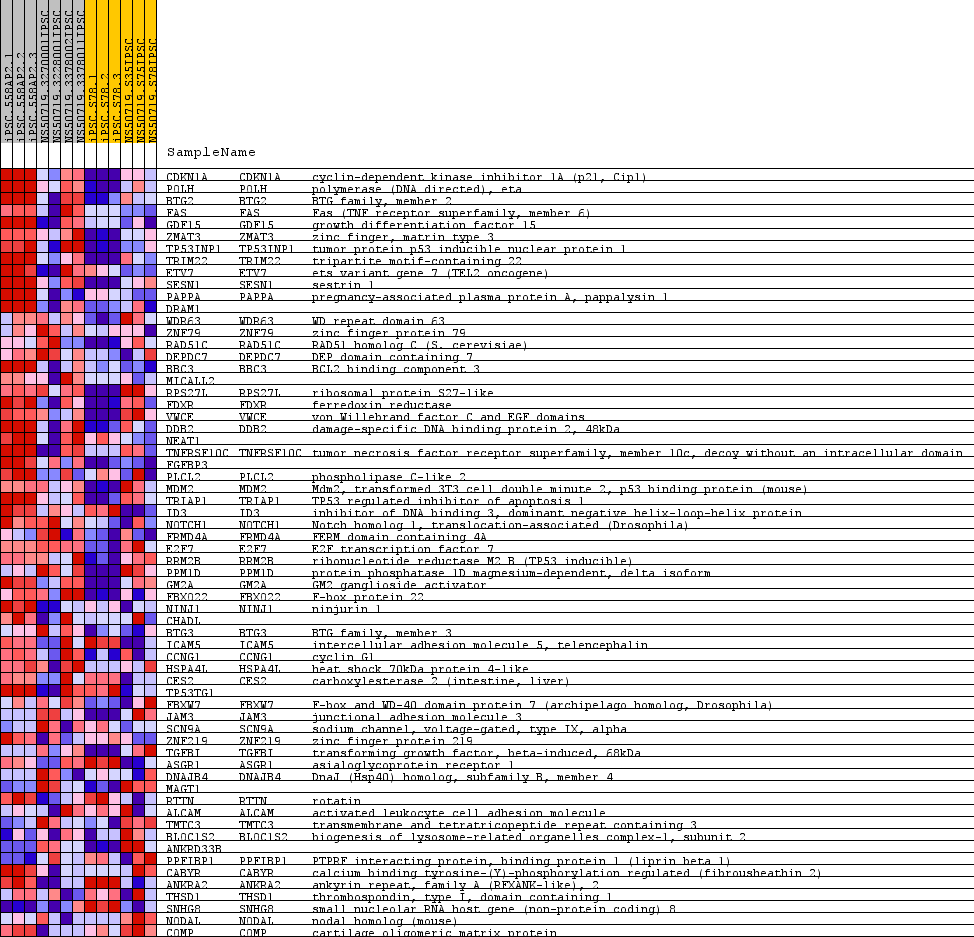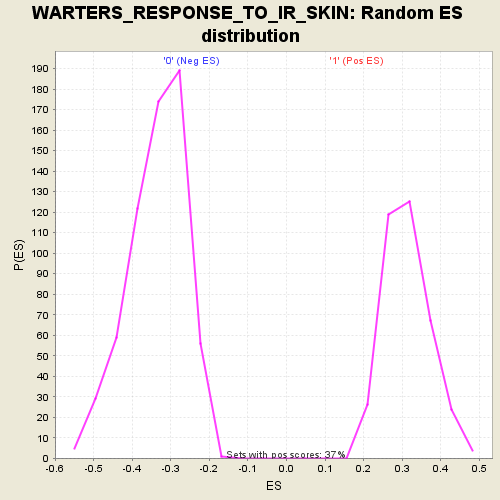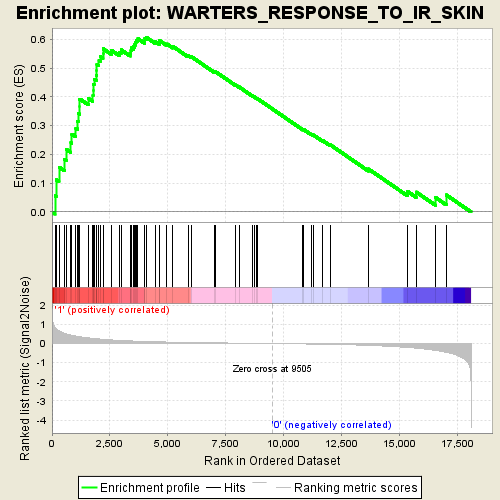
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | NS50719_832_collapsed_to_symbols.NS50719_832.cls #arhinia_versus_control.NS50719_832.cls #arhinia_versus_control_repos |
| Phenotype | NS50719_832.cls#arhinia_versus_control_repos |
| Upregulated in class | 1 |
| GeneSet | WARTERS_RESPONSE_TO_IR_SKIN |
| Enrichment Score (ES) | 0.6066623 |
| Normalized Enrichment Score (NES) | 1.9454496 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.15088971 |
| FWER p-Value | 0.271 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | CDKN1A | CDKN1A Entrez, Source | cyclin-dependent kinase inhibitor 1A (p21, Cip1) | 127 | 0.840 | 0.0565 | Yes |
| 2 | POLH | POLH Entrez, Source | polymerase (DNA directed), eta | 179 | 0.768 | 0.1119 | Yes |
| 3 | BTG2 | BTG2 Entrez, Source | BTG family, member 2 | 300 | 0.668 | 0.1558 | Yes |
| 4 | FAS | FAS Entrez, Source | Fas (TNF receptor superfamily, member 6) | 533 | 0.537 | 0.1835 | Yes |
| 5 | GDF15 | GDF15 Entrez, Source | growth differentiation factor 15 | 612 | 0.507 | 0.2176 | Yes |
| 6 | ZMAT3 | ZMAT3 Entrez, Source | zinc finger, matrin type 3 | 814 | 0.447 | 0.2403 | Yes |
| 7 | TP53INP1 | TP53INP1 Entrez, Source | tumor protein p53 inducible nuclear protein 1 | 855 | 0.437 | 0.2711 | Yes |
| 8 | TRIM22 | TRIM22 Entrez, Source | tripartite motif-containing 22 | 1028 | 0.394 | 0.2914 | Yes |
| 9 | ETV7 | ETV7 Entrez, Source | ets variant gene 7 (TEL2 oncogene) | 1099 | 0.378 | 0.3162 | Yes |
| 10 | SESN1 | SESN1 Entrez, Source | sestrin 1 | 1131 | 0.371 | 0.3425 | Yes |
| 11 | PAPPA | PAPPA Entrez, Source | pregnancy-associated plasma protein A, pappalysin 1 | 1194 | 0.355 | 0.3660 | Yes |
| 12 | DRAM1 | 1196 | 0.354 | 0.3927 | Yes | ||
| 13 | WDR63 | WDR63 Entrez, Source | WD repeat domain 63 | 1565 | 0.295 | 0.3947 | Yes |
| 14 | ZNF79 | ZNF79 Entrez, Source | zinc finger protein 79 | 1766 | 0.269 | 0.4039 | Yes |
| 15 | RAD51C | RAD51C Entrez, Source | RAD51 homolog C (S. cerevisiae) | 1773 | 0.268 | 0.4238 | Yes |
| 16 | DEPDC7 | DEPDC7 Entrez, Source | DEP domain containing 7 | 1776 | 0.267 | 0.4440 | Yes |
| 17 | BBC3 | BBC3 Entrez, Source | BCL2 binding component 3 | 1818 | 0.264 | 0.4617 | Yes |
| 18 | MICALL2 | 1907 | 0.253 | 0.4760 | Yes | ||
