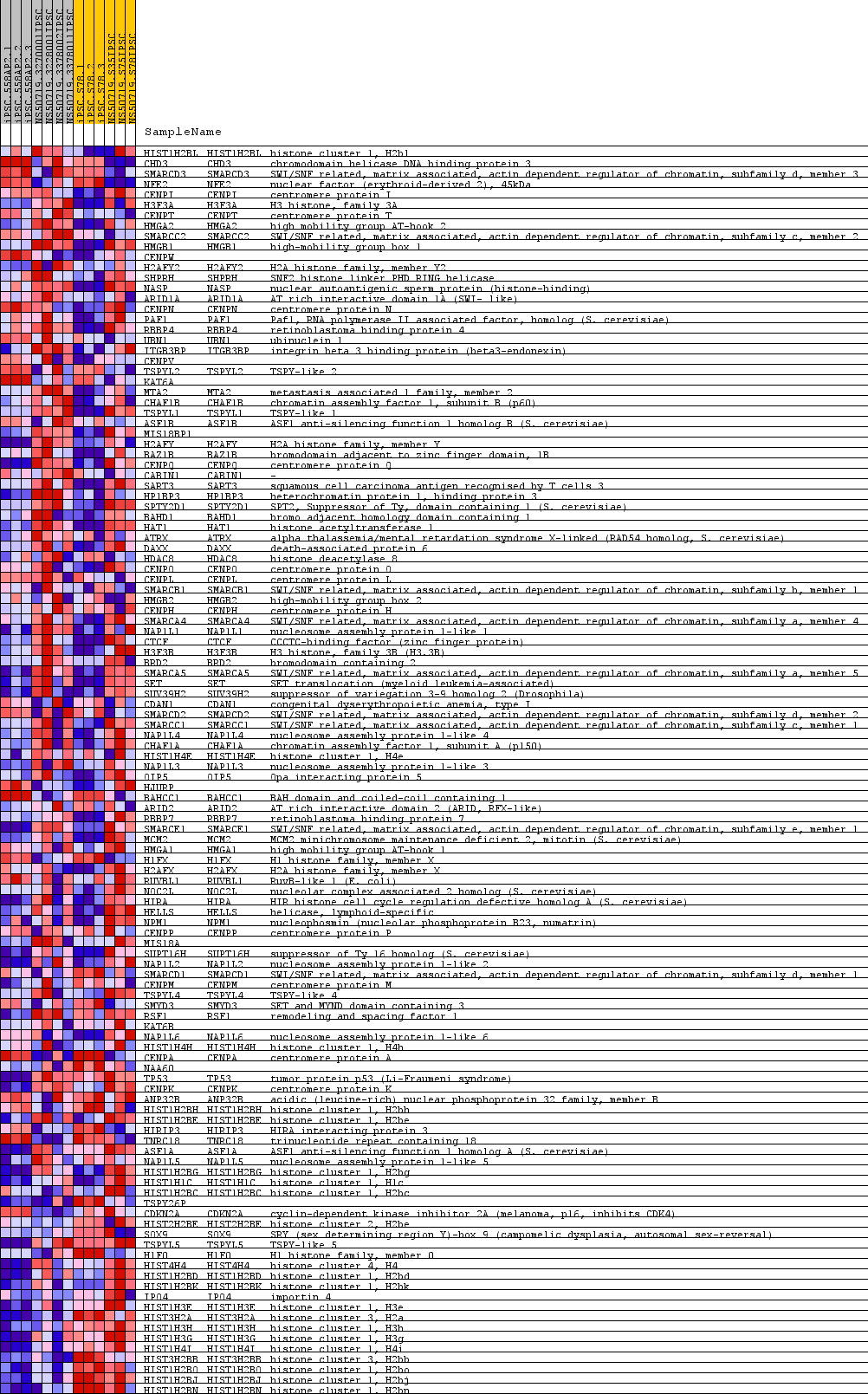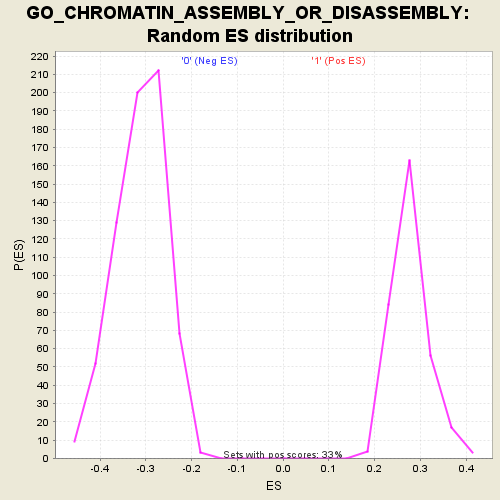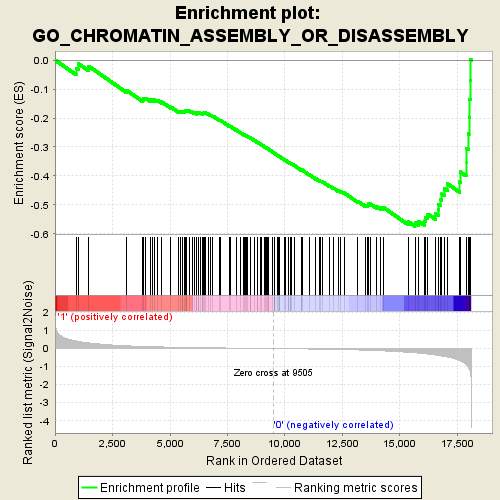
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | NS50719_832_collapsed_to_symbols.NS50719_832.cls #arhinia_versus_control.NS50719_832.cls #arhinia_versus_control_repos |
| Phenotype | NS50719_832.cls#arhinia_versus_control_repos |
| Upregulated in class | 0 |
| GeneSet | GO_CHROMATIN_ASSEMBLY_OR_DISASSEMBLY |
| Enrichment Score (ES) | -0.57363176 |
| Normalized Enrichment Score (NES) | -1.8435796 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.06340994 |
| FWER p-Value | 0.683 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | HIST1H2BL | HIST1H2BL Entrez, Source | histone cluster 1, H2bl | 914 | 0.421 | -0.0282 | No |
| 2 | CHD3 | CHD3 Entrez, Source | chromodomain helicase DNA binding protein 3 | 1021 | 0.396 | -0.0129 | No |
| 3 | SMARCD3 | SMARCD3 Entrez, Source | SWI/SNF related, matrix associated, actin dependent regulator of chromatin, subfamily d, member 3 | 1468 | 0.310 | -0.0211 | No |
| 4 | NFE2 | NFE2 Entrez, Source | nuclear factor (erythroid-derived 2), 45kDa | 3122 | 0.152 | -0.1049 | No |
| 5 | CENPI | CENPI Entrez, Source | centromere protein I | 3806 | 0.117 | -0.1366 | No |
| 6 | H3F3A | H3F3A Entrez, Source | H3 histone, family 3A | 3841 | 0.116 | -0.1323 | No |
| 7 | CENPT | CENPT Entrez, Source | centromere protein T | 3935 | 0.112 | -0.1315 | No |
| 8 | HMGA2 | HMGA2 Entrez, Source | high mobility group AT-hook 2 | 4137 | 0.105 | -0.1370 | No |
| 9 | SMARCC2 | SMARCC2 Entrez, Source | SWI/SNF related, matrix associated, actin dependent regulator of chromatin, subfamily c, member 2 | 4236 | 0.101 | -0.1370 | No |
| 10 | HMGB1 | HMGB1 Entrez, Source | high-mobility group box 1 | 4328 | 0.098 | -0.1368 | No |
| 11 | CENPW | 4465 | 0.094 | -0.1394 | No | ||
| 12 | H2AFY2 | H2AFY2 Entrez, Source | H2A histone family, member Y2 | 4641 | 0.088 | -0.1444 | No |
| 13 | SHPRH | SHPRH Entrez, Source | SNF2 histone linker PHD RING helicase | 5029 | 0.077 | -0.1618 | No |
| 14 | NASP | NASP Entrez, Source | nuclear autoantigenic sperm protein (histone-binding) | 5381 | 0.069 | -0.1776 | No |
| 15 | ARID1A | ARID1A Entrez, Source | AT rich interactive domain 1A (SWI- like) | 5460 | 0.067 | -0.1783 | No |
