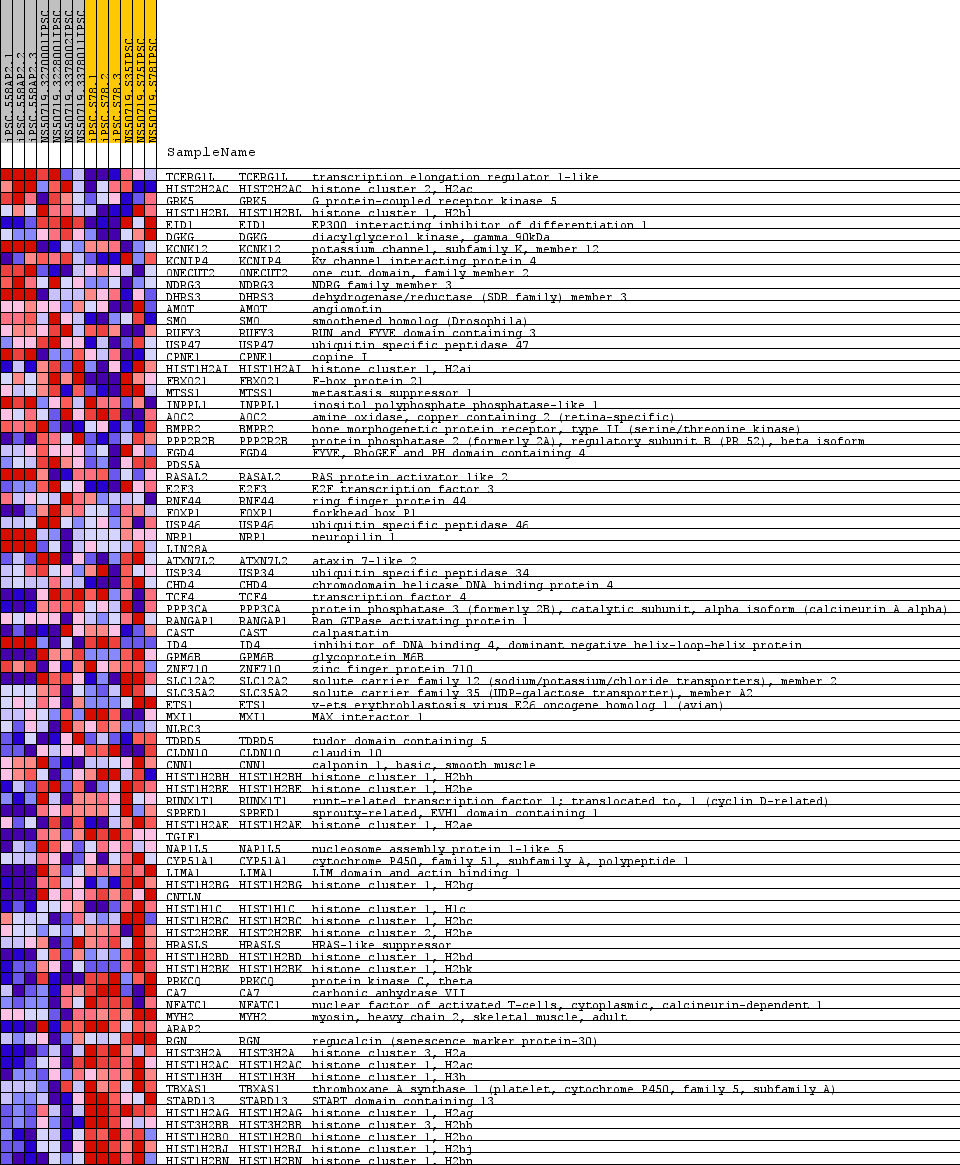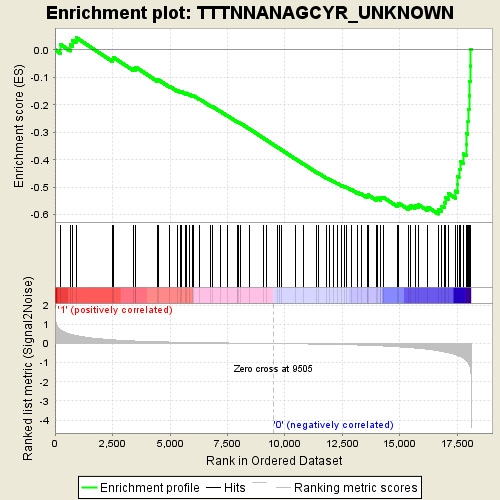
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | NS50719_832_collapsed_to_symbols.NS50719_832.cls #arhinia_versus_control.NS50719_832.cls #arhinia_versus_control_repos |
| Phenotype | NS50719_832.cls#arhinia_versus_control_repos |
| Upregulated in class | 0 |
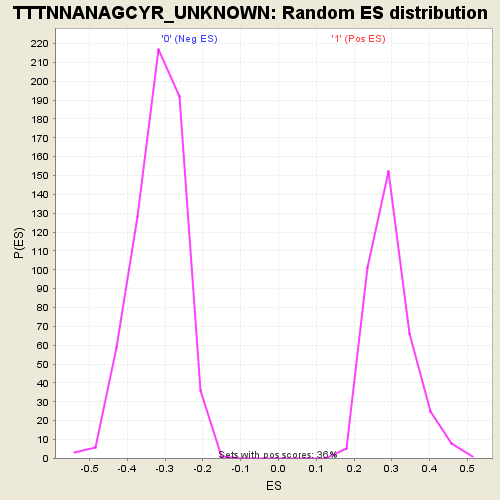
| GeneSet | TTTNNANAGCYR_UNKNOWN |
| Enrichment Score (ES) | -0.5974233 |
| Normalized Enrichment Score (NES) | -1.871911 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.05720628 |
| FWER p-Value | 0.486 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | TCERG1L | TCERG1L Entrez, Source | transcription elongation regulator 1-like | 245 | 0.712 | 0.0190 | No |
| 2 | HIST2H2AC | HIST2H2AC Entrez, Source | histone cluster 2, H2ac | 664 | 0.491 | 0.0183 | No |
| 3 | GRK5 | GRK5 Entrez, Source | G protein-coupled receptor kinase 5 | 757 | 0.464 | 0.0344 | No |
| 4 | HIST1H2BL | HIST1H2BL Entrez, Source | histone cluster 1, H2bl | 914 | 0.421 | 0.0450 | No |
| 5 | EID1 | EID1 Entrez, Source | EP300 interacting inhibitor of differentiation 1 | 2512 | 0.194 | -0.0347 | No |
| 6 | DGKG | DGKG Entrez, Source | diacylglycerol kinase, gamma 90kDa | 2539 | 0.192 | -0.0273 | No |
| 7 | KCNK12 | KCNK12 Entrez, Source | potassium channel, subfamily K, member 12 | 3403 | 0.136 | -0.0690 | No |
| 8 | KCNIP4 | KCNIP4 Entrez, Source | Kv channel interacting protein 4 | 3487 | 0.131 | -0.0676 | No |
| 9 | ONECUT2 | ONECUT2 Entrez, Source | one cut domain, family member 2 | 3504 | 0.130 | -0.0626 | No |
| 10 | NDRG3 | NDRG3 Entrez, Source | NDRG family member 3 | 4462 | 0.094 | -0.1114 | No |
| 11 | DHRS3 | DHRS3 Entrez, Source | dehydrogenase/reductase (SDR family) member 3 | 4482 | 0.093 | -0.1082 | No |
| 12 | AMOT | AMOT Entrez, Source | angiomotin | 4979 | 0.079 | -0.1321 | No |
| 13 | SMO | SMO Entrez, Source | smoothened homolog (Drosophila) | 5334 | 0.070 | -0.1486 | No |
| 14 | RUFY3 | RUFY3 Entrez, Source | RUN and FYVE domain containing 3 | 5451 | 0.067 | -0.1519 | No |
| 15 | USP47 | USP47 Entrez, Source | ubiquitin specific peptidase 47 | 5477 | 0.066 | -0.1503 | No |
| 16 | CPNE1 | CPNE1 Entrez, Source | copine I | 5693 | 0.062 | -0.1594 | No |
