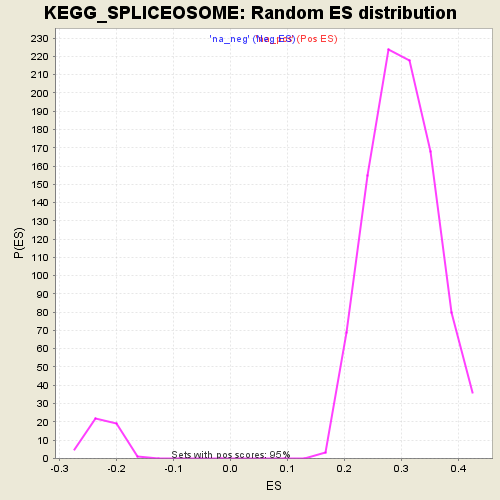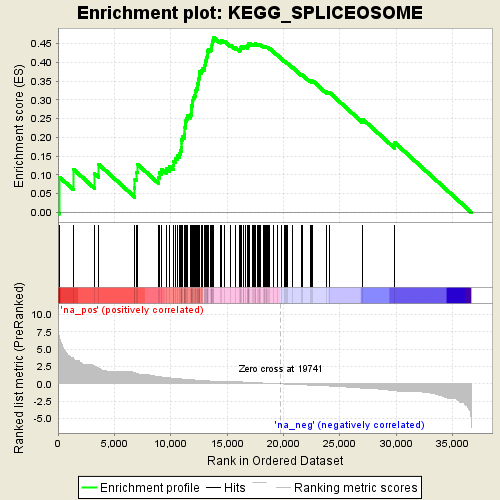
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | NOV0135_D3_log2foldchange_preranked |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_pos |
| GeneSet | KEGG_SPLICEOSOME |
| Enrichment Score (ES) | 0.46626765 |
| Normalized Enrichment Score (NES) | 1.5472292 |
| Nominal p-value | 0.0 |
| FDR q-value | 1.0 |
| FWER p-Value | 1.0 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | HSPA6 | 162 | 6.448 | 0.0932 | Yes | ||
| 2 | HSPA1A | 1380 | 3.616 | 0.1146 | Yes | ||
| 3 | SRSF8 | 3236 | 2.603 | 0.1032 | Yes | ||
| 4 | HSPA1B | 3600 | 2.310 | 0.1282 | Yes | ||
| 5 | HSPA2 | 6802 | 1.619 | 0.0651 | Yes | ||
| 6 | PRPF18 | 6820 | 1.602 | 0.0889 | Yes | ||
| 7 | EIF4A3 | 6983 | 1.520 | 0.1075 | Yes | ||
| 8 | BUD31 | 7046 | 1.481 | 0.1282 | Yes | ||
| 9 | PUF60 | 8933 | 1.037 | 0.0923 | Yes | ||
| 10 | HSPA8 | 9010 | 1.014 | 0.1055 | Yes | ||
| 11 | CCDC12 | 9182 | 0.973 | 0.1156 | Yes | ||
| 12 | PRPF3 | 9593 | 0.913 | 0.1182 | Yes | ||
| 13 | DHX8 | 9899 | 0.858 | 0.1228 | Yes | ||
| 14 | MAGOH | 10240 | 0.792 | 0.1255 | Yes | ||
| 15 | HNRNPU | 10263 | 0.787 | 0.1368 | Yes | ||
| 16 | CTNNBL1 | 10405 | 0.763 | 0.1445 | Yes | ||
| 17 | PRPF38A | 10557 | 0.747 | 0.1516 | Yes | ||
| 18 | PRPF38B | 10800 | 0.710 | 0.1558 | Yes | ||
| 19 | SNRPB2 | 10885 | 0.697 | 0.1640 | Yes | ||
| 20 | BCAS2 | 10934 | 0.692 | 0.1732 | Yes | ||
| 21 | CWC15 | 10952 | 0.688 | 0.1831 | Yes | ||
| 22 | SMNDC1 | 10955 | 0.688 | 0.1935 | Yes | ||
| 23 | DDX5 | 11026 | 0.676 | 0.2018 | Yes | ||
| 24 | SNW1 | 11193 | 0.657 | 0.2072 | Yes | ||
| 25 | SNRPA1 | 11249 | 0.652 | 0.2155 | Yes | ||
| 26 | PRPF4 | 11250 | 0.651 | 0.2254 | Yes | ||
| 27 | CDC40 | 11283 | 0.648 | 0.2343 | Yes | ||
| 28 | TRA2B | 11298 | 0.645 | 0.2437 | Yes | ||
| 29 | PLRG1 | 11370 | 0.634 | 0.2514 | Yes | ||
| 30 | RBM25 | 11500 | 0.619 | 0.2572 | Yes | ||
| 31 | RBM22 | 11708 | 0.596 | 0.2605 | Yes | ||
| 32 | SF3B4 | 11803 | 0.585 | 0.2668 | Yes | ||
| 33 | ISY1 | 11807 | 0.584 | 0.2756 | Yes | ||
| 34 | PRPF40A | 11815 | 0.583 | 0.2842 | Yes | ||
