Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | NS50719_832_corrected2_collapsed_to_symbols.NS50719_832.cls #arhinia_versus_control.NS50719_832.cls #arhinia_versus_control_repos |
| Phenotype | NS50719_832.cls#arhinia_versus_control_repos |
| Upregulated in class | 0 |
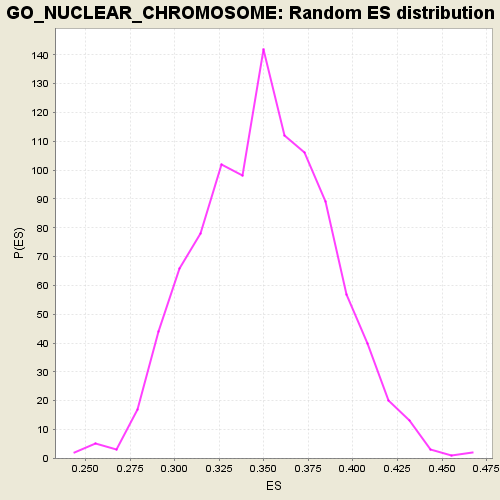
| GeneSet | GO_NUCLEAR_CHROMOSOME |
| Enrichment Score (ES) | -0.31383216 |
| Normalized Enrichment Score (NES) | NaN |
| Nominal p-value | NaN |
| FDR q-value | 1.0 |
| FWER p-Value | 0.0 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | MSH5 | MSH5 Entrez, Source | mutS homolog 5 (E. coli) | 42 | 3.125 | 0.0275 | No |
| 2 | MUC1 | MUC1 Entrez, Source | mucin 1, cell surface associated | 143 | 2.479 | 0.0459 | No |
| 3 | POLR3GL | POLR3GL Entrez, Source | polymerase (RNA) III (DNA directed) polypeptide G (32kD) like | 262 | 2.134 | 0.0601 | No |
| 4 | SCRT2 | SCRT2 Entrez, Source | scratch homolog 2, zinc finger protein (Drosophila) | 604 | 1.657 | 0.0582 | No |
| 5 | SYCE1L | 684 | 1.569 | 0.0690 | No | ||
| 6 | SMC1B | SMC1B Entrez, Source | structural maintenance of chromosomes 1B | 854 | 1.434 | 0.0739 | No |
| 7 | SMARCD3 | SMARCD3 Entrez, Source | SWI/SNF related, matrix associated, actin dependent regulator of chromatin, subfamily d, member 3 | 889 | 1.411 | 0.0855 | No |
| 8 | NLRP2 | 1324 | 1.180 | 0.0743 | No | ||
| 9 | PURA | PURA Entrez, Source | purine-rich element binding protein A | 1418 | 1.146 | 0.0804 | No |
| 10 | SP100 | SP100 Entrez, Source | SP100 nuclear antigen | 1610 | 1.077 | 0.0808 | No |
| 11 | RSPH1 | 2083 | 0.903 | 0.0650 | No | ||
| 12 | HIST1H1B | HIST1H1B Entrez, Source | histone cluster 1, H1b | 2395 | 0.818 | 0.0567 | No |
| 13 | USP3 | USP3 Entrez, Source | ubiquitin specific peptidase 3 | 2537 | 0.775 | 0.0567 | No |
| 14 | KLF1 | KLF1 Entrez, Source | Kruppel-like factor 1 (erythroid) | 2835 | 0.694 | 0.0480 | No |
| 15 | CBX7 | CBX7 Entrez, Source | chromobox homolog 7 | 2877 | 0.684 | 0.0524 | No |
| 16 | POLD4 | POLD4 Entrez, Source | polymerase (DNA-directed), delta 4 | 2918 | 0.674 | 0.0567 | No |
| 17 | ZFP57 | ZFP57 Entrez, Source | zinc finger protein 57 homolog (mouse) | 3018 | 0.653 | 0.0578 | No |
| 18 | JUNB | JUNB Entrez, Source | jun B proto-oncogene | 3223 | 0.605 | 0.0530 | No |
| 19 | HIST1H3J | HIST1H3J Entrez, Source | histone cluster 1, H3j | 3358 | 0.582 | 0.0516 | No |
| 20 | PMS2P3 | 3436 | 0.568 | 0.0530 | No | ||
| 21 | ZNF385A | 3603 | 0.530 | 0.0495 | No | ||
| 22 | SOX18 | SOX18 Entrez, Source | SRY (sex determining region Y)-box 18 | 3959 | 0.467 | 0.0356 | No |
| 23 | RUNX2 | RUNX2 Entrez, Source | runt-related transcription factor 2 | 4161 | 0.438 | 0.0294 | No |
| 24 | H2AFY2 | H2AFY2 Entrez, Source | H2A histone family, member Y2 | 4202 | 0.430 | 0.0314 | No |
| 25 | RUNX3 | RUNX3 Entrez, Source | runt-related transcription factor 3 | 4204 | 0.430 | 0.0354 | No |
| 26 | PLCB1 | PLCB1 Entrez, Source | phospholipase C, beta 1 (phosphoinositide-specific) | 4304 | 0.415 | 0.0342 | No |
| 27 | HIST1H2BL | HIST1H2BL Entrez, Source | histone cluster 1, H2bl | 4459 | 0.392 | 0.0300 | No |
| 28 | SYCE2 | SYCE2 Entrez, Source | synaptonemal complex central element protein 2 | 4472 | 0.390 | 0.0331 | No |
| 29 | H1F0 | H1F0 Entrez, Source | H1 histone family, member 0 | 4538 | 0.381 | 0.0333 | No |
| 30 | STAT6 | STAT6 Entrez, Source | signal transducer and activator of transcription 6, interleukin-4 induced | 4609 | 0.372 | 0.0333 | No |
| 31 | LIG4 | LIG4 Entrez, Source | ligase IV, DNA, ATP-dependent | 4771 | 0.354 | 0.0283 | No |
| 32 | CALCOCO1 | CALCOCO1 Entrez, Source | calcium binding and coiled-coil domain 1 | 4923 | 0.337 | 0.0237 | No |
| 33 | EME1 | EME1 Entrez, Source | essential meiotic endonuclease 1 homolog 1 (S. pombe) | 4925 | 0.337 | 0.0268 | No |
| 34 | SS18L1 | SS18L1 Entrez, Source | synovial sarcoma translocation gene on chromosome 18-like 1 | 5204 | 0.304 | 0.0154 | No |
| 35 | HIST1H4E | HIST1H4E Entrez, Source | histone cluster 1, H4e | 5438 | 0.280 | 0.0060 | No |
| 36 | DDX11 | DDX11 Entrez, Source | DEAD/H (Asp-Glu-Ala-Asp/His) box polypeptide 11 (CHL1-like helicase homolog, S. cerevisiae) | 5703 | 0.257 | -0.0052 | No |
| 37 | NR1D1 | NR1D1 Entrez, Source | nuclear receptor subfamily 1, group D, member 1 | 5739 | 0.253 | -0.0046 | No |
| 38 | SLX4 | 5783 | 0.249 | -0.0045 | No | ||
| 39 | SMARCC2 | SMARCC2 Entrez, Source | SWI/SNF related, matrix associated, actin dependent regulator of chromatin, subfamily c, member 2 | 5888 | 0.241 | -0.0075 | No |
| 40 | PMS1 | PMS1 Entrez, Source | PMS1 postmeiotic segregation increased 1 (S. cerevisiae) | 6012 | 0.231 | -0.0117 | No |
| 41 | RAD9A | RAD9A Entrez, Source | RAD9 homolog A (S. pombe) | 6023 | 0.230 | -0.0100 | No |
| 42 | ATR | ATR Entrez, Source | ataxia telangiectasia and Rad3 related | 6036 | 0.229 | -0.0085 | No |
| 43 | DMC1 | DMC1 Entrez, Source | DMC1 dosage suppressor of mck1 homolog, meiosis-specific homologous recombination (yeast) | 6095 | 0.225 | -0.0093 | No |
| 44 | GABPA | GABPA Entrez, Source | GA binding protein transcription factor, alpha subunit 60kDa | 6160 | 0.220 | -0.0105 | No |
| 45 | TERT | TERT Entrez, Source | telomerase reverse transcriptase | 6178 | 0.219 | -0.0093 | No |
| 46 | UHRF2 | UHRF2 Entrez, Source | ubiquitin-like, containing PHD and RING finger domains, 2 | 6180 | 0.219 | -0.0073 | No |
| 47 | SMAD3 | SMAD3 Entrez, Source | SMAD, mothers against DPP homolog 3 (Drosophila) | 6269 | 0.213 | -0.0098 | No |
| 48 | POT1 | POT1 Entrez, Source | POT1 protection of telomeres 1 homolog (S. pombe) | 6311 | 0.210 | -0.0099 | No |
| 49 | SYCP1 | SYCP1 Entrez, Source | synaptonemal complex protein 1 | 6392 | 0.205 | -0.0121 | No |
| 50 | ING2 | ING2 Entrez, Source | inhibitor of growth family, member 2 | 6427 | 0.202 | -0.0120 | No |
| 51 | NEK2 | NEK2 Entrez, Source | NIMA (never in mitosis gene a)-related kinase 2 | 6467 | 0.199 | -0.0121 | No |
