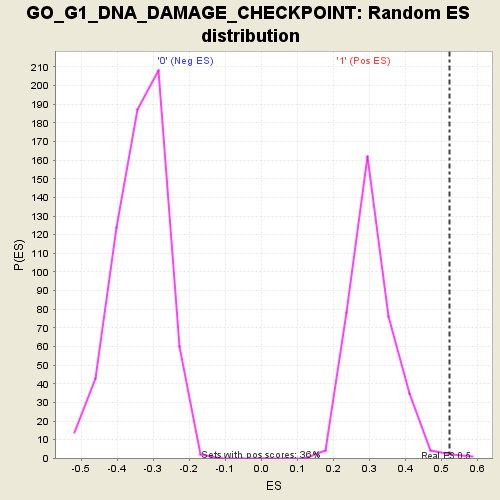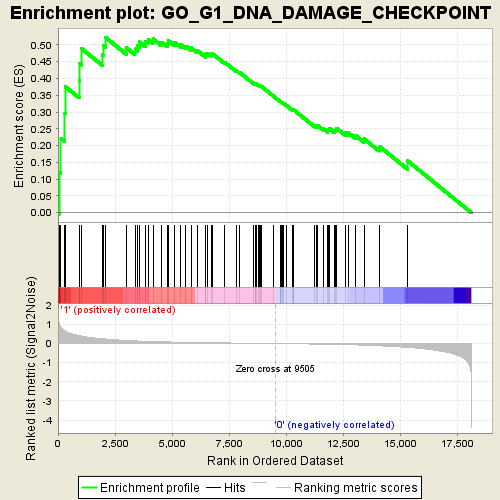
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | NS50719_832_collapsed_to_symbols.NS50719_832.cls #arhinia_versus_control.NS50719_832.cls #arhinia_versus_control_repos |
| Phenotype | NS50719_832.cls#arhinia_versus_control_repos |
| Upregulated in class | 1 |
| GeneSet | GO_G1_DNA_DAMAGE_CHECKPOINT |
| Enrichment Score (ES) | 0.52205116 |
| Normalized Enrichment Score (NES) | 1.6956049 |
| Nominal p-value | 0.002762431 |
| FDR q-value | 0.9237187 |
| FWER p-Value | 1.0 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | TP73 | TP73 Entrez, Source | tumor protein p73 | 70 | 0.995 | 0.1203 | Yes |
| 2 | CDKN1A | CDKN1A Entrez, Source | cyclin-dependent kinase inhibitor 1A (p21, Cip1) | 127 | 0.840 | 0.2219 | Yes |
| 3 | BTG2 | BTG2 Entrez, Source | BTG family, member 2 | 300 | 0.668 | 0.2957 | Yes |
| 4 | SFN | SFN Entrez, Source | stratifin | 327 | 0.649 | 0.3752 | Yes |
| 5 | PLK3 | PLK3 Entrez, Source | polo-like kinase 3 (Drosophila) | 926 | 0.419 | 0.3943 | Yes |
| 6 | CRADD | CRADD Entrez, Source | CASP2 and RIPK1 domain containing adaptor with death domain | 954 | 0.411 | 0.4440 | Yes |
| 7 | PLAGL1 | PLAGL1 Entrez, Source | pleiomorphic adenoma gene-like 1 | 1030 | 0.394 | 0.4891 | Yes |
| 8 | RPS27L | RPS27L Entrez, Source | ribosomal protein S27-like | 1934 | 0.251 | 0.4703 | Yes |
| 9 | ZNF385A | 1978 | 0.246 | 0.4986 | Yes | ||
| 10 | BAX | BAX Entrez, Source | BCL2-associated X protein | 2084 | 0.235 | 0.5221 | Yes |
| 11 | MDM2 | MDM2 Entrez, Source | Mdm2, transformed 3T3 cell double minute 2, p53 binding protein (mouse) | 2982 | 0.159 | 0.4922 | No |
| 12 | TRIAP1 | TRIAP1 Entrez, Source | TP53 regulated inhibitor of apoptosis 1 | 3370 | 0.138 | 0.4879 | No |
| 13 | FBXO31 | FBXO31 Entrez, Source | F-box protein 31 | 3492 | 0.130 | 0.4974 | No |
| 14 | E2F7 | E2F7 Entrez, Source | E2F transcription factor 7 | 3574 | 0.127 | 0.5088 | No |
| 15 | CASP2 | CASP2 Entrez, Source | caspase 2, apoptosis-related cysteine peptidase (neural precursor cell expressed, developmentally down-regulated 2) | 3829 | 0.116 | 0.5092 | No |
| 16 | PCNA | PCNA Entrez, Source | proliferating cell nuclear antigen | 3939 | 0.112 | 0.5171 | No |
| 17 | GTSE1 | GTSE1 Entrez, Source | G-2 and S-phase expressed 1 | 4159 | 0.104 | 0.5179 | No |
| 18 | CENPJ | CENPJ Entrez, Source | centromere protein J | 4535 | 0.091 | 0.5085 | No |
| 19 | E2F4 | E2F4 Entrez, Source | E2F transcription factor 4, p107/p130-binding | 4804 | 0.084 | 0.5041 | No |