| 35 | WBP11 | 11936 | 0.567 | 0.2895 | Yes | ||
| 36 | PQBP1 | 11947 | 0.566 | 0.2978 | Yes | ||
| 37 | U2AF2 | 11969 | 0.563 | 0.3058 | Yes | ||
| 38 | CRNKL1 | 12065 | 0.553 | 0.3115 | Yes | ||
| 39 | SRSF7 | 12184 | 0.539 | 0.3165 | Yes | ||
| 40 | NCBP1 | 12186 | 0.539 | 0.3246 | Yes | ||
| 41 | AQR | 12242 | 0.530 | 0.3311 | Yes | ||
| 42 | LSM3 | 12351 | 0.519 | 0.3360 | Yes | ||
| 43 | DHX38 | 12380 | 0.516 | 0.3430 | Yes | ||
| 44 | RBM17 | 12467 | 0.506 | 0.3483 | Yes | ||
| 45 | SLU7 | 12476 | 0.505 | 0.3558 | Yes | ||
| 46 | DDX46 | 12504 | 0.501 | 0.3626 | Yes | ||
| 47 | PRPF19 | 12525 | 0.498 | 0.3696 | Yes | ||
| 48 | CHERP | 12552 | 0.493 | 0.3763 | Yes | ||
| 49 | HNRNPM | 12757 | 0.472 | 0.3779 | Yes | ||
| 50 | PRPF8 | 12818 | 0.465 | 0.3833 | Yes | ||
| 51 | NCBP2 | 13012 | 0.443 | 0.3847 | Yes | ||
| 52 | SNRNP40 | 13020 | 0.443 | 0.3912 | Yes | ||
| 53 | PPIH | 13038 | 0.441 | 0.3974 | Yes | ||
| 54 | SNRPB | 13053 | 0.439 | 0.4037 | Yes | ||
| 55 | SNRPD3 | 13144 | 0.429 | 0.4077 | Yes | ||
| 56 | U2SURP | 13202 | 0.423 | 0.4126 | Yes | ||
| 57 | SRSF4 | 13228 | 0.421 | 0.4183 | Yes | ||
| 58 | DHX15 | 13243 | 0.420 | 0.4242 | Yes | ||
| 59 | SNRPG | 13244 | 0.420 | 0.4306 | Yes | ||
| 60 | SF3A3 | 13365 | 0.408 | 0.4335 | Yes | ||
| 61 | HNRNPK | 13563 | 0.387 | 0.4339 | Yes | ||
| 62 | PHF5A | 13614 | 0.381 | 0.4384 | Yes | ||
| 63 | SRSF1 | 13641 | 0.378 | 0.4434 | Yes | ||
| 64 | SYF2 | 13652 | 0.378 | 0.4488 | Yes | ||
| 65 | TRA2A | 13702 | 0.371 | 0.4531 | Yes | ||
| 66 | DDX42 | 13736 | 0.366 | 0.4577 | Yes | ||
| 67 | RBM8A | 13775 | 0.363 | 0.4622 | Yes | ||
| 68 | EFTUD2 | 13824 | 0.357 | 0.4663 | Yes | ||
| 69 | SNRNP27 | 14365 | 0.341 | 0.4567 | No | ||
| 70 | SNRPC | 14499 | 0.338 | 0.4581 | No | ||
| 71 | PRPF31 | 14739 | 0.336 | 0.4567 | No | ||
| 72 | DHX16 | 15306 | 0.328 | 0.4461 | No | ||
| 73 | SF3B5 | 15761 | 0.309 | 0.4384 | No | ||
| 74 | ZMAT2 | 16098 | 0.276 | 0.4334 | No | ||
| 75 | LSM5 | 16212 | 0.265 | 0.4343 | No | ||
| 76 | HSPA1L | 16213 | 0.265 | 0.4383 | No | ||
| 77 | THOC1 | 16254 | 0.261 | 0.4412 | No | ||
| 78 | CDC5L | 16445 | 0.245 | 0.4397 | No | ||