| 19 | RPS27L | RPS27L Entrez, Source | ribosomal protein S27-like | 1934 | 0.251 | 0.4935 | Yes |
| 20 | FDXR | FDXR Entrez, Source | ferredoxin reductase | 1937 | 0.250 | 0.5124 | Yes |
| 21 | VWCE | VWCE Entrez, Source | von Willebrand factor C and EGF domains | 2026 | 0.242 | 0.5258 | Yes |
| 22 | DDB2 | DDB2 Entrez, Source | damage-specific DNA binding protein 2, 48kDa | 2079 | 0.235 | 0.5407 | Yes |
| 23 | NEAT1 | 2201 | 0.224 | 0.5509 | Yes | ||
| 24 | TNFRSF10C | TNFRSF10C Entrez, Source | tumor necrosis factor receptor superfamily, member 10c, decoy without an intracellular domain | 2205 | 0.223 | 0.5676 | Yes |
| 25 | FGFBP3 | 2568 | 0.190 | 0.5620 | Yes | ||
| 26 | PLCL2 | PLCL2 Entrez, Source | phospholipase C-like 2 | 2913 | 0.164 | 0.5553 | Yes |
| 27 | MDM2 | MDM2 Entrez, Source | Mdm2, transformed 3T3 cell double minute 2, p53 binding protein (mouse) | 2982 | 0.159 | 0.5636 | Yes |
| 28 | TRIAP1 | TRIAP1 Entrez, Source | TP53 regulated inhibitor of apoptosis 1 | 3370 | 0.138 | 0.5526 | Yes |
| 29 | ID3 | ID3 Entrez, Source | inhibitor of DNA binding 3, dominant negative helix-loop-helix protein | 3407 | 0.135 | 0.5608 | Yes |
| 30 | NOTCH1 | NOTCH1 Entrez, Source | Notch homolog 1, translocation-associated (Drosophila) | 3419 | 0.135 | 0.5704 | Yes |
| 31 | FRMD4A | FRMD4A Entrez, Source | FERM domain containing 4A | 3527 | 0.129 | 0.5743 | Yes |
| 32 | E2F7 | E2F7 Entrez, Source | E2F transcription factor 7 | 3574 | 0.127 | 0.5813 | Yes |
| 33 | RRM2B | RRM2B Entrez, Source | ribonucleotide reductase M2 B (TP53 inducible) | 3596 | 0.126 | 0.5897 | Yes |
| 34 | PPM1D | PPM1D Entrez, Source | protein phosphatase 1D magnesium-dependent, delta isoform | 3630 | 0.125 | 0.5974 | Yes |
| 35 | GM2A | GM2A Entrez, Source | GM2 ganglioside activator | 3710 | 0.121 | 0.6022 | Yes |
| 36 | FBXO22 | FBXO22 Entrez, Source | F-box protein 22 | 4007 | 0.109 | 0.5940 | Yes |
| 37 | NINJ1 | NINJ1 Entrez, Source | ninjurin 1 | 4009 | 0.109 | 0.6022 | Yes |
| 38 | CHADL | 4075 | 0.107 | 0.6067 | Yes | ||
| 39 | BTG3 | BTG3 Entrez, Source | BTG family, member 3 | 4472 | 0.094 | 0.5918 | No |
| 40 | ICAM5 | ICAM5 Entrez, Source | intercellular adhesion molecule 5, telencephalin | 4637 | 0.088 | 0.5894 | No |
| 41 | CCNG1 | CCNG1 Entrez, Source | cyclin G1 | 4643 | 0.088 | 0.5958 | No |
| 42 | HSPA4L | HSPA4L Entrez, Source | heat shock 70kDa protein 4-like | 4963 | 0.079 | 0.5841 | No |