| 52 | NR1H3 | NR1H3 Entrez, Source | nuclear receptor subfamily 1, group H, member 3 | 6529 | 0.194 | -0.0134 | No |
| 53 | HIST2H2AC | HIST2H2AC Entrez, Source | histone cluster 2, H2ac | 6531 | 0.194 | -0.0116 | No |
| 54 | CBX8 | CBX8 Entrez, Source | chromobox homolog 8 (Pc class homolog, Drosophila) | 6595 | 0.190 | -0.0131 | No |
| 55 | ERCC4 | ERCC4 Entrez, Source | excision repair cross-complementing rodent repair deficiency, complementation group 4 | 6695 | 0.184 | -0.0164 | No |
| 56 | H3F3A | H3F3A Entrez, Source | H3 histone, family 3A | 6744 | 0.181 | -0.0172 | No |
| 57 | STAG3 | STAG3 Entrez, Source | stromal antigen 3 | 6830 | 0.176 | -0.0199 | No |
| 58 | SUN2 | 6862 | 0.174 | -0.0199 | No | ||
| 59 | HIST1H2AL | HIST1H2AL Entrez, Source | histone cluster 1, H2al | 6908 | 0.171 | -0.0206 | No |
| 60 | TNKS | TNKS Entrez, Source | tankyrase, TRF1-interacting ankyrin-related ADP-ribose polymerase | 6955 | 0.168 | -0.0213 | No |
| 61 | PPP1CB | PPP1CB Entrez, Source | protein phosphatase 1, catalytic subunit, beta isoform | 6958 | 0.168 | -0.0198 | No |
| 62 | STAT1 | STAT1 Entrez, Source | signal transducer and activator of transcription 1, 91kDa | 6999 | 0.165 | -0.0203 | No |
| 63 | NFATC1 | NFATC1 Entrez, Source | nuclear factor of activated T-cells, cytoplasmic, calcineurin-dependent 1 | 7020 | 0.164 | -0.0198 | No |
| 64 | POLR3G | POLR3G Entrez, Source | polymerase (RNA) III (DNA directed) polypeptide G (32kD) | 7027 | 0.164 | -0.0186 | No |
| 65 | PPARD | PPARD Entrez, Source | peroxisome proliferative activated receptor, delta | 7060 | 0.163 | -0.0187 | No |
| 66 | SIRT2 | SIRT2 Entrez, Source | sirtuin (silent mating type information regulation 2 homolog) 2 (S. cerevisiae) | 7080 | 0.161 | -0.0181 | No |
| 67 | INO80B | 7164 | 0.157 | -0.0209 | No | ||
| 68 | HELB | HELB Entrez, Source | helicase (DNA) B | 7208 | 0.155 | -0.0217 | No |
| 69 | RARG | RARG Entrez, Source | retinoic acid receptor, gamma | 7251 | 0.152 | -0.0224 | No |
| 70 | MBD2 | MBD2 Entrez, Source | methyl-CpG binding domain protein 2 | 7290 | 0.150 | -0.0229 | No |
| 71 | BUB1 | BUB1 Entrez, Source | BUB1 budding uninhibited by benzimidazoles 1 homolog (yeast) | 7328 | 0.149 | -0.0234 | No |
| 72 | ASF1B | ASF1B Entrez, Source | ASF1 anti-silencing function 1 homolog B (S. cerevisiae) | 7451 | 0.142 | -0.0284 | No |
| 73 | GATAD2B | GATAD2B Entrez, Source | GATA zinc finger domain containing 2B | 7460 | 0.142 | -0.0275 | No |
| 74 | TEX11 | TEX11 Entrez, Source | testis expressed sequence 11 | 7489 | 0.141 | -0.0276 | No |
| 75 | SATB1 | SATB1 Entrez, Source | special AT-rich sequence binding protein 1 (binds to nuclear matrix/scaffold-associating DNA's) | 7539 | 0.139 | -0.0288 | No |
| 76 | CEBPB | CEBPB Entrez, Source | CCAAT/enhancer binding protein (C/EBP), beta | 7547 | 0.139 | -0.0278 | No |
| 77 | POU4F1 | POU4F1 Entrez, Source | POU domain, class 4, transcription factor 1 | 7574 | 0.137 | -0.0279 | No |
| 78 | SUZ12 | SUZ12 Entrez, Source | suppressor of zeste 12 homolog (Drosophila) | 7581 | 0.137 | -0.0269 | No |
| 79 | DCLRE1A | DCLRE1A Entrez, Source | DNA cross-link repair 1A (PSO2 homolog, S. cerevisiae) | 7714 | 0.131 | -0.0324 | No |
| 80 | PML | PML Entrez, Source | promyelocytic leukemia | 7732 | 0.130 | -0.0321 | No |
| 81 | CITED2 | CITED2 Entrez, Source | Cbp/p300-interacting transactivator, with Glu/Asp-rich carboxy-terminal domain, 2 | 7765 | 0.129 | -0.0325 | No |
| 82 | TINF2 | TINF2 Entrez, Source | TERF1 (TRF1)-interacting nuclear factor 2 | 7772 | 0.129 | -0.0316 | No |
| 83 | CREB1 | CREB1 Entrez, Source | cAMP responsive element binding protein 1 | 7931 | 0.121 | -0.0386 | No |
| 84 | ZRANB3 | ZRANB3 Entrez, Source | zinc finger, RAN-binding domain containing 3 | 7957 | 0.120 | -0.0388 | No |
| 85 | NCOA3 | NCOA3 Entrez, Source | nuclear receptor coactivator 3 | 8010 | 0.117 | -0.0403 | No |
| 86 | HDAC8 | HDAC8 Entrez, Source | histone deacetylase 8 | 8019 | 0.117 | -0.0396 | No |
| 87 | LRIF1 | 8032 | 0.116 | -0.0391 | No | ||
| 88 | DCLRE1C | DCLRE1C Entrez, Source | DNA cross-link repair 1C (PSO2 homolog, S. cerevisiae) | 8131 | 0.113 | -0.0431 | No |
| 89 | SMAD4 | SMAD4 Entrez, Source | SMAD, mothers against DPP homolog 4 (Drosophila) | 8137 | 0.112 | -0.0423 | No |
| 90 | INO80E | 8204 | 0.110 | -0.0447 | No | ||
| 91 | UBE2B | UBE2B Entrez, Source | ubiquitin-conjugating enzyme E2B (RAD6 homolog) | 8209 | 0.110 | -0.0439 | No |
| 92 | SMCHD1 | SMCHD1 Entrez, Source | structural maintenance of chromosomes flexible hinge domain containing 1 | 8215 | 0.109 | -0.0431 | No |
| 93 | HMGB2 | HMGB2 Entrez, Source | high-mobility group box 2 | 8276 | 0.107 | -0.0452 | No |
| 94 | TONSL | 8278 | 0.107 | -0.0442 | No | ||
| 95 | MPHOSPH8 | 8291 | 0.107 | -0.0438 | No | ||
| 96 | ATRX | ATRX Entrez, Source | alpha thalassemia/mental retardation syndrome X-linked (RAD54 homolog, S. cerevisiae) | 8348 | 0.105 | -0.0457 | No |
| 97 | CENPA | CENPA Entrez, Source | centromere protein A | 8358 | 0.104 | -0.0452 | No |
| 98 | TCF4 | TCF4 Entrez, Source | transcription factor 4 | 8363 | 0.104 | -0.0444 | No |
| 99 | PIF1 | 8436 | 0.101 | -0.0472 | No | ||
| 100 | SUV39H1 | SUV39H1 Entrez, Source | suppressor of variegation 3-9 homolog 1 (Drosophila) | 8453 | 0.100 | -0.0470 | No |
| 101 | JUND | JUND Entrez, Source | jun D proto-oncogene | 8472 | 0.099 | -0.0470 | No |
| 102 | DFFB | DFFB Entrez, Source | DNA fragmentation factor, 40kDa, beta polypeptide (caspase-activated DNase) | 8544 | 0.096 | -0.0498 | No |
| 103 | ERCC1 | ERCC1 Entrez, Source | excision repair cross-complementing rodent repair deficiency, complementation group 1 (includes overlapping antisense sequence) | 8571 | 0.095 | -0.0502 | No |
| 104 | RGS12 | RGS12 Entrez, Source | regulator of G-protein signalling 12 | 8573 | 0.095 | -0.0494 | No |