| 17 | HIST1H2AI | HIST1H2AI Entrez, Source | histone cluster 1, H2ai | 5702 | 0.062 | -0.1570 | No |
| 18 | FBXO21 | FBXO21 Entrez, Source | F-box protein 21 | 5863 | 0.058 | -0.1632 | No |
| 19 | MTSS1 | MTSS1 Entrez, Source | metastasis suppressor 1 | 5959 | 0.056 | -0.1659 | No |
| 20 | INPPL1 | INPPL1 Entrez, Source | inositol polyphosphate phosphatase-like 1 | 6013 | 0.055 | -0.1663 | No |
| 21 | AOC2 | AOC2 Entrez, Source | amine oxidase, copper containing 2 (retina-specific) | 6288 | 0.050 | -0.1793 | No |
| 22 | BMPR2 | BMPR2 Entrez, Source | bone morphogenetic protein receptor, type II (serine/threonine kinase) | 6776 | 0.041 | -0.2044 | No |
| 23 | PPP2R2B | PPP2R2B Entrez, Source | protein phosphatase 2 (formerly 2A), regulatory subunit B (PR 52), beta isoform | 6850 | 0.040 | -0.2066 | No |
| 24 | FGD4 | FGD4 Entrez, Source | FYVE, RhoGEF and PH domain containing 4 | 7202 | 0.034 | -0.2246 | No |
| 25 | PDS5A | 7490 | 0.029 | -0.2392 | No | ||
| 26 | RASAL2 | RASAL2 Entrez, Source | RAS protein activator like 2 | 7939 | 0.022 | -0.2630 | No |
| 27 | E2F3 | E2F3 Entrez, Source | E2F transcription factor 3 | 7940 | 0.022 | -0.2620 | No |
| 28 | RNF44 | RNF44 Entrez, Source | ring finger protein 44 | 7999 | 0.022 | -0.2642 | No |
| 29 | FOXP1 | FOXP1 Entrez, Source | forkhead box P1 | 8060 | 0.021 | -0.2666 | No |
| 30 | USP46 | USP46 Entrez, Source | ubiquitin specific peptidase 46 | 8439 | 0.015 | -0.2869 | No |
| 31 | NRP1 | NRP1 Entrez, Source | neuropilin 1 | 9056 | 0.006 | -0.3208 | No |
| 32 | LIN28A | 9178 | 0.004 | -0.3273 | No | ||
| 33 | ATXN7L2 | ATXN7L2 Entrez, Source | ataxin 7-like 2 | 9676 | -0.002 | -0.3548 | No |
| 34 | USP34 | USP34 Entrez, Source | ubiquitin specific peptidase 34 | 9757 | -0.004 | -0.3591 | No |
| 35 | CHD4 | CHD4 Entrez, Source | chromodomain helicase DNA binding protein 4 | 9856 | -0.005 | -0.3643 | No |
| 36 | TCF4 | TCF4 Entrez, Source | transcription factor 4 | 10448 | -0.015 | -0.3964 | No |
| 37 | PPP3CA | PPP3CA Entrez, Source | protein phosphatase 3 (formerly 2B), catalytic subunit, alpha isoform (calcineurin A alpha) | 10812 | -0.021 | -0.4156 | No |
| 38 | RANGAP1 | RANGAP1 Entrez, Source | Ran GTPase activating protein 1 | 11390 | -0.031 | -0.4462 | No |
| 39 | CAST | CAST Entrez, Source | calpastatin | 11452 | -0.033 | -0.4481 | No |