| 16 | CENPN | CENPN Entrez, Source | centromere protein N | 5536 | 0.065 | -0.1790 | No |
| 17 | PAF1 | PAF1 Entrez, Source | Paf1, RNA polymerase II associated factor, homolog (S. cerevisiae) | 5563 | 0.064 | -0.1770 | No |
| 18 | RBBP4 | RBBP4 Entrez, Source | retinoblastoma binding protein 4 | 5619 | 0.063 | -0.1767 | No |
| 19 | UBN1 | UBN1 Entrez, Source | ubinuclein 1 | 5658 | 0.062 | -0.1755 | No |
| 20 | ITGB3BP | ITGB3BP Entrez, Source | integrin beta 3 binding protein (beta3-endonexin) | 5696 | 0.062 | -0.1742 | No |
| 21 | CENPV | 5733 | 0.061 | -0.1729 | No | ||
| 22 | TSPYL2 | TSPYL2 Entrez, Source | TSPY-like 2 | 5837 | 0.059 | -0.1755 | No |
| 23 | KAT6A | 5958 | 0.056 | -0.1791 | No | ||
| 24 | MTA2 | MTA2 Entrez, Source | metastasis associated 1 family, member 2 | 6084 | 0.054 | -0.1832 | No |
| 25 | CHAF1B | CHAF1B Entrez, Source | chromatin assembly factor 1, subunit B (p60) | 6148 | 0.053 | -0.1839 | No |
| 26 | TSPYL1 | TSPYL1 Entrez, Source | TSPY-like 1 | 6158 | 0.052 | -0.1816 | No |
| 27 | ASF1B | ASF1B Entrez, Source | ASF1 anti-silencing function 1 homolog B (S. cerevisiae) | 6231 | 0.051 | -0.1829 | No |
| 28 | MIS18BP1 | 6235 | 0.051 | -0.1803 | No | ||
| 29 | H2AFY | H2AFY Entrez, Source | H2A histone family, member Y | 6326 | 0.049 | -0.1827 | No |
| 30 | BAZ1B | BAZ1B Entrez, Source | bromodomain adjacent to zinc finger domain, 1B | 6411 | 0.047 | -0.1848 | No |
| 31 | CENPQ | CENPQ Entrez, Source | centromere protein Q | 6423 | 0.047 | -0.1829 | No |
| 32 | CABIN1 | CABIN1 Entrez, Source | - | 6460 | 0.047 | -0.1824 | No |
| 33 | SART3 | SART3 Entrez, Source | squamous cell carcinoma antigen recognised by T cells 3 | 6510 | 0.046 | -0.1827 | No |
| 34 | HP1BP3 | HP1BP3 Entrez, Source | heterochromatin protein 1, binding protein 3 | 6531 | 0.045 | -0.1814 | No |
| 35 | SPTY2D1 | SPTY2D1 Entrez, Source | SPT2, Suppressor of Ty, domain containing 1 (S. cerevisiae) | 6657 | 0.043 | -0.1860 | No |
| 36 | BAHD1 | BAHD1 Entrez, Source | bromo adjacent homology domain containing 1 | 6763 | 0.041 | -0.1897 | No |
| 37 | HAT1 | HAT1 Entrez, Source | histone acetyltransferase 1 | 6863 | 0.040 | -0.1930 | No |
