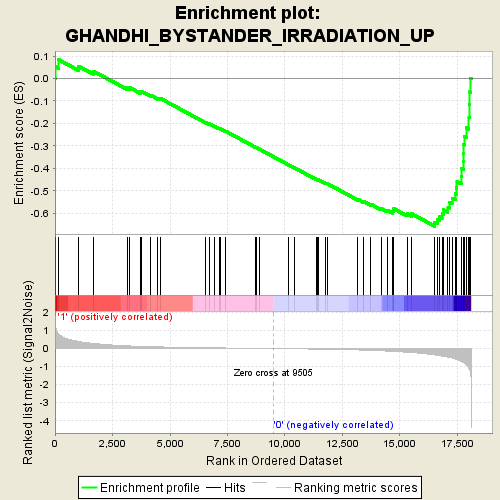
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | NS50719_832_collapsed_to_symbols.NS50719_832.cls #arhinia_versus_control.NS50719_832.cls #arhinia_versus_control_repos |
| Phenotype | NS50719_832.cls#arhinia_versus_control_repos |
| Upregulated in class | 0 |
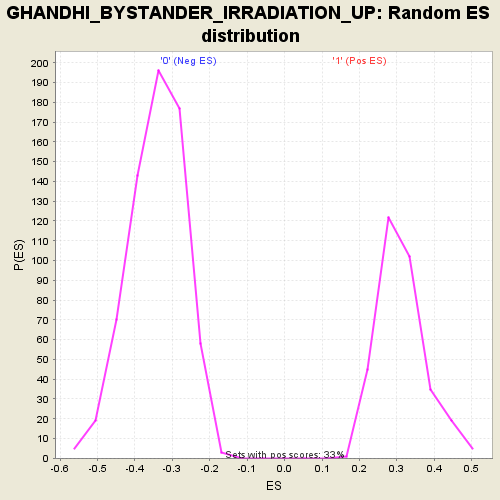
| GeneSet | GHANDHI_BYSTANDER_IRRADIATION_UP |
| Enrichment Score (ES) | -0.6582263 |
| Normalized Enrichment Score (NES) | -1.929196 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.050783467 |
| FWER p-Value | 0.175 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | PTGES | PTGES Entrez, Source | prostaglandin E synthase | 35 | 1.117 | 0.0519 | No |
| 2 | PLEK2 | PLEK2 Entrez, Source | pleckstrin 2 | 147 | 0.801 | 0.0844 | No |
| 3 | LIF | LIF Entrez, Source | leukemia inhibitory factor (cholinergic differentiation factor) | 1019 | 0.396 | 0.0552 | No |
| 4 | B3GNT5 | B3GNT5 Entrez, Source | UDP-GlcNAc:betaGal beta-1,3-N-acetylglucosaminyltransferase 5 | 1657 | 0.282 | 0.0335 | No |
| 5 | NFKBIZ | NFKBIZ Entrez, Source | nuclear factor of kappa light polypeptide gene enhancer in B-cells inhibitor, zeta | 3134 | 0.150 | -0.0411 | No |
| 6 | SRSF12 | 3254 | 0.143 | -0.0408 | No | ||
| 7 | ZCCHC6 | ZCCHC6 Entrez, Source | zinc finger, CCHC domain containing 6 | 3701 | 0.122 | -0.0596 | No |
| 8 | ZC3H12C | ZC3H12C Entrez, Source | zinc finger CCCH-type containing 12C | 3766 | 0.119 | -0.0575 | No |
| 9 | AHR | AHR Entrez, Source | aryl hydrocarbon receptor | 4148 | 0.104 | -0.0736 | No |
| 10 | RGS9 | RGS9 Entrez, Source | regulator of G-protein signalling 9 | 4474 | 0.093 | -0.0871 | No |
| 11 | ICAM1 | ICAM1 Entrez, Source | intercellular adhesion molecule 1 (CD54), human rhinovirus receptor | 4569 | 0.091 | -0.0879 | No |
| 12 | SLC6A15 | SLC6A15 Entrez, Source | solute carrier family 6, member 15 | 6560 | 0.045 | -0.1961 | No |
| 13 | RELB | RELB Entrez, Source | v-rel reticuloendotheliosis viral oncogene homolog B, nuclear factor of kappa light polypeptide gene enhancer in B-cells 3 (avian) | 6706 | 0.042 | -0.2021 | No |
| 14 | SLC39A8 | SLC39A8 Entrez, Source | solute carrier family 39 (zinc transporter), member 8 | 6912 | 0.039 | -0.2116 | No |
| 15 | LRRC8B | LRRC8B Entrez, Source | leucine rich repeat containing 8 family, member B | 7147 | 0.035 | -0.2229 | No |
