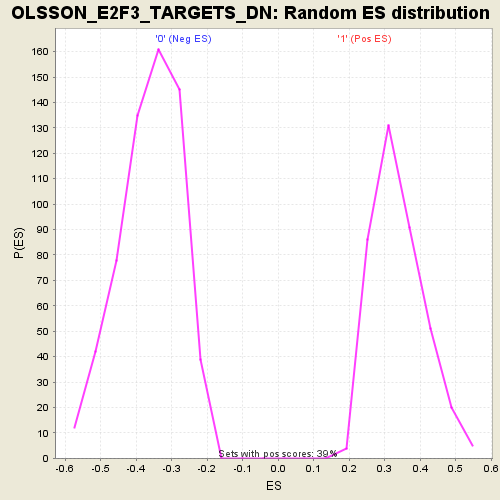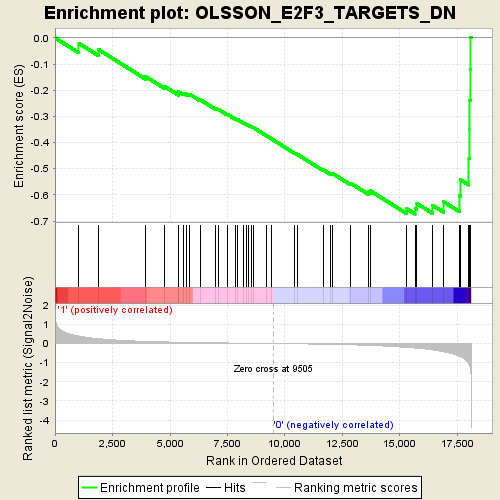
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | NS50719_832_collapsed_to_symbols.NS50719_832.cls #arhinia_versus_control.NS50719_832.cls #arhinia_versus_control_repos |
| Phenotype | NS50719_832.cls#arhinia_versus_control_repos |
| Upregulated in class | 0 |
| GeneSet | OLSSON_E2F3_TARGETS_DN |
| Enrichment Score (ES) | -0.6730337 |
| Normalized Enrichment Score (NES) | -1.866441 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.05841219 |
| FWER p-Value | 0.517 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | TIMP3 | TIMP3 Entrez, Source | TIMP metallopeptidase inhibitor 3 (Sorsby fundus dystrophy, pseudoinflammatory) | 1039 | 0.392 | -0.0211 | No |
| 2 | TAF6L | TAF6L Entrez, Source | TAF6-like RNA polymerase II, p300/CBP-associated factor (PCAF)-associated factor, 65kDa | 1896 | 0.254 | -0.0448 | No |
| 3 | PCNA | PCNA Entrez, Source | proliferating cell nuclear antigen | 3939 | 0.112 | -0.1475 | No |
| 4 | PTMA | PTMA Entrez, Source | prothymosin, alpha (gene sequence 28) | 4778 | 0.085 | -0.1861 | No |
| 5 | MAD2L1 | MAD2L1 Entrez, Source | MAD2 mitotic arrest deficient-like 1 (yeast) | 5368 | 0.069 | -0.2123 | No |
| 6 | CDK1 | 5369 | 0.069 | -0.2058 | No | ||
| 7 | IGF2 | IGF2 Entrez, Source | insulin-like growth factor 2 (somatomedin A) | 5591 | 0.064 | -0.2121 | No |
| 8 | MAGED1 | MAGED1 Entrez, Source | melanoma antigen family D, 1 | 5728 | 0.061 | -0.2140 | No |
| 9 | LBR | LBR Entrez, Source | lamin B receptor | 5854 | 0.058 | -0.2155 | No |
| 10 | PTTG1 | PTTG1 Entrez, Source | pituitary tumor-transforming 1 | 6303 | 0.049 | -0.2357 | No |
| 11 | CCNA2 | CCNA2 Entrez, Source | cyclin A2 | 6986 | 0.037 | -0.2700 | No |
| 12 | UBE2C | UBE2C Entrez, Source | ubiquitin-conjugating enzyme E2C | 7087 | 0.036 | -0.2721 | No |
| 13 | RRM2 | RRM2 Entrez, Source | ribonucleotide reductase M2 polypeptide | 7479 | 0.029 | -0.2911 | No |
| 14 | USP32 | USP32 Entrez, Source | ubiquitin specific peptidase 32 | 7844 | 0.024 | -0.3090 | No |
| 15 | E2F3 | E2F3 Entrez, Source | E2F transcription factor 3 | 7940 | 0.022 | -0.3122 | No |
| 16 | HMGB2 | HMGB2 Entrez, Source | high-mobility group box 2 | 8213 | 0.018 | -0.3256 | No |
| 17 | FST | FST Entrez, Source | follistatin | 8325 | 0.017 | -0.3302 | No |
| 18 | CD9 | CD9 Entrez, Source | CD9 molecule | 8431 | 0.015 | -0.3346 | No |