| 43 | CES2 | CES2 Entrez, Source | carboxylesterase 2 (intestine, liver) | 5219 | 0.073 | 0.5754 | No |
| 44 | TP53TG1 | 5880 | 0.058 | 0.5432 | No | ||
| 45 | FBXW7 | FBXW7 Entrez, Source | F-box and WD-40 domain protein 7 (archipelago homolog, Drosophila) | 6007 | 0.056 | 0.5405 | No |
| 46 | JAM3 | JAM3 Entrez, Source | junctional adhesion molecule 3 | 7027 | 0.037 | 0.4867 | No |
| 47 | SCN9A | SCN9A Entrez, Source | sodium channel, voltage-gated, type IX, alpha | 7057 | 0.036 | 0.4879 | No |
| 48 | ZNF219 | ZNF219 Entrez, Source | zinc finger protein 219 | 7921 | 0.022 | 0.4417 | No |
| 49 | TGFBI | TGFBI Entrez, Source | transforming growth factor, beta-induced, 68kDa | 8075 | 0.020 | 0.4348 | No |
| 50 | ASGR1 | ASGR1 Entrez, Source | asialoglycoprotein receptor 1 | 8636 | 0.012 | 0.4047 | No |
| 51 | DNAJB4 | DNAJB4 Entrez, Source | DnaJ (Hsp40) homolog, subfamily B, member 4 | 8729 | 0.011 | 0.4004 | No |
| 52 | MAGT1 | 8832 | 0.009 | 0.3954 | No | ||
| 53 | RTTN | RTTN Entrez, Source | rotatin | 8885 | 0.009 | 0.3932 | No |
| 54 | ALCAM | ALCAM Entrez, Source | activated leukocyte cell adhesion molecule | 10839 | -0.021 | 0.2865 | No |
| 55 | TMTC3 | TMTC3 Entrez, Source | transmembrane and tetratricopeptide repeat containing 3 | 10845 | -0.021 | 0.2879 | No |
| 56 | BLOC1S2 | BLOC1S2 Entrez, Source | biogenesis of lysosome-related organelles complex-1, subunit 2 | 11191 | -0.028 | 0.2708 | No |
| 57 | ANKRD33B | 11285 | -0.029 | 0.2679 | No | ||
| 58 | PPFIBP1 | PPFIBP1 Entrez, Source | PTPRF interacting protein, binding protein 1 (liprin beta 1) | 11703 | -0.038 | 0.2476 | No |
| 59 | CABYR | CABYR Entrez, Source | calcium binding tyrosine-(Y)-phosphorylation regulated (fibrousheathin 2) | 12012 | -0.044 | 0.2339 | No |
| 60 | ANKRA2 | ANKRA2 Entrez, Source | ankyrin repeat, family A (RFXANK-like), 2 | 13666 | -0.098 | 0.1497 | No |
| 61 | THSD1 | THSD1 Entrez, Source | thrombospondin, type I, domain containing 1 | 15351 | -0.206 | 0.0719 | No |
| 62 | SNHG8 | SNHG8 Entrez, Source | small nucleolar RNA host gene (non-protein coding) 8 | 15727 | -0.243 | 0.0695 | No |
| 63 | NODAL | NODAL Entrez, Source | nodal homolog (mouse) | 16563 | -0.358 | 0.0503 | No |
| 64 | COMP | COMP Entrez, Source | cartilage oligomeric matrix protein | 17029 | -0.459 | 0.0593 | No |