| 16 | DUSP2 | DUSP2 Entrez, Source | dual specificity phosphatase 2 | 7189 | 0.034 | -0.2235 | No |
| 17 | STC1 | STC1 Entrez, Source | stanniocalcin 1 | 7430 | 0.030 | -0.2353 | No |
| 18 | DNAJB4 | DNAJB4 Entrez, Source | DnaJ (Hsp40) homolog, subfamily B, member 4 | 8729 | 0.011 | -0.3067 | No |
| 19 | FGF2 | FGF2 Entrez, Source | fibroblast growth factor 2 (basic) | 8748 | 0.011 | -0.3072 | No |
| 20 | MCTP1 | MCTP1 Entrez, Source | multiple C2 domains, transmembrane 1 | 8875 | 0.009 | -0.3138 | No |
| 21 | ATP6V1C1 | ATP6V1C1 Entrez, Source | ATPase, H+ transporting, lysosomal 42kDa, V1 subunit C1 | 10172 | -0.010 | -0.3851 | No |
| 22 | INHBA | INHBA Entrez, Source | inhibin, beta A (activin A, activin AB alpha polypeptide) | 10424 | -0.015 | -0.3984 | No |
| 23 | NAMPT | 11376 | -0.031 | -0.4496 | No | ||
| 24 | EPB41L4B | EPB41L4B Entrez, Source | erythrocyte membrane protein band 4.1 like 4B | 11425 | -0.032 | -0.4507 | No |
| 25 | OSGIN2 | OSGIN2 Entrez, Source | oxidative stress induced growth inhibitor family member 2 | 11458 | -0.033 | -0.4509 | No |
| 26 | SLC16A3 | SLC16A3 Entrez, Source | solute carrier family 16, member 3 (monocarboxylic acid transporter 4) | 11766 | -0.039 | -0.4660 | No |
| 27 | SOD2 | SOD2 Entrez, Source | superoxide dismutase 2, mitochondrial | 11828 | -0.040 | -0.4675 | No |
| 28 | GCLM | GCLM Entrez, Source | glutamate-cysteine ligase, modifier subunit | 13171 | -0.078 | -0.5381 | No |
| 29 | SAT1 | SAT1 Entrez, Source | spermidine/spermine N1-acetyltransferase 1 | 13415 | -0.087 | -0.5474 | No |
| 30 | SLC39A14 | SLC39A14 Entrez, Source | solute carrier family 39 (zinc transporter), member 14 | 13738 | -0.101 | -0.5604 | No |
| 31 | DUSP5 | DUSP5 Entrez, Source | dual specificity phosphatase 5 | 14184 | -0.125 | -0.5791 | No |
| 32 | MMP10 | MMP10 Entrez, Source | matrix metallopeptidase 10 (stromelysin 2) | 14442 | -0.140 | -0.5866 | No |
| 33 | TNFAIP3 | TNFAIP3 Entrez, Source | tumor necrosis factor, alpha-induced protein 3 | 14692 | -0.155 | -0.5929 | No |
| 34 | LAMB3 | LAMB3 Entrez, Source | laminin, beta 3 | 14706 | -0.156 | -0.5861 | No |
| 35 | PLA2G4A | PLA2G4A Entrez, Source | phospholipase A2, group IVA (cytosolic, calcium-dependent) | 14737 | -0.158 | -0.5801 | No |
| 36 | GCH1 | GCH1 Entrez, Source | GTP cyclohydrolase 1 (dopa-responsive dystonia) | 15319 | -0.204 | -0.6025 | No |