| 105 | RNF212 | 8587 | 0.095 | -0.0491 | No | ||
| 106 | ACD | ACD Entrez, Source | adrenocortical dysplasia homolog (mouse) | 8620 | 0.094 | -0.0499 | No |
| 107 | ADD3 | ADD3 Entrez, Source | adducin 3 (gamma) | 8678 | 0.091 | -0.0520 | No |
| 108 | BUB1B | BUB1B Entrez, Source | BUB1 budding uninhibited by benzimidazoles 1 homolog beta (yeast) | 8692 | 0.091 | -0.0518 | No |
| 109 | HIST1H2AI | HIST1H2AI Entrez, Source | histone cluster 1, H2ai | 8725 | 0.090 | -0.0526 | No |
| 110 | TCF12 | TCF12 Entrez, Source | transcription factor 12 (HTF4, helix-loop-helix transcription factors 4) | 8742 | 0.089 | -0.0526 | No |
| 111 | PURB | PURB Entrez, Source | purine-rich element binding protein B | 8800 | 0.086 | -0.0547 | No |
| 112 | SYN1 | SYN1 Entrez, Source | synapsin I | 8822 | 0.086 | -0.0550 | No |
| 113 | REPIN1 | REPIN1 Entrez, Source | replication initiator 1 | 8845 | 0.085 | -0.0553 | No |
| 114 | TERF2IP | TERF2IP Entrez, Source | telomeric repeat binding factor 2, interacting protein | 8862 | 0.085 | -0.0553 | No |
| 115 | HMGA2 | HMGA2 Entrez, Source | high mobility group AT-hook 2 | 8863 | 0.085 | -0.0545 | No |
| 116 | TOP2A | TOP2A Entrez, Source | topoisomerase (DNA) II alpha 170kDa | 8879 | 0.084 | -0.0545 | No |
| 117 | WRN | WRN Entrez, Source | Werner syndrome | 8926 | 0.083 | -0.0561 | No |
| 118 | TNKS2 | TNKS2 Entrez, Source | tankyrase, TRF1-interacting ankyrin-related ADP-ribose polymerase 2 | 8938 | 0.082 | -0.0559 | No |
| 119 | ESCO2 | ESCO2 Entrez, Source | establishment of cohesion 1 homolog 2 (S. cerevisiae) | 8978 | 0.081 | -0.0571 | No |
| 120 | CHAF1B | CHAF1B Entrez, Source | chromatin assembly factor 1, subunit B (p60) | 9018 | 0.080 | -0.0584 | No |
| 121 | AURKC | AURKC Entrez, Source | aurora kinase C | 9072 | 0.078 | -0.0604 | No |
| 122 | BLM | BLM Entrez, Source | Bloom syndrome | 9078 | 0.078 | -0.0599 | No |
| 123 | KAT2A | 9091 | 0.077 | -0.0598 | No | ||
| 124 | TIMELESS | TIMELESS Entrez, Source | timeless homolog (Drosophila) | 9094 | 0.077 | -0.0591 | No |
| 125 | THOC1 | THOC1 Entrez, Source | THO complex 1 | 9157 | 0.075 | -0.0616 | No |
| 126 | E2F4 | E2F4 Entrez, Source | E2F transcription factor 4, p107/p130-binding | 9173 | 0.075 | -0.0617 | No |
| 127 | ATM | ATM Entrez, Source | ataxia telangiectasia mutated (includes complementation groups A, C and D) | 9186 | 0.074 | -0.0616 | No |
| 128 | DBF4B | DBF4B Entrez, Source | DBF4 homolog B (S. cerevisiae) | 9294 | 0.071 | -0.0664 | No |
| 129 | IRF1 | IRF1 Entrez, Source | interferon regulatory factor 1 | 9297 | 0.071 | -0.0659 | No |
| 130 | PSIP1 | PSIP1 Entrez, Source | PC4 and SFRS1 interacting protein 1 | 9372 | 0.069 | -0.0691 | No |
| 131 | TUBG1 | TUBG1 Entrez, Source | tubulin, gamma 1 | 9425 | 0.067 | -0.0711 | No |
| 132 | BIRC5 | BIRC5 Entrez, Source | baculoviral IAP repeat-containing 5 (survivin) | 9479 | 0.066 | -0.0732 | No |
| 133 | STAT3 | STAT3 Entrez, Source | signal transducer and activator of transcription 3 (acute-phase response factor) | 9514 | 0.065 | -0.0744 | No |
| 134 | PPP1CC | PPP1CC Entrez, Source | protein phosphatase 1, catalytic subunit, gamma isoform | 9565 | 0.063 | -0.0763 | No |
| 135 | POLE | POLE Entrez, Source | polymerase (DNA directed), epsilon | 9572 | 0.063 | -0.0761 | No |
| 136 | POLD3 | POLD3 Entrez, Source | polymerase (DNA-directed), delta 3, accessory subunit | 9642 | 0.061 | -0.0790 | No |
| 137 | POGZ | POGZ Entrez, Source | pogo transposable element with ZNF domain | 9648 | 0.061 | -0.0787 | No |
| 138 | DVL3 | DVL3 Entrez, Source | dishevelled, dsh homolog 3 (Drosophila) | 9650 | 0.061 | -0.0782 | No |
| 139 | SMARCAL1 | SMARCAL1 Entrez, Source | SWI/SNF related, matrix associated, actin dependent regulator of chromatin, subfamily a-like 1 | 9747 | 0.058 | -0.0826 | No |
| 140 | TERF2 | TERF2 Entrez, Source | telomeric repeat binding factor 2 | 9751 | 0.058 | -0.0822 | No |
| 141 | EED | EED Entrez, Source | embryonic ectoderm development | 9795 | 0.057 | -0.0839 | No |
| 142 | RAD21 | RAD21 Entrez, Source | RAD21 homolog (S. pombe) | 9827 | 0.056 | -0.0849 | No |
| 143 | ANP32E | ANP32E Entrez, Source | acidic (leucine-rich) nuclear phosphoprotein 32 family, member E | 9888 | 0.054 | -0.0875 | No |
| 144 | DNA2 | 9917 | 0.053 | -0.0885 | No | ||
| 145 | THOC2 | THOC2 Entrez, Source | THO complex 2 | 9921 | 0.053 | -0.0881 | No |
| 146 | CBX1 | CBX1 Entrez, Source | chromobox homolog 1 (HP1 beta homolog Drosophila ) | 9939 | 0.053 | -0.0885 | No |
| 147 | NPM2 | NPM2 Entrez, Source | nucleophosmin/nucleoplasmin, 2 | 10004 | 0.051 | -0.0913 | No |
| 148 | MSH2 | MSH2 Entrez, Source | mutS homolog 2, colon cancer, nonpolyposis type 1 (E. coli) | 10014 | 0.050 | -0.0913 | No |
| 149 | MTA2 | MTA2 Entrez, Source | metastasis associated 1 family, member 2 | 10031 | 0.050 | -0.0917 | No |
| 150 | HDAC2 | HDAC2 Entrez, Source | histone deacetylase 2 | 10070 | 0.049 | -0.0932 | No |
| 151 | INCENP | INCENP Entrez, Source | inner centromere protein antigens 135/155kDa | 10099 | 0.048 | -0.0942 | No |
| 152 | POLE2 | POLE2 Entrez, Source | polymerase (DNA directed), epsilon 2 (p59 subunit) | 10136 | 0.047 | -0.0956 | No |
| 153 | SP1 | SP1 Entrez, Source | Sp1 transcription factor | 10157 | 0.046 | -0.0962 | No |
| 154 | CDK1 | 10283 | 0.043 | -0.1022 | No | ||
| 155 | WBP2 | WBP2 Entrez, Source | WW domain binding protein 2 | 10291 | 0.043 | -0.1022 | No |
| 156 | H2AFY | H2AFY Entrez, Source | H2A histone family, member Y | 10435 | 0.039 | -0.1092 | No |
| 157 | SMARCD2 | SMARCD2 Entrez, Source | SWI/SNF related, matrix associated, actin dependent regulator of chromatin, subfamily d, member 2 | 10442 | 0.038 | -0.1091 | No |
| 158 | NRIP1 | NRIP1 Entrez, Source | nuclear receptor interacting protein 1 | 10444 | 0.038 | -0.1088 | No |
| 159 | ARID1A | ARID1A Entrez, Source | AT rich interactive domain 1A (SWI- like) | 10450 | 0.038 | -0.1087 | No |