| 40 | ID4 | ID4 Entrez, Source | inhibitor of DNA binding 4, dominant negative helix-loop-helix protein | 11821 | -0.040 | -0.4667 | No |
| 41 | GPM6B | GPM6B Entrez, Source | glycoprotein M6B | 11931 | -0.042 | -0.4708 | No |
| 42 | ZNF710 | ZNF710 Entrez, Source | zinc finger protein 710 | 12110 | -0.047 | -0.4785 | No |
| 43 | SLC12A2 | SLC12A2 Entrez, Source | solute carrier family 12 (sodium/potassium/chloride transporters), member 2 | 12284 | -0.051 | -0.4858 | No |
| 44 | SLC35A2 | SLC35A2 Entrez, Source | solute carrier family 35 (UDP-galactose transporter), member A2 | 12479 | -0.056 | -0.4940 | No |
| 45 | ETS1 | ETS1 Entrez, Source | v-ets erythroblastosis virus E26 oncogene homolog 1 (avian) | 12582 | -0.059 | -0.4970 | No |
| 46 | MXI1 | MXI1 Entrez, Source | MAX interactor 1 | 12685 | -0.062 | -0.4998 | No |
| 47 | NLRC3 | 12891 | -0.069 | -0.5081 | No | ||
| 48 | TDRD5 | TDRD5 Entrez, Source | tudor domain containing 5 | 13173 | -0.078 | -0.5201 | No |
| 49 | CLDN10 | CLDN10 Entrez, Source | claudin 10 | 13314 | -0.083 | -0.5241 | No |
| 50 | CNN1 | CNN1 Entrez, Source | calponin 1, basic, smooth muscle | 13585 | -0.094 | -0.5348 | No |
| 51 | HIST1H2BH | HIST1H2BH Entrez, Source | histone cluster 1, H2bh | 13597 | -0.094 | -0.5311 | No |
| 52 | HIST1H2BE | HIST1H2BE Entrez, Source | histone cluster 1, H2be | 13611 | -0.095 | -0.5274 | No |
| 53 | RUNX1T1 | RUNX1T1 Entrez, Source | runt-related transcription factor 1; translocated to, 1 (cyclin D-related) | 13964 | -0.112 | -0.5418 | No |
| 54 | SPRED1 | SPRED1 Entrez, Source | sprouty-related, EVH1 domain containing 1 | 14009 | -0.115 | -0.5390 | No |
| 55 | HIST1H2AE | HIST1H2AE Entrez, Source | histone cluster 1, H2ae | 14163 | -0.124 | -0.5418 | No |
| 56 | TGIF1 | 14170 | -0.124 | -0.5365 | No | ||
| 57 | NAP1L5 | NAP1L5 Entrez, Source | nucleosome assembly protein 1-like 5 | 14280 | -0.129 | -0.5366 | No |
| 58 | CYP51A1 | CYP51A1 Entrez, Source | cytochrome P450, family 51, subfamily A, polypeptide 1 | 14914 | -0.171 | -0.5639 | No |
| 59 | LIMA1 | LIMA1 Entrez, Source | LIM domain and actin binding 1 | 14940 | -0.173 | -0.5574 | No |
| 60 | HIST1H2BG | HIST1H2BG Entrez, Source | histone cluster 1, H2bg | 15373 | -0.209 | -0.5718 | No |
| 61 | CNTLN | 15476 | -0.217 | -0.5675 | No | ||
