1H2BM | 24664 | -0.214 | 0.2778 | No | ||
| 168 | CSTF1 | 25399 | -0.270 | 0.2602 | No | ||
| 169 | GTF3C2 | 25451 | -0.275 | 0.2608 | No | ||
| 170 | ERCC2 | 25540 | -0.281 | 0.2604 | No | ||
| 171 | CPSF3 | 25570 | -0.283 | 0.2616 | No | ||
| 172 | POLR3GL | 25593 | -0.284 | 0.2630 | No | ||
| 173 | POLR1B | 25795 | -0.298 | 0.2597 | No | ||
| 174 | BRF1 | 25901 | -0.308 | 0.2591 | No | ||
| 175 | SUPT16H | 26092 | -0.321 | 0.2563 | No | ||
| 176 | CPSF2 | 26510 | -0.355 | 0.2477 | No | ||
| 177 | NUDT21 | 26787 | -0.377 | 0.2431 | No | ||
| 178 | RNMT | 26932 | -0.387 | 0.2419 | No | ||
| 179 | LSM11 | 27382 | -0.427 | 0.2330 | No | ||
| 180 | CPSF7 | 27554 | -0.442 | 0.2316 | No | ||
| 181 | HIST1H3D | 27756 | -0.459 | 0.2294 | No | ||
| 182 | POLR2H | 27892 | -0.472 | 0.2291 | No | ||
| 183 | SRSF2 | 28096 | -0.490 | 0.2272 | No | ||
| 184 | UBTF | 28553 | -0.532 | 0.2188 | No | ||
| 185 | POU2F1 | 28646 | -0.542 | 0.2201 | No | ||
| 186 | EHMT2 | 28773 | -0.556 | 0.2207 | No | ||
| 187 | KAT2B | 32917 | -1.204 | 0.1195 | No | ||
| 188 | HIST1H4F | 36591 | -2.244 | 0.0380 | No |