| 160 | MEN1 | MEN1 Entrez, Source | multiple endocrine neoplasia I | 10453 | 0.038 | -0.1085 | No |
| 161 | LDB1 | LDB1 Entrez, Source | LIM domain binding 1 | 10457 | 0.038 | -0.1083 | No |
| 162 | SUDS3 | SUDS3 Entrez, Source | suppressor of defective silencing 3 homolog (S. cerevisiae) | 10465 | 0.038 | -0.1083 | No |
| 163 | PHF12 | PHF12 Entrez, Source | PHD finger protein 12 | 10479 | 0.037 | -0.1086 | No |
| 164 | MBD1 | MBD1 Entrez, Source | methyl-CpG binding domain protein 1 | 10529 | 0.036 | -0.1108 | No |
| 165 | CBX5 | CBX5 Entrez, Source | chromobox homolog 5 (HP1 alpha homolog, Drosophila) | 10577 | 0.034 | -0.1129 | No |
| 166 | SIN3A | SIN3A Entrez, Source | SIN3 homolog A, transcription regulator (yeast) | 10591 | 0.034 | -0.1132 | No |
| 167 | UCHL5 | UCHL5 Entrez, Source | ubiquitin carboxyl-terminal hydrolase L5 | 10631 | 0.033 | -0.1149 | No |
| 168 | MEF2A | MEF2A Entrez, Source | MADS box transcription enhancer factor 2, polypeptide A (myocyte enhancer factor 2A) | 10644 | 0.033 | -0.1152 | No |
| 169 | ZMIZ2 | 10654 | 0.033 | -0.1154 | No | ||
| 170 | CCNB1IP1 | CCNB1IP1 Entrez, Source | cyclin B1 interacting protein 1 | 10662 | 0.032 | -0.1154 | No |
| 171 | LRPPRC | LRPPRC Entrez, Source | leucine-rich PPR-motif containing | 10676 | 0.032 | -0.1158 | No |
| 172 | TARDBP | TARDBP Entrez, Source | TAR DNA binding protein | 10729 | 0.030 | -0.1182 | No |
| 173 | KLHDC3 | KLHDC3 Entrez, Source | kelch domain containing 3 | 10746 | 0.030 | -0.1187 | No |
| 174 | TRIM24 | TRIM24 Entrez, Source | tripartite motif-containing 24 | 10763 | 0.030 | -0.1193 | No |
| 175 | FAM60A | FAM60A Entrez, Source | family with sequence similarity 60, member A | 10795 | 0.029 | -0.1206 | No |
| 176 | NCOR1 | NCOR1 Entrez, Source | nuclear receptor co-repressor 1 | 10802 | 0.029 | -0.1207 | No |
| 177 | EP400 | EP400 Entrez, Source | E1A binding protein p400 | 10811 | 0.028 | -0.1208 | No |
| 178 | THOC7 | THOC7 Entrez, Source | THO complex 7 homolog (Drosophila) | 10826 | 0.028 | -0.1213 | No |
| 179 | TCF7L2 | TCF7L2 Entrez, Source | transcription factor 7-like 2 (T-cell specific, HMG-box) | 10940 | 0.025 | -0.1269 | No |
| 180 | POLD1 | POLD1 Entrez, Source | polymerase (DNA directed), delta 1, catalytic subunit 125kDa | 10961 | 0.024 | -0.1277 | No |
| 181 | INO80C | 10986 | 0.024 | -0.1287 | No | ||
| 182 | SMC1A | SMC1A Entrez, Source | structural maintenance of chromosomes 1A | 11023 | 0.022 | -0.1303 | No |
| 183 | MECP2 | MECP2 Entrez, Source | methyl CpG binding protein 2 (Rett syndrome) | 11027 | 0.022 | -0.1303 | No |
| 184 | CTC1 | 11075 | 0.021 | -0.1325 | No | ||
| 185 | LRWD1 | 11104 | 0.020 | -0.1338 | No | ||
| 186 | NCAPD2 | NCAPD2 Entrez, Source | non-SMC condensin I complex, subunit D2 | 11126 | 0.019 | -0.1347 | No |
| 187 | GINS1 | GINS1 Entrez, Source | GINS complex subunit 1 (Psf1 homolog) | 11127 | 0.019 | -0.1345 | No |
| 188 | HMBOX1 | HMBOX1 Entrez, Source | homeobox containing 1 | 11149 | 0.019 | -0.1354 | No |
| 189 | SMARCA5 | SMARCA5 Entrez, Source | SWI/SNF related, matrix associated, actin dependent regulator of chromatin, subfamily a, member 5 | 11153 | 0.019 | -0.1354 | No |
| 190 | ACTL6A | ACTL6A Entrez, Source | actin-like 6A | 11180 | 0.018 | -0.1365 | No |
| 191 | PCGF2 | PCGF2 Entrez, Source | polycomb group ring finger 2 | 11211 | 0.017 | -0.1379 | No |
| 192 | ETV3 | ETV3 Entrez, Source | ets variant gene 3 | 11217 | 0.017 | -0.1380 | No |
| 193 | E2F1 | E2F1 Entrez, Source | E2F transcription factor 1 | 11314 | 0.015 | -0.1428 | No |
| 194 | H3F3B | H3F3B Entrez, Source | H3 histone, family 3B (H3.3B) | 11379 | 0.013 | -0.1460 | No |
| 195 | TNKS1BP1 | TNKS1BP1 Entrez, Source | tankyrase 1 binding protein 1, 182kDa | 11490 | 0.011 | -0.1516 | No |
| 196 | TCF3 | TCF3 Entrez, Source | transcription factor 3 (E2A immunoglobulin enhancer binding factors E12/E47) | 11492 | 0.011 | -0.1516 | No |
| 197 | RAD50 | RAD50 Entrez, Source | RAD50 homolog (S. cerevisiae) | 11497 | 0.010 | -0.1517 | No |
| 198 | EHMT2 | EHMT2 Entrez, Source | euchromatic histone-lysine N-methyltransferase 2 | 11498 | 0.010 | -0.1516 | No |
| 199 | HDAC1 | HDAC1 Entrez, Source | histone deacetylase 1 | 11502 | 0.010 | -0.1516 | No |
| 200 | SMARCB1 | SMARCB1 Entrez, Source | SWI/SNF related, matrix associated, actin dependent regulator of chromatin, subfamily b, member 1 | 11509 | 0.010 | -0.1518 | No |
| 201 | CDC45 | 11519 | 0.010 | -0.1522 | No | ||
| 202 | ORC4 | 11539 | 0.009 | -0.1531 | No | ||
| 203 | RAD51D | 11567 | 0.009 | -0.1544 | No | ||
| 204 | ZSCAN4 | ZSCAN4 Entrez, Source | zinc finger and SCAN domain containing 4 | 11573 | 0.009 | -0.1546 | No |
| 205 | POLR3D | POLR3D Entrez, Source | polymerase (RNA) III (DNA directed) polypeptide D, 44kDa | 11617 | 0.008 | -0.1567 | No |
| 206 | TP53 | TP53 Entrez, Source | tumor protein p53 (Li-Fraumeni syndrome) | 11625 | 0.008 | -0.1570 | No |
| 207 | YY1 | YY1 Entrez, Source | YY1 transcription factor | 11646 | 0.007 | -0.1580 | No |
| 208 | KDM1A | 11731 | 0.005 | -0.1623 | No | ||
| 209 | ORC2 | 11769 | 0.004 | -0.1641 | No | ||
| 210 | CTNNB1 | CTNNB1 Entrez, Source | catenin (cadherin-associated protein), beta 1, 88kDa | 11815 | 0.003 | -0.1664 | No |
| 211 | SFR1 | 11830 | 0.003 | -0.1671 | No | ||
| 212 | CHRAC1 | CHRAC1 Entrez, Source | chromatin accessibility complex 1 | 11861 | 0.002 | -0.1687 | No |
| 213 | BAZ1B | BAZ1B Entrez, Source | bromodomain adjacent to zinc finger domain, 1B | 11883 | 0.001 | -0.1697 | No |
| 214 | BRCA2 | BRCA2 Entrez, Source | breast cancer 2, early onset | 11922 | 0.000 | -0.1717 | No |
| 215 | ORC3 | 11924 | -0.000 | -0.1718 | No | ||
| 216 | NASP | NASP Entrez, Source | nuclear autoantigenic sperm protein (histone-binding) | 11946 | -0.001 | -0.1728 | No |
| 217 | ACTR8 | ACTR8 Entrez, Source | ARP8 actin-related protein 8 homolog (yeast) | 11969 | -0.001 | -0.1740 | No |