| 38 | ATRX | ATRX Entrez, Source | alpha thalassemia/mental retardation syndrome X-linked (RAD54 homolog, S. cerevisiae) | 7156 | 0.035 | -0.2074 | No |
| 39 | DAXX | DAXX Entrez, Source | death-associated protein 6 | 7204 | 0.034 | -0.2082 | No |
| 40 | HDAC8 | HDAC8 Entrez, Source | histone deacetylase 8 | 7567 | 0.028 | -0.2268 | No |
| 41 | CENPO | CENPO Entrez, Source | centromere protein O | 7639 | 0.027 | -0.2293 | No |
| 42 | CENPL | CENPL Entrez, Source | centromere protein L | 7873 | 0.023 | -0.2411 | No |
| 43 | SMARCB1 | SMARCB1 Entrez, Source | SWI/SNF related, matrix associated, actin dependent regulator of chromatin, subfamily b, member 1 | 8056 | 0.021 | -0.2501 | No |
| 44 | HMGB2 | HMGB2 Entrez, Source | high-mobility group box 2 | 8213 | 0.018 | -0.2578 | No |
| 45 | CENPH | CENPH Entrez, Source | centromere protein H | 8242 | 0.018 | -0.2584 | No |
| 46 | SMARCA4 | SMARCA4 Entrez, Source | SWI/SNF related, matrix associated, actin dependent regulator of chromatin, subfamily a, member 4 | 8281 | 0.017 | -0.2596 | No |
| 47 | NAP1L1 | NAP1L1 Entrez, Source | nucleosome assembly protein 1-like 1 | 8343 | 0.017 | -0.2621 | No |
| 48 | CTCF | CTCF Entrez, Source | CCCTC-binding factor (zinc finger protein) | 8379 | 0.016 | -0.2631 | No |
| 49 | H3F3B | H3F3B Entrez, Source | H3 histone, family 3B (H3.3B) | 8507 | 0.014 | -0.2695 | No |
| 50 | BRD2 | BRD2 Entrez, Source | bromodomain containing 2 | 8521 | 0.014 | -0.2694 | No |
| 51 | SMARCA5 | SMARCA5 Entrez, Source | SWI/SNF related, matrix associated, actin dependent regulator of chromatin, subfamily a, member 5 | 8681 | 0.012 | -0.2777 | No |
| 52 | SET | SET Entrez, Source | SET translocation (myeloid leukemia-associated) | 8806 | 0.010 | -0.2840 | No |
| 53 | SUV39H2 | SUV39H2 Entrez, Source | suppressor of variegation 3-9 homolog 2 (Drosophila) | 8816 | 0.010 | -0.2840 | No |
| 54 | CDAN1 | CDAN1 Entrez, Source | congenital dyserythropoietic anemia, type I | 8941 | 0.008 | -0.2905 | No |
| 55 | SMARCD2 | SMARCD2 Entrez, Source | SWI/SNF related, matrix associated, actin dependent regulator of chromatin, subfamily d, member 2 | 8965 | 0.008 | -0.2914 | No |
| 56 | SMARCC1 | SMARCC1 Entrez, Source | SWI/SNF related, matrix associated, actin dependent regulator of chromatin, subfamily c, member 1 | 9104 | 0.005 | -0.2987 | No |