| 62 | HIST1H1C | HIST1H1C Entrez, Source | histone cluster 1, H1c | 15656 | -0.236 | -0.5667 | No |
| 63 | HIST1H2BC | HIST1H2BC Entrez, Source | histone cluster 1, H2bc | 15812 | -0.254 | -0.5637 | No |
| 64 | HIST2H2BE | HIST2H2BE Entrez, Source | histone cluster 2, H2be | 16216 | -0.303 | -0.5722 | No |
| 65 | HRASLS | HRASLS Entrez, Source | HRAS-like suppressor | 16672 | -0.379 | -0.5801 | Yes |
| 66 | HIST1H2BD | HIST1H2BD Entrez, Source | histone cluster 1, H2bd | 16826 | -0.413 | -0.5696 | Yes |
| 67 | HIST1H2BK | HIST1H2BK Entrez, Source | histone cluster 1, H2bk | 16932 | -0.436 | -0.5555 | Yes |
| 68 | PRKCQ | PRKCQ Entrez, Source | protein kinase C, theta | 16972 | -0.446 | -0.5372 | Yes |
| 69 | CA7 | CA7 Entrez, Source | carbonic anhydrase VII | 17098 | -0.476 | -0.5223 | Yes |
| 70 | NFATC1 | NFATC1 Entrez, Source | nuclear factor of activated T-cells, cytoplasmic, calcineurin-dependent 1 | 17421 | -0.581 | -0.5136 | Yes |
| 71 | MYH2 | MYH2 Entrez, Source | myosin, heavy chain 2, skeletal muscle, adult | 17512 | -0.628 | -0.4899 | Yes |
| 72 | ARAP2 | 17519 | -0.632 | -0.4613 | Yes | ||
| 73 | RGN | RGN Entrez, Source | regucalcin (senescence marker protein-30) | 17572 | -0.655 | -0.4342 | Yes |
| 74 | HIST3H2A | HIST3H2A Entrez, Source | histone cluster 3, H2a | 17627 | -0.679 | -0.4061 | Yes |
| 75 | HIST1H2AC | HIST1H2AC Entrez, Source | histone cluster 1, H2ac | 17773 | -0.774 | -0.3787 | Yes |
| 76 | HIST1H3H | HIST1H3H Entrez, Source | histone cluster 1, H3h | 17882 | -0.890 | -0.3439 | Yes |
| 77 | TBXAS1 | TBXAS1 Entrez, Source | thromboxane A synthase 1 (platelet, cytochrome P450, family 5, subfamily A) | 17887 | -0.898 | -0.3031 | Yes |
| 78 | STARD13 | STARD13 Entrez, Source | START domain containing 13 | 17957 | -0.990 | -0.2616 | Yes |
| 79 | HIST1H2AG | HIST1H2AG Entrez, Source | histone cluster 1, H2ag | 17970 | -1.012 | -0.2159 | Yes |
| 80 | HIST3H2BB | HIST3H2BB Entrez, Source | histone cluster 3, H2bb | 18007 | -1.109 | -0.1671 | Yes |
| 81 | HIST1H2BO | HIST1H2BO Entrez, Source | histone cluster 1, H2bo | 18024 | -1.161 | -0.1149 | Yes |
| 82 | HIST1H2BJ | HIST1H2BJ Entrez, Source | histone cluster 1, H2bj | 18062 | -1.272 | -0.0587 | Yes |
| 83 | HIST1H2BN | HIST1H2BN Entrez, Source | histone cluster 1, H2bn | 18071 | -1.326 | 0.0016 | Yes |