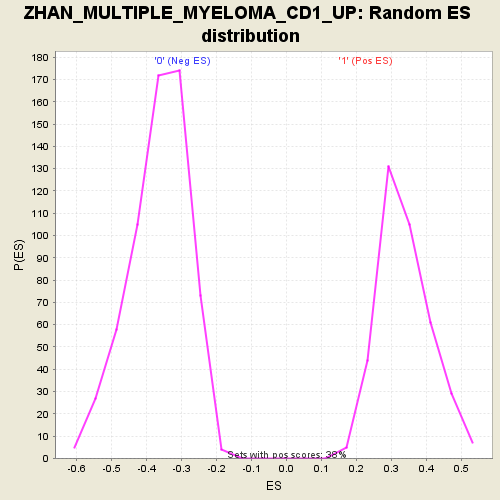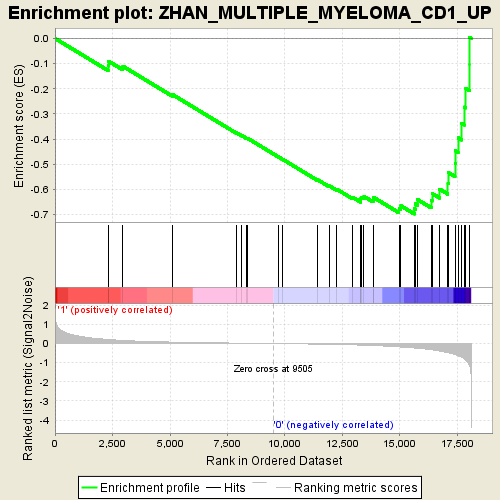
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | NS50719_832_collapsed_to_symbols.NS50719_832.cls #arhinia_versus_control.NS50719_832.cls #arhinia_versus_control_repos |
| Phenotype | NS50719_832.cls#arhinia_versus_control_repos |
| Upregulated in class | 0 |
| GeneSet | ZHAN_MULTIPLE_MYELOMA_CD1_UP |
| Enrichment Score (ES) | -0.69777685 |
| Normalized Enrichment Score (NES) | -1.9056861 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.04385159 |
| FWER p-Value | 0.284 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | SYTL1 | SYTL1 Entrez, Source | synaptotagmin-like 1 | 2309 | 0.212 | -0.1087 | No |
| 2 | RBP1 | RBP1 Entrez, Source | retinol binding protein 1, cellular | 2326 | 0.210 | -0.0905 | No |
| 3 | MTG1 | MTG1 Entrez, Source | mitochondrial GTPase 1 homolog (S. cerevisiae) | 2950 | 0.161 | -0.1104 | No |
| 4 | GPR63 | GPR63 Entrez, Source | G protein-coupled receptor 63 | 5100 | 0.075 | -0.2226 | No |
| 5 | RAB33A | RAB33A Entrez, Source | RAB33A, member RAS oncogene family | 7891 | 0.023 | -0.3750 | No |
| 6 | TMC8 | TMC8 Entrez, Source | transmembrane channel-like 8 | 8123 | 0.020 | -0.3860 | No |
| 7 | TMC6 | TMC6 Entrez, Source | transmembrane channel-like 6 | 8342 | 0.017 | -0.3966 | No |
| 8 | NID2 | NID2 Entrez, Source | nidogen 2 (osteonidogen) | 8359 | 0.016 | -0.3960 | No |
| 9 | TBC1D16 | TBC1D16 Entrez, Source | TBC1 domain family, member 16 | 9700 | -0.003 | -0.4699 | No |
| 10 | HADH | HADH Entrez, Source | hydroxyacyl-Coenzyme A dehydrogenase | 9901 | -0.006 | -0.4805 | No |
| 11 | PPP2R5E | PPP2R5E Entrez, Source | protein phosphatase 2, regulatory subunit B (B56), epsilon isoform | 11429 | -0.032 | -0.5621 | No |
| 12 | VPS37A | VPS37A Entrez, Source | vacuolar protein sorting 37 homolog A (S. cerevisiae) | 11933 | -0.043 | -0.5861 | No |
| 13 | PHGDH | PHGDH Entrez, Source | phosphoglycerate dehydrogenase | 12236 | -0.050 | -0.5984 | No |
| 14 | RAB31 | RAB31 Entrez, Source | RAB31, member RAS oncogene family | 12936 | -0.070 | -0.6307 | No |
| 15 | TOX2 | 13293 | -0.082 | -0.6430 | No | ||
| 16 | FYN | FYN Entrez, Source | FYN oncogene related to SRC, FGR, YES | 13300 | -0.082 | -0.6359 | No |