| 37 | MCTP2 | MCTP2 Entrez, Source | multiple C2 domains, transmembrane 2 | 15513 | -0.221 | -0.6026 | No |
| 38 | TESC | TESC Entrez, Source | tescalcin | 16518 | -0.350 | -0.6414 | Yes |
| 39 | GPR68 | GPR68 Entrez, Source | G protein-coupled receptor 68 | 16634 | -0.374 | -0.6297 | Yes |
| 40 | CXCL2 | CXCL2 Entrez, Source | chemokine (C-X-C motif) ligand 2 | 16710 | -0.388 | -0.6151 | Yes |
| 41 | C2CD4B | 16832 | -0.416 | -0.6018 | Yes | ||
| 42 | MT1H | MT1H Entrez, Source | metallothionein 1H | 16899 | -0.430 | -0.5848 | Yes |
| 43 | SLC7A11 | SLC7A11 Entrez, Source | solute carrier family 7, (cationic amino acid transporter, y+ system) member 11 | 17090 | -0.475 | -0.5724 | Yes |
| 44 | MT1E | MT1E Entrez, Source | metallothionein 1E (functional) | 17172 | -0.498 | -0.5529 | Yes |
| 45 | IRAK3 | IRAK3 Entrez, Source | interleukin-1 receptor-associated kinase 3 | 17267 | -0.524 | -0.5329 | Yes |
| 46 | CXCL5 | CXCL5 Entrez, Source | chemokine (C-X-C motif) ligand 5 | 17403 | -0.575 | -0.5127 | Yes |
| 47 | CCK | CCK Entrez, Source | cholecystokinin | 17451 | -0.595 | -0.4866 | Yes |
| 48 | POPDC3 | POPDC3 Entrez, Source | popeye domain containing 3 | 17481 | -0.609 | -0.4589 | Yes |
| 49 | GADD45G | GADD45G Entrez, Source | growth arrest and DNA-damage-inducible, gamma | 17662 | -0.706 | -0.4348 | Yes |
| 50 | G0S2 | G0S2 Entrez, Source | G0/G1switch 2 | 17689 | -0.721 | -0.4015 | Yes |
| 51 | MT1G | MT1G Entrez, Source | metallothionein 1G | 17761 | -0.766 | -0.3685 | Yes |
| 52 | IL1A | IL1A Entrez, Source | interleukin 1, alpha | 17769 | -0.771 | -0.3317 | Yes |
| 53 | CREB5 | CREB5 Entrez, Source | cAMP responsive element binding protein 5 | 17778 | -0.779 | -0.2947 | Yes |
| 54 | TSLP | TSLP Entrez, Source | - | 17806 | -0.808 | -0.2572 | Yes |
| 55 | CXCL3 | CXCL3 Entrez, Source | chemokine (C-X-C motif) ligand 3 | 17881 | -0.889 | -0.2185 | Yes |
| 56 | CXCL1 | CXCL1 Entrez, Source | chemokine (C-X-C motif) ligand 1 (melanoma growth stimulating activity, alpha) | 18003 | -1.099 | -0.1722 | Yes |
| 57 | PTGS2 | PTGS2 Entrez, Source | prostaglandin-endoperoxide synthase 2 (prostaglandin G/H synthase and cyclooxygenase) | 18030 | -1.172 | -0.1172 | Yes |
| 58 | MT1X | MT1X Entrez, Source | metallothionein 1X | 18042 | -1.203 | -0.0598 | Yes |
| 59 | RNF128 | RNF128 Entrez, Source | ring finger protein 128 | 18066 | -1.304 | 0.0018 | Yes |