| 20 | RBM38 | RBM38 Entrez, Source | RNA binding motif protein 38 | 4831 | 0.083 | 0.5130 | No |
| 21 | PML | PML Entrez, Source | promyelocytic leukemia | 5087 | 0.076 | 0.5083 | No |
| 22 | CDK1 | 5369 | 0.069 | 0.5014 | No | ||
| 23 | CDK2 | CDK2 Entrez, Source | cyclin-dependent kinase 2 | 5601 | 0.063 | 0.4965 | No |
| 24 | CCNB1 | CCNB1 Entrez, Source | cyclin B1 | 5822 | 0.059 | 0.4916 | No |
| 25 | E2F1 | E2F1 Entrez, Source | E2F transcription factor 1 | 6103 | 0.054 | 0.4828 | No |
| 26 | GIGYF2 | 6467 | 0.046 | 0.4684 | No | ||
| 27 | SOX4 | SOX4 Entrez, Source | SRY (sex determining region Y)-box 4 | 6474 | 0.046 | 0.4739 | No |
| 28 | CNOT6 | CNOT6 Entrez, Source | CCR4-NOT transcription complex, subunit 6 | 6562 | 0.045 | 0.4746 | No |
| 29 | CNOT8 | CNOT8 Entrez, Source | CCR4-NOT transcription complex, subunit 8 | 6711 | 0.042 | 0.4717 | No |
| 30 | MDM4 | MDM4 Entrez, Source | Mdm4, transformed 3T3 cell double minute 4, p53 binding protein (mouse) | 6778 | 0.041 | 0.4731 | No |
| 31 | PRMT1 | PRMT1 Entrez, Source | protein arginine methyltransferase 1 | 7303 | 0.032 | 0.4481 | No |
| 32 | UBB | UBB Entrez, Source | ubiquitin B | 7813 | 0.024 | 0.4229 | No |
| 33 | RFWD3 | RFWD3 Entrez, Source | ring finger and WD repeat domain 3 | 7945 | 0.022 | 0.4184 | No |
| 34 | ARID3A | ARID3A Entrez, Source | AT rich interactive domain 3A (BRIGHT- like) | 8549 | 0.013 | 0.3866 | No |
| 35 | CNOT2 | CNOT2 Entrez, Source | CCR4-NOT transcription complex, subunit 2 | 8652 | 0.012 | 0.3825 | No |
| 36 | CARM1 | CARM1 Entrez, Source | coactivator-associated arginine methyltransferase 1 | 8662 | 0.012 | 0.3835 | No |
| 37 | TNKS1BP1 | TNKS1BP1 Entrez, Source | tankyrase 1 binding protein 1, 182kDa | 8669 | 0.012 | 0.3846 | No |
| 38 | EP300 | EP300 Entrez, Source | E1A binding protein p300 | 8772 | 0.010 | 0.3802 | No |
| 39 | TFDP2 | TFDP2 Entrez, Source | transcription factor Dp-2 (E2F dimerization partner 2) | 8841 | 0.009 | 0.3776 | No |
| 40 | CHEK2 | CHEK2 Entrez, Source | CHK2 checkpoint homolog (S. pombe) | 8860 | 0.009 | 0.3777 | No |
| 41 | PRKDC | PRKDC Entrez, Source | protein kinase, DNA-activated, catalytic polypeptide | 8898 | 0.009 | 0.3767 | No |
| 42 | ATM | ATM Entrez, Source | ataxia telangiectasia mutated (includes complementation groups A, C and D) | 9415 | 0.001 | 0.3483 | No |
| 43 | CNOT1 | CNOT1 Entrez, Source | CCR4-NOT transcription complex, subunit 1 | 9728 | -0.003 | 0.3314 | No |