| 57 | NAP1L4 | NAP1L4 Entrez, Source | nucleosome assembly protein 1-like 4 | 9139 | 0.005 | -0.3004 | No |
| 58 | CHAF1A | CHAF1A Entrez, Source | chromatin assembly factor 1, subunit A (p150) | 9205 | 0.004 | -0.3038 | No |
| 59 | HIST1H4E | HIST1H4E Entrez, Source | histone cluster 1, H4e | 9219 | 0.004 | -0.3043 | No |
| 60 | NAP1L3 | NAP1L3 Entrez, Source | nucleosome assembly protein 1-like 3 | 9285 | 0.003 | -0.3078 | No |
| 61 | OIP5 | OIP5 Entrez, Source | Opa interacting protein 5 | 9304 | 0.003 | -0.3086 | No |
| 62 | HJURP | 9449 | 0.001 | -0.3166 | No | ||
| 63 | BAHCC1 | BAHCC1 Entrez, Source | BAH domain and coiled-coil containing 1 | 9539 | -0.000 | -0.3215 | No |
| 64 | ARID2 | ARID2 Entrez, Source | AT rich interactive domain 2 (ARID, RFX-like) | 9546 | -0.001 | -0.3218 | No |
| 65 | RBBP7 | RBBP7 Entrez, Source | retinoblastoma binding protein 7 | 9559 | -0.001 | -0.3225 | No |
| 66 | SMARCE1 | SMARCE1 Entrez, Source | SWI/SNF related, matrix associated, actin dependent regulator of chromatin, subfamily e, member 1 | 9683 | -0.002 | -0.3292 | No |
| 67 | MCM2 | MCM2 Entrez, Source | MCM2 minichromosome maintenance deficient 2, mitotin (S. cerevisiae) | 9711 | -0.003 | -0.3305 | No |
| 68 | HMGA1 | HMGA1 Entrez, Source | high mobility group AT-hook 1 | 9756 | -0.004 | -0.3328 | No |
| 69 | H1FX | H1FX Entrez, Source | H1 histone family, member X | 9780 | -0.004 | -0.3339 | No |
| 70 | H2AFX | H2AFX Entrez, Source | H2A histone family, member X | 9986 | -0.007 | -0.3449 | No |
| 71 | RUVBL1 | RUVBL1 Entrez, Source | RuvB-like 1 (E. coli) | 10022 | -0.008 | -0.3464 | No |
| 72 | NOC2L | NOC2L Entrez, Source | nucleolar complex associated 2 homolog (S. cerevisiae) | 10143 | -0.010 | -0.3526 | No |
| 73 | HIRA | HIRA Entrez, Source | HIR histone cell cycle regulation defective homolog A (S. cerevisiae) | 10174 | -0.010 | -0.3537 | No |
| 74 | HELLS | HELLS Entrez, Source | helicase, lymphoid-specific | 10258 | -0.012 | -0.3577 | No |
| 75 | NPM1 | NPM1 Entrez, Source | nucleophosmin (nucleolar phosphoprotein B23, numatrin) | 10260 | -0.012 | -0.3571 | No |
| 76 | CENPP | CENPP Entrez, Source | centromere protein P | 10282 | -0.012 | -0.3576 | No |
| 77 | MIS18A | 10407 | -0.014 | -0.3638 | No | ||