| 218 | PRKDC | PRKDC Entrez, Source | protein kinase, DNA-activated, catalytic polypeptide | 11978 | -0.001 | -0.1744 | No |
| 219 | KAT5 | 12001 | -0.002 | -0.1755 | No | ||
| 220 | MIS18BP1 | 12002 | -0.002 | -0.1755 | No | ||
| 221 | TRNP1 | 12022 | -0.003 | -0.1764 | No | ||
| 222 | RNF2 | RNF2 Entrez, Source | ring finger protein 2 | 12034 | -0.003 | -0.1770 | No |
| 223 | XRCC5 | XRCC5 Entrez, Source | X-ray repair complementing defective repair in Chinese hamster cells 5 (double-strand-break rejoining; Ku autoantigen, 80kDa) | 12051 | -0.003 | -0.1777 | No |
| 224 | RPA1 | RPA1 Entrez, Source | replication protein A1, 70kDa | 12081 | -0.004 | -0.1792 | No |
| 225 | MCM3 | MCM3 Entrez, Source | MCM3 minichromosome maintenance deficient 3 (S. cerevisiae) | 12107 | -0.005 | -0.1805 | No |
| 226 | SMAD2 | SMAD2 Entrez, Source | SMAD, mothers against DPP homolog 2 (Drosophila) | 12117 | -0.005 | -0.1809 | No |
| 227 | UBE2I | UBE2I Entrez, Source | ubiquitin-conjugating enzyme E2I (UBC9 homolog, yeast) | 12129 | -0.005 | -0.1814 | No |
| 228 | MORF4L1 | MORF4L1 Entrez, Source | mortality factor 4 like 1 | 12149 | -0.006 | -0.1823 | No |
| 229 | SMARCE1 | SMARCE1 Entrez, Source | SWI/SNF related, matrix associated, actin dependent regulator of chromatin, subfamily e, member 1 | 12196 | -0.007 | -0.1846 | No |
| 230 | TTC21B | TTC21B Entrez, Source | tetratricopeptide repeat domain 21B | 12245 | -0.008 | -0.1870 | No |
| 231 | DFFA | DFFA Entrez, Source | DNA fragmentation factor, 45kDa, alpha polypeptide | 12261 | -0.009 | -0.1877 | No |
| 232 | RPA3 | RPA3 Entrez, Source | replication protein A3, 14kDa | 12264 | -0.009 | -0.1877 | No |
| 233 | RBBP4 | RBBP4 Entrez, Source | retinoblastoma binding protein 4 | 12266 | -0.009 | -0.1877 | No |
| 234 | NCAPD3 | NCAPD3 Entrez, Source | non-SMC condensin II complex, subunit D3 | 12288 | -0.009 | -0.1887 | No |
| 235 | SMARCC1 | SMARCC1 Entrez, Source | SWI/SNF related, matrix associated, actin dependent regulator of chromatin, subfamily c, member 1 | 12315 | -0.010 | -0.1900 | No |
| 236 | HAT1 | HAT1 Entrez, Source | histone acetyltransferase 1 | 12355 | -0.011 | -0.1919 | No |
| 237 | GINS2 | GINS2 Entrez, Source | GINS complex subunit 2 (Psf2 homolog) | 12384 | -0.011 | -0.1932 | No |
| 238 | SAP130 | SAP130 Entrez, Source | Sin3A-associated protein, 130kDa | 12417 | -0.012 | -0.1947 | No |
| 239 | MMS22L | 12422 | -0.012 | -0.1948 | No | ||
| 240 | DDB1 | DDB1 Entrez, Source | damage-specific DNA binding protein 1, 127kDa | 12441 | -0.013 | -0.1956 | No |
| 241 | NACC2 | 12491 | -0.014 | -0.1980 | No | ||
| 242 | RING1 | RING1 Entrez, Source | ring finger protein 1 | 12549 | -0.015 | -0.2008 | No |
| 243 | TOP1MT | TOP1MT Entrez, Source | topoisomerase (DNA) I, mitochondrial | 12555 | -0.015 | -0.2009 | No |
| 244 | DCLRE1B | DCLRE1B Entrez, Source | DNA cross-link repair 1B (PSO2 homolog, S. cerevisiae) | 12569 | -0.016 | -0.2015 | No |
| 245 | NCOR2 | NCOR2 Entrez, Source | nuclear receptor co-repressor 2 | 12600 | -0.016 | -0.2029 | No |
| 246 | RAD51 | RAD51 Entrez, Source | RAD51 homolog (RecA homolog, E. coli) (S. cerevisiae) | 12626 | -0.017 | -0.2040 | No |
| 247 | MSH6 | MSH6 Entrez, Source | mutS homolog 6 (E. coli) | 12627 | -0.017 | -0.2038 | No |
| 248 | CSNK2A1 | CSNK2A1 Entrez, Source | casein kinase 2, alpha 1 polypeptide | 12684 | -0.019 | -0.2065 | No |
| 249 | CBX3 | CBX3 Entrez, Source | chromobox homolog 3 (HP1 gamma homolog, Drosophila) | 12696 | -0.019 | -0.2069 | No |
| 250 | SMC3 | SMC3 Entrez, Source | structural maintenance of chromosomes 3 | 12703 | -0.019 | -0.2071 | No |
| 251 | HIST1H2BH | HIST1H2BH Entrez, Source | histone cluster 1, H2bh | 12717 | -0.019 | -0.2076 | No |
| 252 | NFRKB | NFRKB Entrez, Source | nuclear factor related to kappaB binding protein | 12722 | -0.020 | -0.2076 | No |
| 253 | TFPT | TFPT Entrez, Source | TCF3 (E2A) fusion partner (in childhood Leukemia) | 12753 | -0.020 | -0.2089 | No |
| 254 | XRCC6 | XRCC6 Entrez, Source | X-ray repair complementing defective repair in Chinese hamster cells 6 (Ku autoantigen, 70kDa) | 12829 | -0.022 | -0.2126 | No |
| 255 | H2AFV | H2AFV Entrez, Source | H2A histone family, member V | 12833 | -0.022 | -0.2125 | No |
| 256 | RUVBL1 | RUVBL1 Entrez, Source | RuvB-like 1 (E. coli) | 12917 | -0.025 | -0.2166 | No |
| 257 | SSB | SSB Entrez, Source | Sjogren syndrome antigen B (autoantigen La) | 13022 | -0.027 | -0.2217 | No |
| 258 | ZNHIT1 | ZNHIT1 Entrez, Source | zinc finger, HIT type 1 | 13060 | -0.028 | -0.2233 | No |
| 259 | KDM4C | 13095 | -0.029 | -0.2248 | No | ||
| 260 | EXOSC9 | EXOSC9 Entrez, Source | exosome component 9 | 13120 | -0.029 | -0.2258 | No |
| 261 | POLR2B | POLR2B Entrez, Source | polymerase (RNA) II (DNA directed) polypeptide B, 140kDa | 13127 | -0.030 | -0.2258 | No |
| 262 | SMARCA4 | SMARCA4 Entrez, Source | SWI/SNF related, matrix associated, actin dependent regulator of chromatin, subfamily a, member 4 | 13142 | -0.030 | -0.2263 | No |
| 263 | MLH1 | MLH1 Entrez, Source | mutL homolog 1, colon cancer, nonpolyposis type 2 (E. coli) | 13200 | -0.031 | -0.2289 | No |
| 264 | CDC5L | CDC5L Entrez, Source | CDC5 cell division cycle 5-like (S. pombe) | 13222 | -0.032 | -0.2297 | No |
| 265 | ORC6 | 13229 | -0.032 | -0.2297 | No | ||
| 266 | SETX | SETX Entrez, Source | senataxin | 13283 | -0.034 | -0.2321 | No |
| 267 | ACTR5 | ACTR5 Entrez, Source | ARP5 actin-related protein 5 homolog (yeast) | 13290 | -0.034 | -0.2321 | No |
| 268 | NAT10 | NAT10 Entrez, Source | N-acetyltransferase 10 | 13320 | -0.035 | -0.2333 | No |
| 269 | WDR82 | WDR82 Entrez, Source | WD repeat domain 82 | 13326 | -0.035 | -0.2332 | No |
| 270 | NDC80 | 13352 | -0.036 | -0.2341 | No | ||
| 271 | HNRNPC | 13353 | -0.036 | -0.2338 | No | ||
