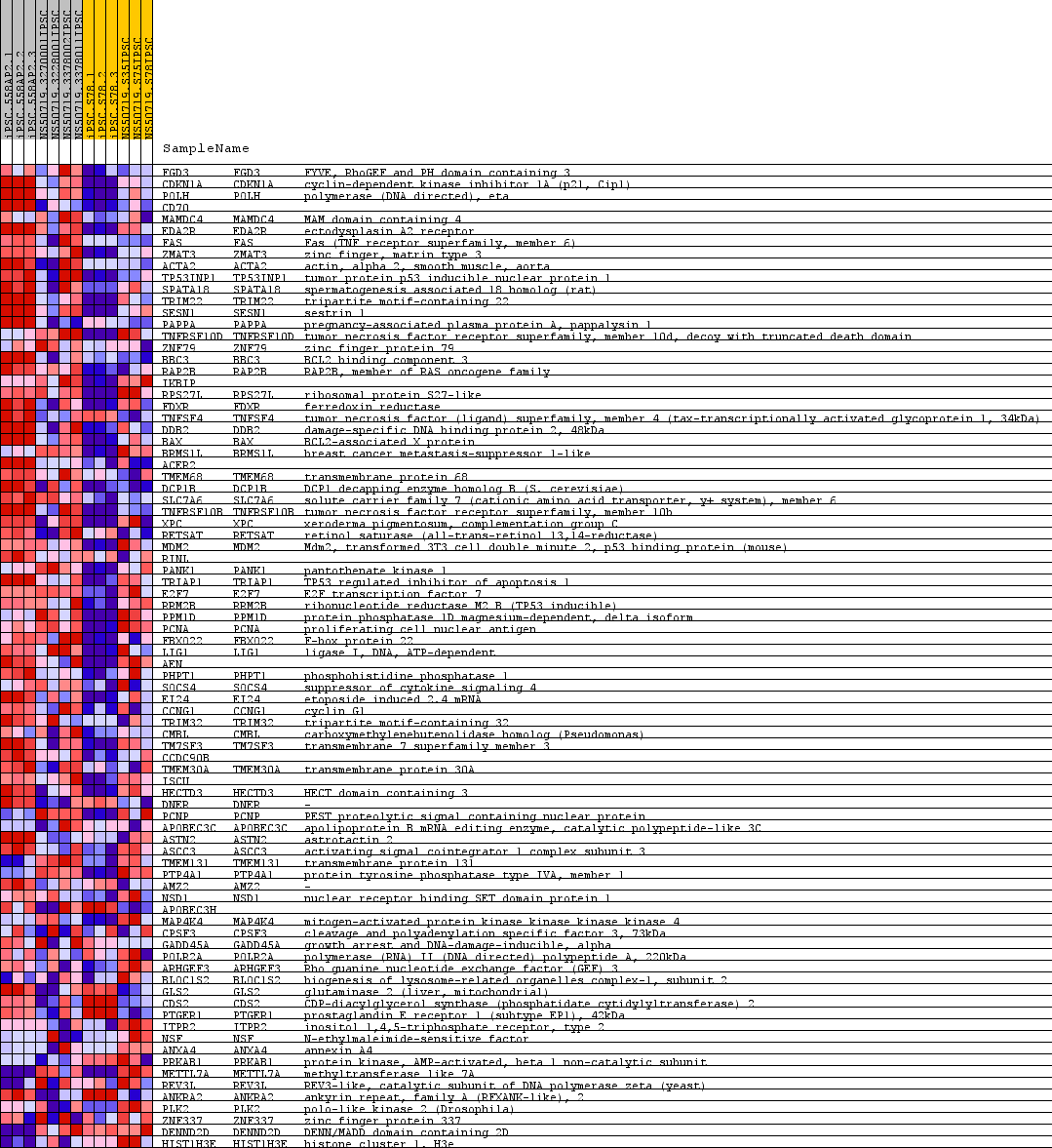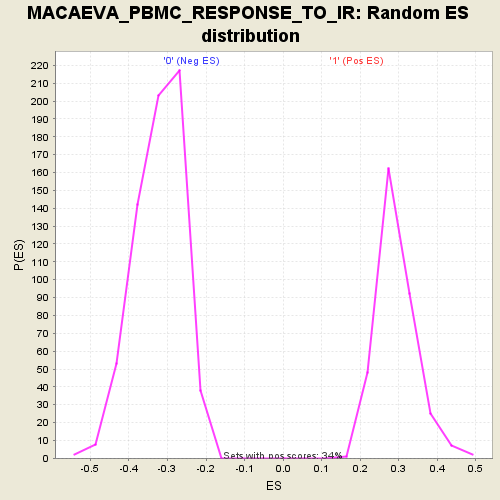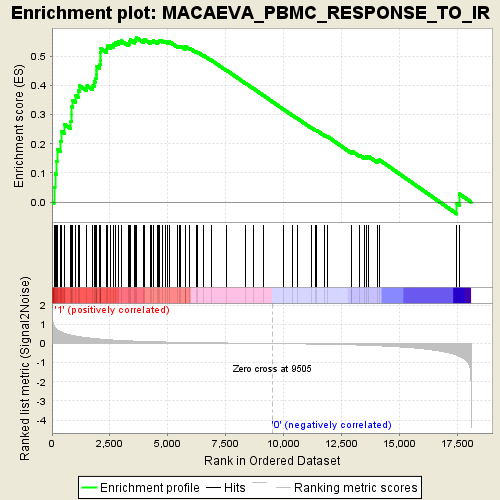
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | NS50719_832_collapsed_to_symbols.NS50719_832.cls #arhinia_versus_control.NS50719_832.cls #arhinia_versus_control_repos |
| Phenotype | NS50719_832.cls#arhinia_versus_control_repos |
| Upregulated in class | 1 |
| GeneSet | MACAEVA_PBMC_RESPONSE_TO_IR |
| Enrichment Score (ES) | 0.56477505 |
| Normalized Enrichment Score (NES) | 1.9362487 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.11609014 |
| FWER p-Value | 0.306 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | FGD3 | FGD3 Entrez, Source | FYVE, RhoGEF and PH domain containing 3 | 84 | 0.949 | 0.0510 | Yes |
| 2 | CDKN1A | CDKN1A Entrez, Source | cyclin-dependent kinase inhibitor 1A (p21, Cip1) | 127 | 0.840 | 0.0980 | Yes |
| 3 | POLH | POLH Entrez, Source | polymerase (DNA directed), eta | 179 | 0.768 | 0.1402 | Yes |
| 4 | CD70 | 216 | 0.737 | 0.1815 | Yes | ||
| 5 | MAMDC4 | MAMDC4 Entrez, Source | MAM domain containing 4 | 367 | 0.618 | 0.2094 | Yes |
| 6 | EDA2R | EDA2R Entrez, Source | ectodysplasin A2 receptor | 402 | 0.597 | 0.2426 | Yes |
| 7 | FAS | FAS Entrez, Source | Fas (TNF receptor superfamily, member 6) | 533 | 0.537 | 0.2668 | Yes |
| 8 | ZMAT3 | ZMAT3 Entrez, Source | zinc finger, matrin type 3 | 814 | 0.447 | 0.2775 | Yes |
| 9 | ACTA2 | ACTA2 Entrez, Source | actin, alpha 2, smooth muscle, aorta | 853 | 0.437 | 0.3010 | Yes |
| 10 | TP53INP1 | TP53INP1 Entrez, Source | tumor protein p53 inducible nuclear protein 1 | 855 | 0.437 | 0.3266 | Yes |
| 11 | SPATA18 | SPATA18 Entrez, Source | spermatogenesis associated 18 homolog (rat) | 897 | 0.425 | 0.3493 | Yes |
| 12 | TRIM22 | TRIM22 Entrez, Source | tripartite motif-containing 22 | 1028 | 0.394 | 0.3652 | Yes |