| 78 | SUPT16H | SUPT16H Entrez, Source | suppressor of Ty 16 homolog (S. cerevisiae) | 10711 | -0.019 | -0.3796 | No |
| 79 | NAP1L2 | NAP1L2 Entrez, Source | nucleosome assembly protein 1-like 2 | 10720 | -0.019 | -0.3790 | No |
| 80 | SMARCD1 | SMARCD1 Entrez, Source | SWI/SNF related, matrix associated, actin dependent regulator of chromatin, subfamily d, member 1 | 10741 | -0.020 | -0.3791 | No |
| 81 | CENPM | CENPM Entrez, Source | centromere protein M | 11063 | -0.025 | -0.3956 | No |
| 82 | TSPYL4 | TSPYL4 Entrez, Source | TSPY-like 4 | 11345 | -0.031 | -0.4095 | No |
| 83 | SMYD3 | SMYD3 Entrez, Source | SET and MYND domain containing 3 | 11516 | -0.034 | -0.4172 | No |
| 84 | RSF1 | RSF1 Entrez, Source | remodeling and spacing factor 1 | 11556 | -0.035 | -0.4175 | No |
| 85 | KAT6B | 11619 | -0.036 | -0.4190 | No | ||
| 86 | NAP1L6 | NAP1L6 Entrez, Source | nucleosome assembly protein 1-like 6 | 11957 | -0.043 | -0.4355 | No |
| 87 | HIST1H4H | HIST1H4H Entrez, Source | histone cluster 1, H4h | 12125 | -0.047 | -0.4422 | No |
| 88 | CENPA | CENPA Entrez, Source | centromere protein A | 12307 | -0.052 | -0.4495 | No |
| 89 | NAA60 | 12417 | -0.054 | -0.4527 | No | ||
| 90 | TP53 | TP53 Entrez, Source | tumor protein p53 (Li-Fraumeni syndrome) | 12574 | -0.058 | -0.4582 | No |
| 91 | CENPK | CENPK Entrez, Source | centromere protein K | 13164 | -0.077 | -0.4868 | No |
| 92 | ANP32B | ANP32B Entrez, Source | acidic (leucine-rich) nuclear phosphoprotein 32 family, member B | 13522 | -0.092 | -0.5017 | No |
| 93 | HIST1H2BH | HIST1H2BH Entrez, Source | histone cluster 1, H2bh | 13597 | -0.094 | -0.5008 | No |
| 94 | HIST1H2BE | HIST1H2BE Entrez, Source | histone cluster 1, H2be | 13611 | -0.095 | -0.4964 | No |
| 95 | HIRIP3 | HIRIP3 Entrez, Source | HIRA interacting protein 3 | 13709 | -0.100 | -0.4965 | No |
| 96 | TNRC18 | TNRC18 Entrez, Source | trinucleotide repeat containing 18 | 13986 | -0.113 | -0.5057 | No |
| 97 | ASF1A | ASF1A Entrez, Source | ASF1 anti-silencing function 1 homolog A (S. cerevisiae) | 14150 | -0.123 | -0.5082 | No |
| 98 | NAP1L5 | NAP1L5 Entrez, Source | nucleosome assembly protein 1-like 5 | 14280 | -0.129 | -0.5084 | No |
