Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | NS50719_832_collapsed_to_symbols.NS50719_832.cls #arhinia_versus_control.NS50719_832.cls #arhinia_versus_control_repos |
| Phenotype | NS50719_832.cls#arhinia_versus_control_repos |
| Upregulated in class | 0 |
| GeneSet | GO_NEGATIVE_REGULATION_OF_PROTEIN_KINASE_B_SIGNALING |
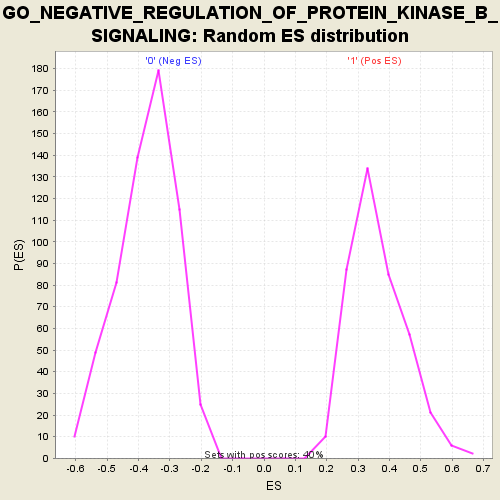
| Enrichment Score (ES) | -0.6874319 |
| Normalized Enrichment Score (NES) | -1.8453623 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.06442266 |
| FWER p-Value | 0.668 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | PHLDA3 | PHLDA3 Entrez, Source | pleckstrin homology-like domain, family A, member 3 | 995 | 0.400 | 0.0125 | No |
| 2 | PLEKHA1 | PLEKHA1 Entrez, Source | pleckstrin homology domain containing, family A (phosphoinositide binding specific) member 1 | 3988 | 0.109 | -0.1346 | No |
| 3 | TSC2 | TSC2 Entrez, Source | tuberous sclerosis 2 | 4860 | 0.082 | -0.1690 | No |
| 4 | ARRB2 | ARRB2 Entrez, Source | arrestin, beta 2 | 5259 | 0.072 | -0.1789 | No |
| 5 | LMBRD1 | LMBRD1 Entrez, Source | LMBR1 domain containing 1 | 5804 | 0.059 | -0.1989 | No |
| 6 | CIB1 | CIB1 Entrez, Source | calcium and integrin binding 1 (calmyrin) | 6739 | 0.042 | -0.2436 | No |
| 7 | LEMD2 | LEMD2 Entrez, Source | LEM domain containing 2 | 6851 | 0.040 | -0.2430 | No |
| 8 | PTEN | PTEN Entrez, Source | phosphatase and tensin homolog (mutated in multiple advanced cancers 1) | 7074 | 0.036 | -0.2492 | No |
| 9 | MTM1 | MTM1 Entrez, Source | myotubularin 1 | 7119 | 0.036 | -0.2456 | No |
| 10 | INPP5K | 7433 | 0.030 | -0.2578 | No | ||
| 11 | HYAL2 | HYAL2 Entrez, Source | hyaluronoglucosaminidase 2 | 7469 | 0.029 | -0.2548 | No |
| 12 | DLG1 | DLG1 Entrez, Source | discs, large homolog 1 (Drosophila) | 8413 | 0.015 | -0.3044 | No |
| 13 | SLC9A3R1 | SLC9A3R1 Entrez, Source | solute carrier family 9 (sodium/hydrogen exchanger), member 3 regulator 1 | 8598 | 0.013 | -0.3124 | No |
| 14 | PTPRJ | PTPRJ Entrez, Source | protein tyrosine phosphatase, receptor type, J | 9244 | 0.003 | -0.3475 | No |
| 15 | EPHA2 | EPHA2 Entrez, Source | EPH receptor A2 | 9547 | -0.001 | -0.3641 | No |
| 16 | MUL1 | 9872 | -0.005 | -0.3811 | No | ||
| 17 | FLCN | FLCN Entrez, Source | folliculin | 9882 | -0.006 | -0.3807 | No |
| 18 | PDCD6 | PDCD6 Entrez, Source | programmed cell death 6 | 11119 | -0.026 | -0.4447 | No |
| 19 | DAG1 | DAG1 Entrez, Source | dystroglycan 1 (dystrophin-associated glycoprotein 1) | 11144 | -0.027 | -0.4415 | No |
| 20 | SIRT1 | SIRT1 Entrez, Source | sirtuin (silent mating type information regulation 2 homolog) 1 (S. cerevisiae) | 11717 | -0.038 | -0.4667 | No |
| 21 | MAGI2 | MAGI2 Entrez, Source | membrane associated guanylate kinase, WW and PDZ domain containing 2 | 12131 | -0.047 | -0.4816 | No |
| 22 | DRD2 | DRD2 Entrez, Source | dopamine receptor D2 | 12539 | -0.057 | -0.4944 | No |
| 23 | PHLPP1 | 12769 | -0.064 | -0.4962 | No | ||
| 24 | RAPGEF1 | RAPGEF1 Entrez, Source | Rap guanine nucleotide exchange factor (GEF) 1 | 13929 | -0.111 | -0.5416 | No |
| 25 | SFRP5 | SFRP5 Entrez, Source | secreted frizzled-related protein 5 | 15239 | -0.197 | -0.5808 | No |
| 26 | XDH | XDH Entrez, Source | xanthine dehydrogenase | 17167 | -0.495 | -0.6038 | Yes |
| 27 | DDIT3 | DDIT3 Entrez, Source | DNA-damage-inducible transcript 3 | 17584 | -0.660 | -0.5154 | Yes |
| 28 | BANK1 | BANK1 Entrez, Source | B-cell scaffold protein with ankyrin repeats 1 | 17982 | -1.032 | -0.3630 | Yes |
| 29 | KLF4 | KLF4 Entrez, Source | Kruppel-like factor 4 (gut) | 17991 | -1.051 | -0.1859 | Yes |
| 30 | PKHD1 | PKHD1 Entrez, Source | polycystic kidney and hepatic disease 1 (autosomal recessive) | 18016 | -1.136 | 0.0046 | Yes |