| 272 | TERF1 | TERF1 Entrez, Source | telomeric repeat binding factor (NIMA-interacting) 1 | 13443 | -0.038 | -0.2380 | No |
| 273 | NCOA1 | NCOA1 Entrez, Source | nuclear receptor coactivator 1 | 13448 | -0.038 | -0.2379 | No |
| 274 | CHEK1 | CHEK1 Entrez, Source | CHK1 checkpoint homolog (S. pombe) | 13477 | -0.039 | -0.2389 | No |
| 275 | CHD4 | CHD4 Entrez, Source | chromodomain helicase DNA binding protein 4 | 13485 | -0.039 | -0.2389 | No |
| 276 | PCNA | PCNA Entrez, Source | proliferating cell nuclear antigen | 13511 | -0.040 | -0.2398 | No |
| 277 | MCM4 | MCM4 Entrez, Source | MCM4 minichromosome maintenance deficient 4 (S. cerevisiae) | 13517 | -0.040 | -0.2397 | No |
| 278 | SIN3B | SIN3B Entrez, Source | SIN3 homolog B, transcription regulator (yeast) | 13519 | -0.040 | -0.2394 | No |
| 279 | SIRT7 | SIRT7 Entrez, Source | sirtuin (silent mating type information regulation 2 homolog) 7 (S. cerevisiae) | 13549 | -0.041 | -0.2405 | No |
| 280 | CDC73 | CDC73 Entrez, Source | cell division cycle 73, Paf1/RNA polymerase II complex component, homolog (S. cerevisiae) | 13552 | -0.041 | -0.2402 | No |
| 281 | CHAF1A | CHAF1A Entrez, Source | chromatin assembly factor 1, subunit A (p150) | 13575 | -0.041 | -0.2410 | No |
| 282 | TBP | TBP Entrez, Source | TATA box binding protein | 13587 | -0.042 | -0.2411 | No |
| 283 | MCRS1 | MCRS1 Entrez, Source | microspherule protein 1 | 13655 | -0.044 | -0.2442 | No |
| 284 | NHP2 | 13764 | -0.046 | -0.2493 | No | ||
| 285 | UHRF1 | UHRF1 Entrez, Source | ubiquitin-like, containing PHD and RING finger domains, 1 | 13792 | -0.047 | -0.2503 | No |
| 286 | TCP1 | TCP1 Entrez, Source | t-complex 1 | 13800 | -0.047 | -0.2502 | No |
| 287 | RBMX | RBMX Entrez, Source | RNA binding motif protein, X-linked | 13827 | -0.048 | -0.2510 | No |
| 288 | RUVBL2 | RUVBL2 Entrez, Source | RuvB-like 2 (E. coli) | 13894 | -0.050 | -0.2540 | No |
| 289 | XRCC3 | XRCC3 Entrez, Source | X-ray repair complementing defective repair in Chinese hamster cells 3 | 13919 | -0.051 | -0.2547 | No |
| 290 | ASH2L | ASH2L Entrez, Source | ash2 (absent, small, or homeotic)-like (Drosophila) | 13952 | -0.052 | -0.2559 | No |
| 291 | BRD8 | BRD8 Entrez, Source | bromodomain containing 8 | 14007 | -0.053 | -0.2582 | No |
| 292 | EZH2 | EZH2 Entrez, Source | enhancer of zeste homolog 2 (Drosophila) | 14060 | -0.055 | -0.2603 | No |
| 293 | BUD31 | BUD31 Entrez, Source | BUD31 homolog (yeast) | 14070 | -0.056 | -0.2603 | No |
| 294 | HNRNPK | 14078 | -0.056 | -0.2601 | No | ||
| 295 | TOP1 | TOP1 Entrez, Source | topoisomerase (DNA) I | 14171 | -0.059 | -0.2643 | No |
| 296 | TOPBP1 | TOPBP1 Entrez, Source | topoisomerase (DNA) II binding protein 1 | 14172 | -0.059 | -0.2637 | No |
| 297 | MIS12 | MIS12 Entrez, Source | MIS12 homolog (yeast) | 14207 | -0.060 | -0.2649 | No |
| 298 | AURKB | AURKB Entrez, Source | aurora kinase B | 14220 | -0.060 | -0.2650 | No |
| 299 | TIPIN | TIPIN Entrez, Source | TIMELESS interacting protein | 14259 | -0.061 | -0.2664 | No |
| 300 | POLA2 | POLA2 Entrez, Source | polymerase (DNA directed), alpha 2 (70kD subunit) | 14264 | -0.061 | -0.2660 | No |
| 301 | TRRAP | TRRAP Entrez, Source | transformation/transcription domain-associated protein | 14280 | -0.062 | -0.2662 | No |
| 302 | POLE3 | POLE3 Entrez, Source | polymerase (DNA directed), epsilon 3 (p17 subunit) | 14312 | -0.063 | -0.2672 | No |
| 303 | THOC6 | THOC6 Entrez, Source | THO complex 6 homolog (Drosophila) | 14339 | -0.063 | -0.2679 | No |
| 304 | PPP1CA | PPP1CA Entrez, Source | protein phosphatase 1, catalytic subunit, alpha isoform | 14345 | -0.064 | -0.2676 | No |
| 305 | RAD1 | RAD1 Entrez, Source | RAD1 homolog (S. pombe) | 14368 | -0.064 | -0.2681 | No |
| 306 | BRMS1L | BRMS1L Entrez, Source | breast cancer metastasis-suppressor 1-like | 14407 | -0.066 | -0.2694 | No |
| 307 | HIRA | HIRA Entrez, Source | HIR histone cell cycle regulation defective homolog A (S. cerevisiae) | 14412 | -0.066 | -0.2690 | No |
| 308 | HSPA2 | HSPA2 Entrez, Source | heat shock 70kDa protein 2 | 14430 | -0.066 | -0.2693 | No |
| 309 | NUFIP1 | NUFIP1 Entrez, Source | nuclear fragile X mental retardation protein interacting protein 1 | 14495 | -0.068 | -0.2719 | No |
| 310 | PARP1 | PARP1 Entrez, Source | poly (ADP-ribose) polymerase family, member 1 | 14506 | -0.069 | -0.2718 | No |
| 311 | GAR1 | 14511 | -0.069 | -0.2713 | No | ||
| 312 | MCM7 | MCM7 Entrez, Source | MCM7 minichromosome maintenance deficient 7 (S. cerevisiae) | 14517 | -0.069 | -0.2709 | No |
| 313 | PLK1 | PLK1 Entrez, Source | polo-like kinase 1 (Drosophila) | 14539 | -0.070 | -0.2714 | No |
| 314 | SRF | SRF Entrez, Source | serum response factor (c-fos serum response element-binding transcription factor) | 14540 | -0.070 | -0.2707 | No |
| 315 | FEN1 | FEN1 Entrez, Source | flap structure-specific endonuclease 1 | 14558 | -0.070 | -0.2709 | No |
| 316 | H2AFZ | H2AFZ Entrez, Source | H2A histone family, member Z | 14561 | -0.071 | -0.2703 | No |
| 317 | DNMT3A | DNMT3A Entrez, Source | DNA (cytosine-5-)-methyltransferase 3 alpha | 14589 | -0.071 | -0.2711 | No |
| 318 | WRNIP1 | WRNIP1 Entrez, Source | Werner helicase interacting protein 1 | 14598 | -0.071 | -0.2708 | No |
| 319 | ASXL1 | ASXL1 Entrez, Source | additional sex combs like 1 (Drosophila) | 14635 | -0.072 | -0.2720 | No |
| 320 | BRD4 | BRD4 Entrez, Source | bromodomain containing 4 | 14655 | -0.073 | -0.2723 | No |
| 321 | ACTB | ACTB Entrez, Source | actin, beta | 14674 | -0.074 | -0.2725 | No |
| 322 | BRCA1 | BRCA1 Entrez, Source | breast cancer 1, early onset | 14710 | -0.076 | -0.2736 | No |
| 323 | RXRA | RXRA Entrez, Source | retinoid X receptor, alpha | 14740 | -0.077 | -0.2743 | No |
| 324 | CDCA5 | CDCA5 Entrez, Source | cell division cycle associated 5 | 14743 | -0.077 | -0.2737 | No |
| 325 | PLRG1 | PLRG1 Entrez, Source | pleiotropic regulator 1 (PRL1 homolog, Arabidopsis) | 14768 | -0.077 | -0.2742 | No |