| 99 | HIST1H2BG | HIST1H2BG Entrez, Source | histone cluster 1, H2bg | 15373 | -0.209 | -0.5579 | No |
| 100 | HIST1H1C | HIST1H1C Entrez, Source | histone cluster 1, H1c | 15656 | -0.236 | -0.5610 | Yes |
| 101 | HIST1H2BC | HIST1H2BC Entrez, Source | histone cluster 1, H2bc | 15812 | -0.254 | -0.5560 | Yes |
| 102 | TSPY26P | 16074 | -0.284 | -0.5553 | Yes | ||
| 103 | CDKN2A | CDKN2A Entrez, Source | cyclin-dependent kinase inhibitor 2A (melanoma, p16, inhibits CDK4) | 16106 | -0.287 | -0.5416 | Yes |
| 104 | HIST2H2BE | HIST2H2BE Entrez, Source | histone cluster 2, H2be | 16216 | -0.303 | -0.5314 | Yes |
| 105 | SOX9 | SOX9 Entrez, Source | SRY (sex determining region Y)-box 9 (campomelic dysplasia, autosomal sex-reversal) | 16540 | -0.353 | -0.5304 | Yes |
| 106 | TSPYL5 | TSPYL5 Entrez, Source | TSPY-like 5 | 16662 | -0.377 | -0.5169 | Yes |
| 107 | H1F0 | H1F0 Entrez, Source | H1 histone family, member 0 | 16674 | -0.380 | -0.4972 | Yes |
| 108 | HIST4H4 | HIST4H4 Entrez, Source | histone cluster 4, H4 | 16754 | -0.396 | -0.4803 | Yes |
| 109 | HIST1H2BD | HIST1H2BD Entrez, Source | histone cluster 1, H2bd | 16826 | -0.413 | -0.4621 | Yes |
| 110 | HIST1H2BK | HIST1H2BK Entrez, Source | histone cluster 1, H2bk | 16932 | -0.436 | -0.4445 | Yes |
| 111 | IPO4 | IPO4 Entrez, Source | importin 4 | 17063 | -0.469 | -0.4266 | Yes |
| 112 | HIST1H3E | HIST1H3E Entrez, Source | histone cluster 1, H3e | 17595 | -0.663 | -0.4205 | Yes |
| 113 | HIST3H2A | HIST3H2A Entrez, Source | histone cluster 3, H2a | 17627 | -0.679 | -0.3858 | Yes |
| 114 | HIST1H3H | HIST1H3H Entrez, Source | histone cluster 1, H3h | 17882 | -0.890 | -0.3522 | Yes |
| 115 | HIST1H3G | HIST1H3G Entrez, Source | histone cluster 1, H3g | 17884 | -0.892 | -0.3044 | Yes |
| 116 | HIST1H4I | HIST1H4I Entrez, Source | histone cluster 1, H4i | 17977 | -1.021 | -0.2547 | Yes |
| 117 | HIST3H2BB | HIST3H2BB Entrez, Source | histone cluster 3, H2bb | 18007 | -1.109 | -0.1968 | Yes |
| 118 | HIST1H2BO | HIST1H2BO Entrez, Source | histone cluster 1, H2bo | 18024 | -1.161 | -0.1354 | Yes |
| 119 | HIST1H2BJ | HIST1H2BJ Entrez, Source | histone cluster 1, H2bj | 18062 | -1.272 | -0.0692 | Yes |
| 120 | HIST1H2BN | HIST1H2BN Entrez, Source | histone cluster 1, H2bn | 18071 | -1.326 | 0.0016 | Yes |