| 79 | SF3A1 | 16460 | 0.244 | 0.4430 | No | ||
| 80 | XAB2 | 16630 | 0.228 | 0.4418 | No | ||
| 81 | SRSF6 | 16793 | 0.215 | 0.4406 | No | ||
| 82 | SRSF3 | 16795 | 0.215 | 0.4439 | No | ||
| 83 | SNRPF | 16865 | 0.209 | 0.4451 | No | ||
| 84 | THOC2 | 16889 | 0.207 | 0.4476 | No | ||
| 85 | USP39 | 16908 | 0.205 | 0.4502 | No | ||
| 86 | SF3B2 | 16987 | 0.198 | 0.4511 | No | ||
| 87 | THOC3 | 17202 | 0.177 | 0.4479 | No | ||
| 88 | SNRNP70 | 17311 | 0.168 | 0.4475 | No | ||
| 89 | SRSF10 | 17425 | 0.158 | 0.4468 | No | ||
| 90 | LSM6 | 17485 | 0.153 | 0.4475 | No | ||
| 91 | SNRPD2 | 17500 | 0.152 | 0.4494 | No | ||
| 92 | SNRNP200 | 17670 | 0.140 | 0.4469 | No | ||
| 93 | SNRPE | 17768 | 0.134 | 0.4463 | No | ||
| 94 | TCERG1 | 17853 | 0.128 | 0.4460 | No | ||
| 95 | SART1 | 17868 | 0.127 | 0.4475 | No | ||
| 96 | SF3B1 | 17934 | 0.122 | 0.4476 | No | ||
| 97 | RBMX | 18201 | 0.105 | 0.4419 | No | ||
| 98 | PPIE | 18295 | 0.098 | 0.4408 | No | ||
| 99 | PRPF6 | 18304 | 0.097 | 0.4421 | No | ||
| 100 | TXNL4A | 18387 | 0.091 | 0.4412 | No | ||
| 101 | HNRNPC | 18490 | 0.084 | 0.4397 | No | ||
| 102 | PPIL1 | 18501 | 0.083 | 0.4407 | No | ||
| 103 | SNRPD1 | 18580 | 0.079 | 0.4397 | No | ||
| 104 | MAGOHB | 18700 | 0.072 | 0.4376 | No | ||
| 105 | DDX23 | 18791 | 0.067 | 0.4361 | No | ||
| 106 | HNRNPA3 | 19078 | 0.046 | 0.4290 | No | ||
| 107 | SRSF9 | 19482 | 0.017 | 0.4182 | No | ||
| 108 | SF3A2 | 19826 | -0.006 | 0.4089 | No | ||
| 109 | SRSF2 | 20091 | -0.029 | 0.4021 | No | ||
| 110 | SF3B3 | 20148 | -0.035 | 0.4011 | No | ||
| 111 | ACIN1 | 20253 | -0.044 | 0.3989 | No | ||
| 112 | DDX39B | 20358 | -0.053 | 0.3969 | No | ||
| 113 | SRSF5 | 20829 | -0.083 | 0.3853 | No | ||
| 114 | LSM2 | 21603 | -0.137 | 0.3662 | No | ||
| 115 | PCBP1 | 21675 | -0.142 | 0.3664 | No | ||
| 116 | LSM4 | 22360 | -0.192 | 0.3506 | No | ||
| 117 | LSM7 | 22516 | -0.206 | 0.3495 | No | ||
| 118 | HNRNPA1L2 | 22607 | -0.216 | 0.3503 | No | ||
| 119 | SNRPA | 23827 | -0.299 | 0.3214 | No | ||
| 120 | HNRNPA1 | 24039 | -0.318 | 0.3205 | No | ||
| 121 | NAA38 | 27009 | -0.639 | 0.2489 | No | ||
| 122 | PRPF40B | 29883 | -1.004 | 0.1855 | No |