| 19 | NUCKS1 | NUCKS1 Entrez, Source | nuclear casein kinase and cyclin-dependent kinase substrate 1 | 8527 | 0.014 | -0.3386 | No |
| 20 | TMPO | TMPO Entrez, Source | thymopoietin | 8616 | 0.013 | -0.3423 | No |
| 21 | MYBL2 | MYBL2 Entrez, Source | v-myb myeloblastosis viral oncogene homolog (avian)-like 2 | 9194 | 0.004 | -0.3738 | No |
| 22 | BAIAP2L1 | BAIAP2L1 Entrez, Source | BAI1-associated protein 2-like 1 | 9417 | 0.001 | -0.3860 | No |
| 23 | INHBA | INHBA Entrez, Source | inhibin, beta A (activin A, activin AB alpha polypeptide) | 10424 | -0.015 | -0.4404 | No |
| 24 | MAL2 | MAL2 Entrez, Source | mal, T-cell differentiation protein 2 | 10546 | -0.016 | -0.4456 | No |
| 25 | PLK1 | PLK1 Entrez, Source | polo-like kinase 1 (Drosophila) | 11673 | -0.037 | -0.5045 | No |
| 26 | TPX2 | TPX2 Entrez, Source | TPX2, microtubule-associated, homolog (Xenopus laevis) | 12000 | -0.044 | -0.5184 | No |
| 27 | NFKBIA | NFKBIA Entrez, Source | nuclear factor of kappa light polypeptide gene enhancer in B-cells inhibitor, alpha | 12062 | -0.046 | -0.5175 | No |
| 28 | VIM | VIM Entrez, Source | vimentin | 12844 | -0.067 | -0.5545 | No |
| 29 | LPL | LPL Entrez, Source | lipoprotein lipase | 13622 | -0.096 | -0.5886 | No |
| 30 | THBS1 | THBS1 Entrez, Source | thrombospondin 1 | 13713 | -0.100 | -0.5843 | No |
| 31 | EDN1 | EDN1 Entrez, Source | endothelin 1 | 15301 | -0.202 | -0.6534 | Yes |
| 32 | HIST1H1C | HIST1H1C Entrez, Source | histone cluster 1, H1c | 15656 | -0.236 | -0.6511 | Yes |
| 33 | CAV2 | CAV2 Entrez, Source | caveolin 2 | 15740 | -0.244 | -0.6330 | Yes |
| 34 | THBD | THBD Entrez, Source | thrombomodulin | 16424 | -0.334 | -0.6398 | Yes |
| 35 | MT1H | MT1H Entrez, Source | metallothionein 1H | 16899 | -0.430 | -0.6261 | Yes |
| 36 | HIST1H3E | HIST1H3E Entrez, Source | histone cluster 1, H3e | 17595 | -0.663 | -0.6029 | Yes |
| 37 | HIST3H2A | HIST3H2A Entrez, Source | histone cluster 3, H2a | 17627 | -0.679 | -0.5415 | Yes |
| 38 | CXCL1 | CXCL1 Entrez, Source | chemokine (C-X-C motif) ligand 1 (melanoma growth stimulating activity, alpha) | 18003 | -1.099 | -0.4600 | Yes |
| 39 | MT1F | MT1F Entrez, Source | metallothionein 1F (functional) | 18040 | -1.200 | -0.3504 | Yes |
| 40 | MT1X | MT1X Entrez, Source | metallothionein 1X | 18042 | -1.203 | -0.2386 | Yes |
| 41 | HIST1H2BJ | HIST1H2BJ Entrez, Source | histone cluster 1, H2bj | 18062 | -1.272 | -0.1214 | Yes |
| 42 | HIST1H2BN | HIST1H2BN Entrez, Source | histone cluster 1, H2bn | 18071 | -1.326 | 0.0016 | Yes |