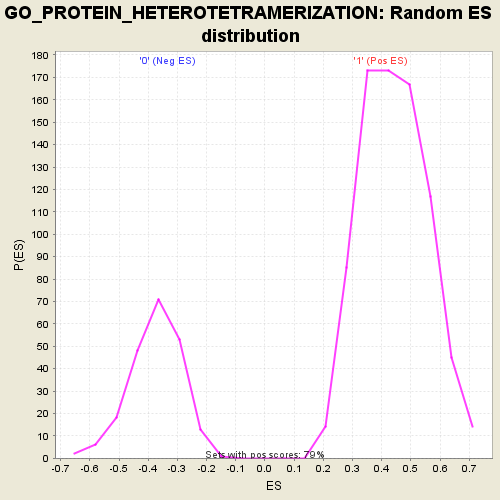
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | NS50719_832_corrected2_collapsed_to_symbols.NS50719_832.cls #arhinia_versus_control.NS50719_832.cls #arhinia_versus_control_repos |
| Phenotype | NS50719_832.cls#arhinia_versus_control_repos |
| Upregulated in class | 0 |
| GeneSet | GO_PROTEIN_HETEROTETRAMERIZATION |
| Enrichment Score (ES) | -0.782594 |
| Normalized Enrichment Score (NES) | -2.0881126 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.002962923 |
| FWER p-Value | 0.041 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | HIST1H3J | HIST1H3J Entrez, Source | histone cluster 1, H3j | 3358 | 0.582 | -0.1300 | No |
| 2 | HIST1H4E | HIST1H4E Entrez, Source | histone cluster 1, H4e | 5438 | 0.280 | -0.2160 | No |
| 3 | LRP4 | LRP4 Entrez, Source | low density lipoprotein receptor-related protein 4 | 8217 | 0.109 | -0.3490 | No |
| 4 | NUP54 | NUP54 Entrez, Source | nucleoporin 54kDa | 10474 | 0.037 | -0.4605 | No |
| 5 | PDSS2 | PDSS2 Entrez, Source | prenyl (decaprenyl) diphosphate synthase, subunit 2 | 10637 | 0.033 | -0.4664 | No |
| 6 | RRM1 | RRM1 Entrez, Source | ribonucleotide reductase M1 polypeptide | 11158 | 0.018 | -0.4915 | No |
| 7 | XRCC6 | XRCC6 Entrez, Source | X-ray repair complementing defective repair in Chinese hamster cells 6 (Ku autoantigen, 70kDa) | 12829 | -0.022 | -0.5744 | No |
| 8 | INSR | INSR Entrez, Source | insulin receptor | 13235 | -0.032 | -0.5926 | No |
| 9 | NLGN1 | NLGN1 Entrez, Source | neuroligin 1 | 16056 | -0.140 | -0.7257 | No |
| 10 | RRM2 | RRM2 Entrez, Source | ribonucleotide reductase M2 polypeptide | 16878 | -0.223 | -0.7520 | No |
| 11 | PDSS1 | PDSS1 Entrez, Source | prenyl (decaprenyl) diphosphate synthase, subunit 1 | 17368 | -0.300 | -0.7562 | Yes |
| 12 | HIST1H3D | HIST1H3D Entrez, Source | histone cluster 1, H3d | 17891 | -0.423 | -0.7537 | Yes |
| 13 | HIST1H3C | HIST1H3C Entrez, Source | histone cluster 1, H3c | 18447 | -0.619 | -0.7394 | Yes |
| 14 | S100A10 | S100A10 Entrez, Source | S100 calcium binding protein A10 | 18471 | -0.630 | -0.6975 | Yes |
| 15 | ANXA2 | ANXA2 Entrez, Source | annexin A2 | 18476 | -0.634 | -0.6544 | Yes |
| 16 | HIST1H3B | HIST1H3B Entrez, Source | histone cluster 1, H3b | 18676 | -0.729 | -0.6147 | Yes |
| 17 | HIST4H4 | HIST4H4 Entrez, Source | histone cluster 4, H4 | 19052 | -0.943 | -0.5691 | Yes |
| 18 | HIST1H3A | HIST1H3A Entrez, Source | histone cluster 1, H3a | 19137 | -0.996 | -0.5052 | Yes |
| 19 | HIST1H3H | HIST1H3H Entrez, Source | histone cluster 1, H3h | 19207 | -1.051 | -0.4368 | Yes |
| 20 | HIST1H3G | HIST1H3G Entrez, Source | histone cluster 1, H3g | 19519 | -1.394 | -0.3572 | Yes |
| 21 | HIST1H4H | HIST1H4H Entrez, Source | histone cluster 1, H4h | 19623 | -1.605 | -0.2527 | Yes |
| 22 | HIST1H4I | HIST1H4I Entrez, Source | histone cluster 1, H4i | 19684 | -1.758 | -0.1355 | Yes |
| 23 | HIST1H3E | HIST1H3E Entrez, Source | histone cluster 1, H3e | 19736 | -2.067 | 0.0032 | Yes |