| 13 | SESN1 | SESN1 Entrez, Source | sestrin 1 | 1131 | 0.371 | 0.3813 | Yes |
| 14 | PAPPA | PAPPA Entrez, Source | pregnancy-associated plasma protein A, pappalysin 1 | 1194 | 0.355 | 0.3987 | Yes |
| 15 | TNFRSF10D | TNFRSF10D Entrez, Source | tumor necrosis factor receptor superfamily, member 10d, decoy with truncated death domain | 1506 | 0.303 | 0.3993 | Yes |
| 16 | ZNF79 | ZNF79 Entrez, Source | zinc finger protein 79 | 1766 | 0.269 | 0.4006 | Yes |
| 17 | BBC3 | BBC3 Entrez, Source | BCL2 binding component 3 | 1818 | 0.264 | 0.4133 | Yes |
| 18 | RAP2B | RAP2B Entrez, Source | RAP2B, member of RAS oncogene family | 1886 | 0.256 | 0.4246 | Yes |
| 19 | IKBIP | 1916 | 0.253 | 0.4378 | Yes | ||
| 20 | RPS27L | RPS27L Entrez, Source | ribosomal protein S27-like | 1934 | 0.251 | 0.4516 | Yes |
| 21 | FDXR | FDXR Entrez, Source | ferredoxin reductase | 1937 | 0.250 | 0.4661 | Yes |
| 22 | TNFSF4 | TNFSF4 Entrez, Source | tumor necrosis factor (ligand) superfamily, member 4 (tax-transcriptionally activated glycoprotein 1, 34kDa) | 2064 | 0.237 | 0.4731 | Yes |
| 23 | DDB2 | DDB2 Entrez, Source | damage-specific DNA binding protein 2, 48kDa | 2079 | 0.235 | 0.4861 | Yes |
| 24 | BAX | BAX Entrez, Source | BCL2-associated X protein | 2084 | 0.235 | 0.4996 | Yes |
| 25 | BRMS1L | BRMS1L Entrez, Source | breast cancer metastasis-suppressor 1-like | 2085 | 0.235 | 0.5134 | Yes |
| 26 | ACER2 | 2108 | 0.232 | 0.5258 | Yes | ||
| 27 | TMEM68 | TMEM68 Entrez, Source | transmembrane protein 68 | 2347 | 0.209 | 0.5249 | Yes |
| 28 | DCP1B | DCP1B Entrez, Source | DCP1 decapping enzyme homolog B (S. cerevisiae) | 2372 | 0.207 | 0.5357 | Yes |
| 29 | SLC7A6 | SLC7A6 Entrez, Source | solute carrier family 7 (cationic amino acid transporter, y+ system), member 6 | 2531 | 0.193 | 0.5382 | Yes |
| 30 | TNFRSF10B | TNFRSF10B Entrez, Source | tumor necrosis factor receptor superfamily, member 10b | 2651 | 0.183 | 0.5424 | Yes |