| 17 | KLHL4 | KLHL4 Entrez, Source | kelch-like 4 (Drosophila) | 13334 | -0.083 | -0.6302 | No |
| 18 | ZNF70 | ZNF70 Entrez, Source | zinc finger protein 70 | 13434 | -0.088 | -0.6277 | No |
| 19 | NSUN7 | NSUN7 Entrez, Source | NOL1/NOP2/Sun domain family, member 7 | 13834 | -0.105 | -0.6403 | No |
| 20 | EIF4EBP1 | EIF4EBP1 Entrez, Source | eukaryotic translation initiation factor 4E binding protein 1 | 13861 | -0.107 | -0.6321 | No |
| 21 | PTGER2 | PTGER2 Entrez, Source | prostaglandin E receptor 2 (subtype EP2), 53kDa | 14959 | -0.174 | -0.6771 | Yes |
| 22 | CLEC2D | CLEC2D Entrez, Source | C-type lectin domain family 2, member D | 15043 | -0.181 | -0.6653 | Yes |
| 23 | STC2 | STC2 Entrez, Source | stanniocalcin 2 | 15630 | -0.233 | -0.6767 | Yes |
| 24 | GPT2 | GPT2 Entrez, Source | glutamic pyruvate transaminase (alanine aminotransferase) 2 | 15671 | -0.237 | -0.6575 | Yes |
| 25 | SESN2 | SESN2 Entrez, Source | sestrin 2 | 15765 | -0.247 | -0.6403 | Yes |
| 26 | PCK2 | PCK2 Entrez, Source | phosphoenolpyruvate carboxykinase 2 (mitochondrial) | 16361 | -0.324 | -0.6439 | Yes |
| 27 | CTH | CTH Entrez, Source | cystathionase (cystathionine gamma-lyase) | 16401 | -0.331 | -0.6162 | Yes |
| 28 | CEBPB | CEBPB Entrez, Source | CCAAT/enhancer binding protein (C/EBP), beta | 16730 | -0.392 | -0.5990 | Yes |
| 29 | SLC7A11 | SLC7A11 Entrez, Source | solute carrier family 7, (cationic amino acid transporter, y+ system) member 11 | 17090 | -0.475 | -0.5759 | Yes |
| 30 | MFAP4 | MFAP4 Entrez, Source | microfibrillar-associated protein 4 | 17119 | -0.483 | -0.5339 | Yes |
| 31 | TM6SF1 | TM6SF1 Entrez, Source | transmembrane 6 superfamily member 1 | 17397 | -0.571 | -0.4976 | Yes |
| 32 | INHBE | INHBE Entrez, Source | inhibin, beta E | 17417 | -0.579 | -0.4463 | Yes |
| 33 | TRIB3 | TRIB3 Entrez, Source | tribbles homolog 3 (Drosophila) | 17533 | -0.636 | -0.3951 | Yes |
| 34 | G0S2 | G0S2 Entrez, Source | G0/G1switch 2 | 17689 | -0.721 | -0.3386 | Yes |
| 35 | ALPK2 | ALPK2 Entrez, Source | alpha-kinase 2 | 17792 | -0.794 | -0.2724 | Yes |
| 36 | ATF5 | ATF5 Entrez, Source | activating transcription factor 5 | 17867 | -0.867 | -0.1982 | Yes |
| 37 | BATF3 | 18027 | -1.163 | -0.1018 | Yes | ||
| 38 | ASB2 | ASB2 Entrez, Source | ankyrin repeat and SOCS box-containing 2 | 18029 | -1.170 | 0.0039 | Yes |