| 326 | XPA | XPA Entrez, Source | xeroderma pigmentosum, complementation group A | 14791 | -0.078 | -0.2746 | No |
| 327 | TRIM28 | TRIM28 Entrez, Source | tripartite motif-containing 28 | 14830 | -0.080 | -0.2758 | No |
| 328 | SMARCAD1 | SMARCAD1 Entrez, Source | SWI/SNF-related, matrix-associated actin-dependent regulator of chromatin, subfamily a, containing DEAD/H box 1 | 14834 | -0.080 | -0.2752 | No |
| 329 | RCC1 | RCC1 Entrez, Source | regulator of chromosome condensation 1 | 14858 | -0.081 | -0.2756 | No |
| 330 | AURKA | AURKA Entrez, Source | aurora kinase A | 14932 | -0.083 | -0.2786 | No |
| 331 | UPF1 | UPF1 Entrez, Source | UPF1 regulator of nonsense transcripts homolog (yeast) | 14951 | -0.084 | -0.2787 | No |
| 332 | RPA2 | RPA2 Entrez, Source | replication protein A2, 32kDa | 14965 | -0.085 | -0.2786 | No |
| 333 | UBE2A | UBE2A Entrez, Source | ubiquitin-conjugating enzyme E2A (RAD6 homolog) | 15045 | -0.088 | -0.2819 | No |
| 334 | TFIP11 | TFIP11 Entrez, Source | tuftelin interacting protein 11 | 15086 | -0.090 | -0.2831 | No |
| 335 | DPF2 | DPF2 Entrez, Source | D4, zinc and double PHD fingers family 2 | 15109 | -0.091 | -0.2833 | No |
| 336 | MCM2 | MCM2 Entrez, Source | MCM2 minichromosome maintenance deficient 2, mitotin (S. cerevisiae) | 15143 | -0.092 | -0.2842 | No |
| 337 | BCAS2 | BCAS2 Entrez, Source | breast carcinoma amplified sequence 2 | 15201 | -0.095 | -0.2862 | No |
| 338 | SWI5 | 15202 | -0.095 | -0.2853 | No | ||
| 339 | CHMP1A | 15220 | -0.095 | -0.2853 | No | ||
| 340 | GINS4 | GINS4 Entrez, Source | GINS complex subunit 4 (Sld5 homolog) | 15233 | -0.096 | -0.2850 | No |
| 341 | APTX | APTX Entrez, Source | aprataxin | 15242 | -0.096 | -0.2845 | No |
| 342 | CCNB1 | CCNB1 Entrez, Source | cyclin B1 | 15299 | -0.098 | -0.2865 | No |
| 343 | PRPF19 | PRPF19 Entrez, Source | PRP19/PSO4 pre-mRNA processing factor 19 homolog (S. cerevisiae) | 15381 | -0.102 | -0.2897 | No |
| 344 | NBN | NBN Entrez, Source | nibrin | 15406 | -0.103 | -0.2899 | No |
| 345 | INO80 | 15438 | -0.105 | -0.2905 | No | ||
| 346 | POLA1 | POLA1 Entrez, Source | polymerase (DNA directed), alpha 1 | 15461 | -0.106 | -0.2907 | No |
| 347 | SETD3 | SETD3 Entrez, Source | SET domain containing 3 | 15512 | -0.109 | -0.2922 | No |
| 348 | ASF1A | ASF1A Entrez, Source | ASF1 anti-silencing function 1 homolog A (S. cerevisiae) | 15531 | -0.110 | -0.2921 | No |
| 349 | ORC5 | 15685 | -0.119 | -0.2989 | No | ||
| 350 | HIST2H2BE | HIST2H2BE Entrez, Source | histone cluster 2, H2be | 15708 | -0.119 | -0.2989 | No |
| 351 | APEX1 | APEX1 Entrez, Source | APEX nuclease (multifunctional DNA repair enzyme) 1 | 15713 | -0.120 | -0.2980 | No |
| 352 | MCM5 | MCM5 Entrez, Source | MCM5 minichromosome maintenance deficient 5, cell division cycle 46 (S. cerevisiae) | 15747 | -0.122 | -0.2985 | No |
| 353 | PBRM1 | 15773 | -0.123 | -0.2986 | No | ||
| 354 | ERCC5 | ERCC5 Entrez, Source | me)) | 15774 | -0.123 | -0.2975 | No |
| 355 | MCM10 | MCM10 Entrez, Source | MCM10 minichromosome maintenance deficient 10 (S. cerevisiae) | 15783 | -0.124 | -0.2967 | No |
| 356 | SMC2 | SMC2 Entrez, Source | structural maintenance of chromosomes 2 | 15870 | -0.128 | -0.2999 | No |
| 357 | CREBBP | CREBBP Entrez, Source | CREB binding protein (Rubinstein-Taybi syndrome) | 15895 | -0.130 | -0.2999 | No |
| 358 | RRS1 | RRS1 Entrez, Source | RRS1 ribosome biogenesis regulator homolog (S. cerevisiae) | 15903 | -0.130 | -0.2991 | No |
| 359 | RAD51C | RAD51C Entrez, Source | RAD51 homolog C (S. cerevisiae) | 15904 | -0.130 | -0.2978 | No |
| 360 | PINX1 | PINX1 Entrez, Source | - | 15924 | -0.131 | -0.2976 | No |
| 361 | SIRT1 | SIRT1 Entrez, Source | sirtuin (silent mating type information regulation 2 homolog) 1 (S. cerevisiae) | 15927 | -0.131 | -0.2964 | No |
| 362 | THOC5 | THOC5 Entrez, Source | THO complex 5 | 15946 | -0.132 | -0.2961 | No |
| 363 | SIRT6 | SIRT6 Entrez, Source | sirtuin (silent mating type information regulation 2 homolog) 6 (S. cerevisiae) | 16049 | -0.139 | -0.3000 | No |
| 364 | MBD3 | MBD3 Entrez, Source | methyl-CpG binding domain protein 3 | 16054 | -0.139 | -0.2989 | No |
| 365 | TOX4 | 16107 | -0.143 | -0.3002 | No | ||
| 366 | TCF7 | TCF7 Entrez, Source | transcription factor 7 (T-cell specific, HMG-box) | 16108 | -0.143 | -0.2989 | No |
| 367 | RFX3 | RFX3 Entrez, Source | regulatory factor X, 3 (influences HLA class II expression) | 16150 | -0.146 | -0.2996 | No |
| 368 | RARA | RARA Entrez, Source | retinoic acid receptor, alpha | 16232 | -0.152 | -0.3024 | No |
| 369 | MLH3 | MLH3 Entrez, Source | mutL homolog 3 (E. coli) | 16258 | -0.154 | -0.3022 | No |
| 370 | PPP1R10 | PPP1R10 Entrez, Source | protein phosphatase 1, regulatory subunit 10 | 16348 | -0.162 | -0.3052 | No |
| 371 | BEND3 | 16376 | -0.164 | -0.3051 | No | ||
| 372 | ALKBH1 | ALKBH1 Entrez, Source | alkB, alkylation repair homolog 1 (E. coli) | 16420 | -0.168 | -0.3057 | No |
| 373 | ENC1 | ENC1 Entrez, Source | ectodermal-neural cortex (with BTB-like domain) | 16465 | -0.171 | -0.3063 | No |
| 374 | H2AFX | H2AFX Entrez, Source | H2A histone family, member X | 16468 | -0.172 | -0.3048 | No |
| 375 | NOL6 | NOL6 Entrez, Source | nucleolar protein family 6 (RNA-associated) | 16531 | -0.177 | -0.3063 | No |
| 376 | ING3 | ING3 Entrez, Source | inhibitor of growth family, member 3 | 16563 | -0.182 | -0.3062 | No |
| 377 | BRMS1 | BRMS1 Entrez, Source | breast cancer metastasis suppressor 1 | 16620 | -0.189 | -0.3073 | No |
| 378 | MCM6 | MCM6 Entrez, Source | MCM6 minichromosome maintenance deficient 6 (MIS5 homolog, S. pombe) (S. cerevisiae) | 16691 | -0.197 | -0.3090 | No |
| 379 | ORC1 | 16733 | -0.203 | -0.3092 | No | ||
| 380 | H2AFJ | H2AFJ Entrez, Source | H2A histone family, member J | 16802 | -0.212 | -0.3107 | No |
| 381 | HIST1H1E | HIST1H1E Entrez, Source | histone cluster 1, H1e | 16814 | -0.214 | -0.3093 | No |