| 31 | XPC | XPC Entrez, Source | xeroderma pigmentosum, complementation group C | 2729 | 0.178 | 0.5485 | Yes |
| 32 | RETSAT | RETSAT Entrez, Source | retinol saturase (all-trans-retinol 13,14-reductase) | 2880 | 0.166 | 0.5499 | Yes |
| 33 | MDM2 | MDM2 Entrez, Source | Mdm2, transformed 3T3 cell double minute 2, p53 binding protein (mouse) | 2982 | 0.159 | 0.5537 | Yes |
| 34 | RINL | 3295 | 0.141 | 0.5446 | Yes | ||
| 35 | PANK1 | PANK1 Entrez, Source | pantothenate kinase 1 | 3344 | 0.139 | 0.5501 | Yes |
| 36 | TRIAP1 | TRIAP1 Entrez, Source | TP53 regulated inhibitor of apoptosis 1 | 3370 | 0.138 | 0.5568 | Yes |
| 37 | E2F7 | E2F7 Entrez, Source | E2F transcription factor 7 | 3574 | 0.127 | 0.5530 | Yes |
| 38 | RRM2B | RRM2B Entrez, Source | ribonucleotide reductase M2 B (TP53 inducible) | 3596 | 0.126 | 0.5593 | Yes |
| 39 | PPM1D | PPM1D Entrez, Source | protein phosphatase 1D magnesium-dependent, delta isoform | 3630 | 0.125 | 0.5648 | Yes |
| 40 | PCNA | PCNA Entrez, Source | proliferating cell nuclear antigen | 3939 | 0.112 | 0.5542 | No |
| 41 | FBXO22 | FBXO22 Entrez, Source | F-box protein 22 | 4007 | 0.109 | 0.5569 | No |
| 42 | LIG1 | LIG1 Entrez, Source | ligase I, DNA, ATP-dependent | 4238 | 0.101 | 0.5501 | No |
| 43 | AEN | 4305 | 0.099 | 0.5522 | No | ||
| 44 | PHPT1 | PHPT1 Entrez, Source | phosphohistidine phosphatase 1 | 4375 | 0.097 | 0.5541 | No |
| 45 | SOCS4 | SOCS4 Entrez, Source | suppressor of cytokine signaling 4 | 4558 | 0.091 | 0.5493 | No |
| 46 | EI24 | EI24 Entrez, Source | etoposide induced 2.4 mRNA | 4605 | 0.089 | 0.5520 | No |
| 47 | CCNG1 | CCNG1 Entrez, Source | cyclin G1 | 4643 | 0.088 | 0.5551 | No |
| 48 | TRIM32 | TRIM32 Entrez, Source | tripartite motif-containing 32 | 4774 | 0.085 | 0.5528 | No |