| 44 | UBA52 | UBA52 Entrez, Source | ubiquitin A-52 residue ribosomal protein fusion product 1 | 9732 | -0.003 | 0.3316 | No |
| 45 | MUC1 | MUC1 Entrez, Source | mucin 1, cell surface associated | 9779 | -0.004 | 0.3295 | No |
| 46 | CNOT3 | CNOT3 Entrez, Source | CCR4-NOT transcription complex, subunit 3 | 9807 | -0.004 | 0.3285 | No |
| 47 | CDKN1B | CDKN1B Entrez, Source | cyclin-dependent kinase inhibitor 1B (p27, Kip1) | 9881 | -0.006 | 0.3252 | No |
| 48 | GADD45A | GADD45A Entrez, Source | growth arrest and DNA-damage-inducible, alpha | 10013 | -0.008 | 0.3189 | No |
| 49 | NPM1 | NPM1 Entrez, Source | nucleophosmin (nucleolar phosphoprotein B23, numatrin) | 10260 | -0.012 | 0.3067 | No |
| 50 | RPS27A | RPS27A Entrez, Source | ribosomal protein S27a | 10308 | -0.013 | 0.3057 | No |
| 51 | DGKZ | DGKZ Entrez, Source | diacylglycerol kinase, zeta 104kDa | 10322 | -0.013 | 0.3065 | No |
| 52 | CNOT7 | CNOT7 Entrez, Source | CCR4-NOT transcription complex, subunit 7 | 11247 | -0.029 | 0.2589 | No |
| 53 | CNOT10 | CNOT10 Entrez, Source | CCR4-NOT transcription complex, subunit 10 | 11331 | -0.030 | 0.2580 | No |
| 54 | UBC | UBC Entrez, Source | ubiquitin C | 11369 | -0.031 | 0.2598 | No |
| 55 | AURKA | AURKA Entrez, Source | aurora kinase A | 11631 | -0.036 | 0.2499 | No |
| 56 | CNOT4 | CNOT4 Entrez, Source | CCR4-NOT transcription complex, subunit 4 | 11813 | -0.040 | 0.2448 | No |
| 57 | RBL2 | RBL2 Entrez, Source | retinoblastoma-like 2 (p130) | 11833 | -0.040 | 0.2488 | No |
| 58 | TFDP1 | TFDP1 Entrez, Source | transcription factor Dp-1 | 11894 | -0.042 | 0.2506 | No |
| 59 | PPP2R5C | PPP2R5C Entrez, Source | protein phosphatase 2, regulatory subunit B (B56), gamma isoform | 12123 | -0.047 | 0.2439 | No |
| 60 | WAC | WAC Entrez, Source | WW domain containing adaptor with coiled-coil | 12165 | -0.048 | 0.2476 | No |
| 61 | CCND1 | CCND1 Entrez, Source | cyclin D1 | 12190 | -0.048 | 0.2523 | No |
| 62 | TP53 | TP53 Entrez, Source | tumor protein p53 (Li-Fraumeni syndrome) | 12574 | -0.058 | 0.2383 | No |
| 63 | RPA2 | RPA2 Entrez, Source | replication protein A2, 32kDa | 12717 | -0.063 | 0.2383 | No |
| 64 | CDC25C | CDC25C Entrez, Source | cell division cycle 25C | 13024 | -0.073 | 0.2304 | No |
| 65 | CNOT6L | CNOT6L Entrez, Source | CCR4-NOT transcription complex, subunit 6-like | 13404 | -0.087 | 0.2202 | No |
| 66 | PLK2 | PLK2 Entrez, Source | polo-like kinase 2 (Drosophila) | 14061 | -0.118 | 0.1985 | No |
| 67 | PCBP4 | PCBP4 Entrez, Source | poly(rC) binding protein 4 | 15313 | -0.203 | 0.1545 | No |