| 382 | HUS1 | HUS1 Entrez, Source | HUS1 checkpoint homolog (S. pombe) | 16863 | -0.221 | -0.3096 | No |
| 383 | THOC3 | THOC3 Entrez, Source | THO complex 3 | 16869 | -0.222 | -0.3078 | No |
| 384 | TTN | TTN Entrez, Source | titin | 16898 | -0.226 | -0.3071 | No |
| 385 | HIST3H2A | HIST3H2A Entrez, Source | histone cluster 3, H2a | 16938 | -0.231 | -0.3069 | No |
| 386 | FOXH1 | FOXH1 Entrez, Source | forkhead box H1 | 16982 | -0.237 | -0.3069 | No |
| 387 | FER | FER Entrez, Source | fer (fps/fes related) tyrosine kinase (phosphoprotein NCP94) | 17028 | -0.242 | -0.3069 | No |
| 388 | REC8 | 17041 | -0.244 | -0.3052 | No | ||
| 389 | FIGNL1 | FIGNL1 Entrez, Source | fidgetin-like 1 | 17082 | -0.250 | -0.3049 | No |
| 390 | MSH3 | MSH3 Entrez, Source | mutS homolog 3 (E. coli) | 17087 | -0.250 | -0.3027 | No |
| 391 | T | T Entrez, Source | T, brachyury homolog (mouse) | 17160 | -0.261 | -0.3040 | No |
| 392 | KAT2B | 17222 | -0.270 | -0.3046 | No | ||
| 393 | FOXD3 | FOXD3 Entrez, Source | forkhead box D3 | 17358 | -0.297 | -0.3087 | No |
| 394 | MXD1 | MXD1 Entrez, Source | MAX dimerization protein 1 | 17458 | -0.320 | -0.3108 | Yes |
| 395 | HEY2 | HEY2 Entrez, Source | hairy/enhancer-of-split related with YRPW motif 2 | 17476 | -0.323 | -0.3086 | Yes |
| 396 | THRB | THRB Entrez, Source | thyroid hormone receptor, beta (erythroblastic leukemia viral (v-erb-a) oncogene homolog 2, avian) | 17498 | -0.328 | -0.3066 | Yes |
| 397 | PPARGC1A | PPARGC1A Entrez, Source | peroxisome proliferative activated receptor, gamma, coactivator 1, alpha | 17509 | -0.330 | -0.3040 | Yes |
| 398 | AR | AR Entrez, Source | androgen receptor (dihydrotestosterone receptor; testicular feminization; spinal and bulbar muscular atrophy; Kennedy disease) | 17517 | -0.331 | -0.3012 | Yes |
| 399 | HIST1H2BF | HIST1H2BF Entrez, Source | histone cluster 1, H2bf | 17743 | -0.382 | -0.3092 | Yes |
| 400 | POLE4 | POLE4 Entrez, Source | polymerase (DNA-directed), epsilon 4 (p12 subunit) | 17784 | -0.392 | -0.3075 | Yes |
| 401 | HIST1H3D | HIST1H3D Entrez, Source | histone cluster 1, H3d | 17891 | -0.423 | -0.3090 | Yes |
| 402 | HIST1H1D | HIST1H1D Entrez, Source | histone cluster 1, H1d | 17930 | -0.433 | -0.3068 | Yes |
| 403 | KIFAP3 | KIFAP3 Entrez, Source | kinesin-associated protein 3 | 17940 | -0.435 | -0.3032 | Yes |
| 404 | FOXC1 | FOXC1 Entrez, Source | forkhead box C1 | 18086 | -0.485 | -0.3060 | Yes |
| 405 | SMARCA2 | SMARCA2 Entrez, Source | SWI/SNF related, matrix associated, actin dependent regulator of chromatin, subfamily a, member 2 | 18190 | -0.518 | -0.3064 | Yes |
| 406 | H3F3C | 18325 | -0.567 | -0.3080 | Yes | ||
| 407 | HIST1H2BE | HIST1H2BE Entrez, Source | histone cluster 1, H2be | 18411 | -0.608 | -0.3066 | Yes |
| 408 | HIST1H3C | HIST1H3C Entrez, Source | histone cluster 1, H3c | 18447 | -0.619 | -0.3025 | Yes |
| 409 | HIST1H2AC | HIST1H2AC Entrez, Source | histone cluster 1, H2ac | 18496 | -0.643 | -0.2989 | Yes |
| 410 | MIXL1 | MIXL1 Entrez, Source | Mix1 homeobox-like 1 (Xenopus laevis) | 18537 | -0.660 | -0.2947 | Yes |
| 411 | HIST3H2BB | HIST3H2BB Entrez, Source | histone cluster 3, H2bb | 18599 | -0.688 | -0.2913 | Yes |
| 412 | JUN | JUN Entrez, Source | jun oncogene | 18625 | -0.705 | -0.2859 | Yes |
| 413 | HIST1H3B | HIST1H3B Entrez, Source | histone cluster 1, H3b | 18676 | -0.729 | -0.2816 | Yes |
| 414 | HIST1H1C | HIST1H1C Entrez, Source | histone cluster 1, H1c | 18954 | -0.882 | -0.2875 | Yes |
| 415 | HIST1H2AG | HIST1H2AG Entrez, Source | histone cluster 1, H2ag | 18958 | -0.883 | -0.2793 | Yes |
| 416 | KLF4 | KLF4 Entrez, Source | Kruppel-like factor 4 (gut) | 19008 | -0.915 | -0.2731 | Yes |
| 417 | HIST4H4 | HIST4H4 Entrez, Source | histone cluster 4, H4 | 19052 | -0.943 | -0.2664 | Yes |
| 418 | HIST1H3A | HIST1H3A Entrez, Source | histone cluster 1, H3a | 19137 | -0.996 | -0.2613 | Yes |
| 419 | HIST1H3H | HIST1H3H Entrez, Source | histone cluster 1, H3h | 19207 | -1.051 | -0.2548 | Yes |
| 420 | SYCP2 | SYCP2 Entrez, Source | synaptonemal complex protein 2 | 19214 | -1.062 | -0.2451 | Yes |
| 421 | HIST1H2BO | HIST1H2BO Entrez, Source | histone cluster 1, H2bo | 19226 | -1.071 | -0.2354 | Yes |
| 422 | HIST1H2BJ | HIST1H2BJ Entrez, Source | histone cluster 1, H2bj | 19248 | -1.094 | -0.2261 | Yes |
| 423 | HIST1H2AE | HIST1H2AE Entrez, Source | histone cluster 1, H2ae | 19289 | -1.131 | -0.2175 | Yes |
| 424 | HIST1H2BN | HIST1H2BN Entrez, Source | histone cluster 1, H2bn | 19368 | -1.195 | -0.2101 | Yes |
| 425 | IRF4 | IRF4 Entrez, Source | interferon regulatory factor 4 | 19504 | -1.373 | -0.2041 | Yes |
| 426 | HIST1H3G | HIST1H3G Entrez, Source | histone cluster 1, H3g | 19519 | -1.394 | -0.1916 | Yes |
| 427 | SNAI2 | SNAI2 Entrez, Source | snail homolog 2 (Drosophila) | 19521 | -1.396 | -0.1784 | Yes |
| 428 | HIST1H2BD | HIST1H2BD Entrez, Source | histone cluster 1, H2bd | 19576 | -1.507 | -0.1668 | Yes |
| 429 | HIST1H4H | HIST1H4H Entrez, Source | histone cluster 1, H4h | 19623 | -1.605 | -0.1540 | Yes |
| 430 | PAX6 | PAX6 Entrez, Source | paired box gene 6 (aniridia, keratitis) | 19656 | -1.674 | -0.1397 | Yes |
| 431 | HIST1H4I | HIST1H4I Entrez, Source | histone cluster 1, H4i | 19684 | -1.758 | -0.1244 | Yes |
| 432 | HIST1H2BK | HIST1H2BK Entrez, Source | histone cluster 1, H2bk | 19715 | -1.903 | -0.1079 | Yes |
| 433 | HIST1H3E | HIST1H3E Entrez, Source | histone cluster 1, H3e | 19736 | -2.067 | -0.0893 | Yes |
| 434 | HIST1H2BG | HIST1H2BG Entrez, Source | histone cluster 1, H2bg | 19741 | -2.101 | -0.0696 | Yes |
| 435 | IPO4 | IPO4 Entrez, Source | importin 4 | 19764 | -2.325 | -0.0486 | Yes |
| 436 | HIST1H2BC | HIST1H2BC Entrez, Source | histone cluster 1, H2bc | 19771 | -2.393 | -0.0262 | Yes |
| 437 | GATA3 | GATA3 Entrez, Source | GATA binding protein 3 | 19788 | -2.906 | 0.0006 | Yes |