| 49 | CMBL | CMBL Entrez, Source | carboxymethylenebutenolidase homolog (Pseudomonas) | 4892 | 0.081 | 0.5511 | No |
| 50 | TM7SF3 | TM7SF3 Entrez, Source | transmembrane 7 superfamily member 3 | 5001 | 0.078 | 0.5497 | No |
| 51 | CCDC90B | 5079 | 0.076 | 0.5499 | No | ||
| 52 | TMEM30A | TMEM30A Entrez, Source | transmembrane protein 30A | 5420 | 0.068 | 0.5350 | No |
| 53 | ISCU | 5487 | 0.066 | 0.5352 | No | ||
| 54 | HECTD3 | HECTD3 Entrez, Source | HECT domain containing 3 | 5558 | 0.064 | 0.5351 | No |
| 55 | DNER | DNER Entrez, Source | - | 5747 | 0.061 | 0.5282 | No |
| 56 | PCNP | PCNP Entrez, Source | PEST proteolytic signal containing nuclear protein | 5765 | 0.060 | 0.5308 | No |
| 57 | APOBEC3C | APOBEC3C Entrez, Source | apolipoprotein B mRNA editing enzyme, catalytic polypeptide-like 3C | 5767 | 0.060 | 0.5343 | No |
| 58 | ASTN2 | ASTN2 Entrez, Source | astrotactin 2 | 5936 | 0.057 | 0.5283 | No |
| 59 | ASCC3 | ASCC3 Entrez, Source | activating signal cointegrator 1 complex subunit 3 | 6221 | 0.051 | 0.5155 | No |
| 60 | TMEM131 | TMEM131 Entrez, Source | transmembrane protein 131 | 6296 | 0.050 | 0.5143 | No |
| 61 | PTP4A1 | PTP4A1 Entrez, Source | protein tyrosine phosphatase type IVA, member 1 | 6557 | 0.045 | 0.5025 | No |
| 62 | AMZ2 | AMZ2 Entrez, Source | - | 6870 | 0.040 | 0.4875 | No |
| 63 | NSD1 | NSD1 Entrez, Source | nuclear receptor binding SET domain protein 1 | 7530 | 0.028 | 0.4526 | No |
| 64 | APOBEC3H | 8346 | 0.016 | 0.4084 | No | ||
| 65 | MAP4K4 | MAP4K4 Entrez, Source | mitogen-activated protein kinase kinase kinase kinase 4 | 8699 | 0.011 | 0.3895 | No |
| 66 | CPSF3 | CPSF3 Entrez, Source | cleavage and polyadenylation specific factor 3, 73kDa | 9145 | 0.005 | 0.3650 | No |
| 67 | GADD45A | GADD45A Entrez, Source | growth arrest and DNA-damage-inducible, alpha | 10013 | -0.008 | 0.3174 | No |
| 68 | POLR2A | POLR2A Entrez, Source | polymerase (RNA) II (DNA directed) polypeptide A, 220kDa | 10365 | -0.014 | 0.2987 | No |
| 69 | ARHGEF3 | ARHGEF3 Entrez, Source | Rho guanine nucleotide exchange factor (GEF) 3 | 10587 | -0.017 | 0.2874 | No |
| 70 | BLOC1S2 | BLOC1S2 Entrez, Source | biogenesis of lysosome-related organelles complex-1, subunit 2 | 11191 | -0.028 | 0.2556 | No |
| 71 | GLS2 | GLS2 Entrez, Source | glutaminase 2 (liver, mitochondrial) | 11380 | -0.031 | 0.2470 | No |
| 72 | CDS2 | CDS2 Entrez, Source | CDP-diacylglycerol synthase (phosphatidate cytidylyltransferase) 2 | 11433 | -0.032 | 0.2460 | No |
| 73 | PTGER1 | PTGER1 Entrez, Source | prostaglandin E receptor 1 (subtype EP1), 42kDa | 11787 | -0.039 | 0.2287 | No |
| 74 | ITPR2 | ITPR2 Entrez, Source | inositol 1,4,5-triphosphate receptor, type 2 | 11889 | -0.042 | 0.2255 | No |
| 75 | NSF | NSF Entrez, Source | N-ethylmaleimide-sensitive factor | 12928 | -0.070 | 0.1720 | No |
| 76 | ANXA4 | ANXA4 Entrez, Source | annexin A4 | 12952 | -0.070 | 0.1748 | No |
| 77 | PRKAB1 | PRKAB1 Entrez, Source | protein kinase, AMP-activated, beta 1 non-catalytic subunit | 13299 | -0.082 | 0.1605 | No |
| 78 | METTL7A | METTL7A Entrez, Source | methyltransferase like 7A | 13516 | -0.092 | 0.1538 | No |
| 79 | REV3L | REV3L Entrez, Source | REV3-like, catalytic subunit of DNA polymerase zeta (yeast) | 13586 | -0.094 | 0.1555 | No |
| 80 | ANKRA2 | ANKRA2 Entrez, Source | ankyrin repeat, family A (RFXANK-like), 2 | 13666 | -0.098 | 0.1569 | No |
| 81 | PLK2 | PLK2 Entrez, Source | polo-like kinase 2 (Drosophila) | 14061 | -0.118 | 0.1419 | No |
| 82 | ZNF337 | ZNF337 Entrez, Source | zinc finger protein 337 | 14142 | -0.122 | 0.1447 | No |
| 83 | DENND2D | DENND2D Entrez, Source | DENN/MADD domain containing 2D | 17489 | -0.613 | -0.0051 | No |
| 84 | HIST1H3E | HIST1H3E Entrez, Source | histone cluster 1, H3e | 17595 | -0.663 | 0.0280 | No |