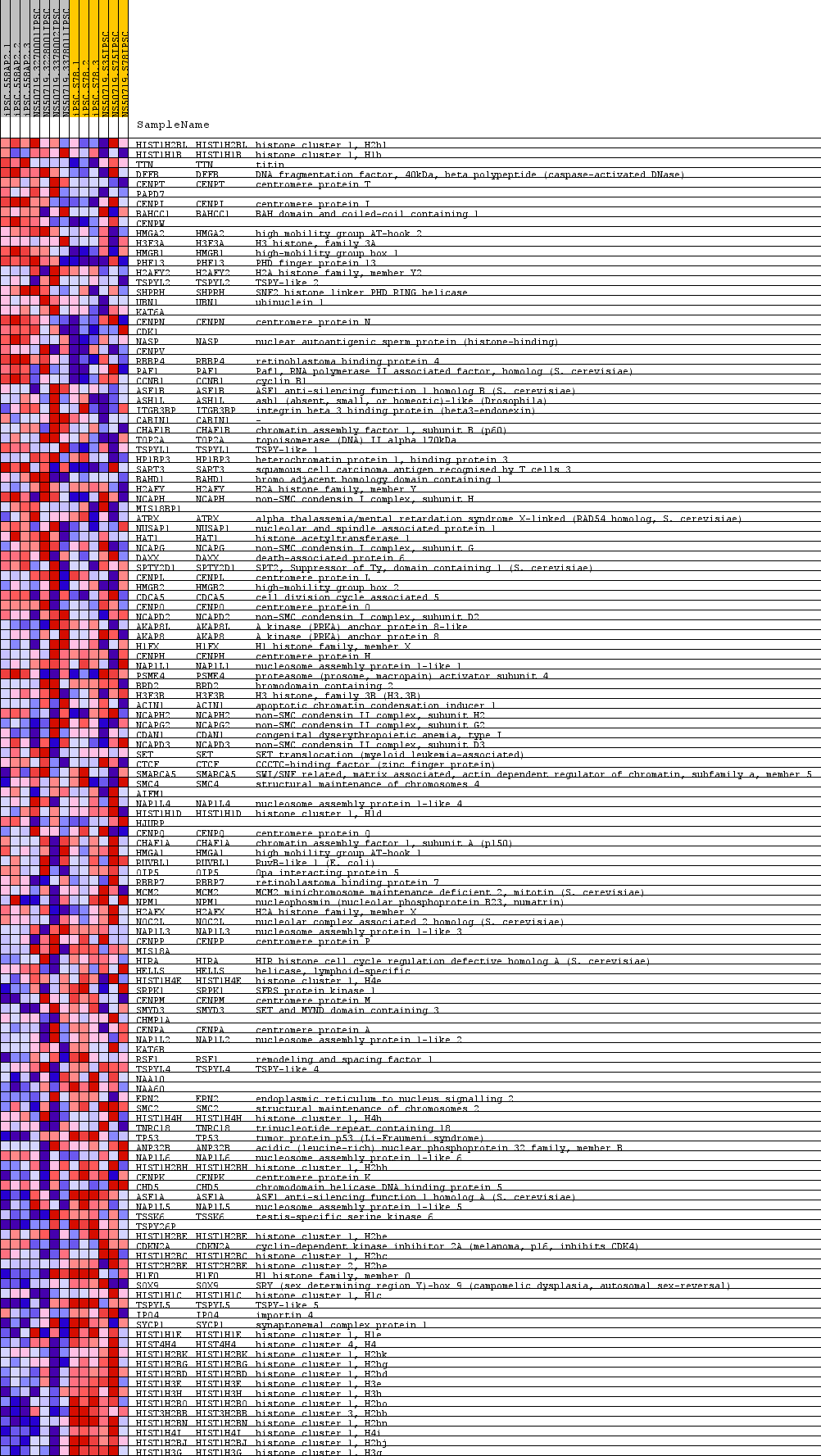
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | NS50719_832_collapsed_to_symbols.NS50719_832.cls #arhinia_versus_control.NS50719_832.cls #arhinia_versus_control_repos |
| Phenotype | NS50719_832.cls#arhinia_versus_control_repos |
| Upregulated in class | 0 |
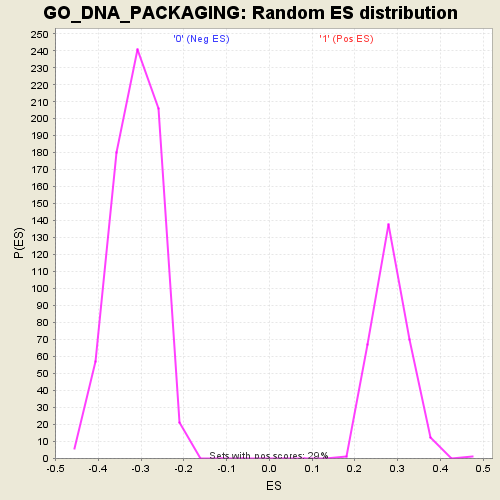
| GeneSet | GO_DNA_PACKAGING |
| Enrichment Score (ES) | -0.62926793 |
| Normalized Enrichment Score (NES) | -2.015373 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.0016398052 |
| FWER p-Value | 0.008 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | HIST1H2BL | HIST1H2BL Entrez, Source | histone cluster 1, H2bl | 1113 | 0.447 | -0.0411 | No |
| 2 | HIST1H1B | HIST1H1B Entrez, Source | histone cluster 1, H1b | 1636 | 0.338 | -0.0548 | No |
| 3 | TTN | TTN Entrez, Source | titin | 2874 | 0.189 | -0.1140 | No |
| 4 | DFFB | DFFB Entrez, Source | DNA fragmentation factor, 40kDa, beta polypeptide (caspase-activated DNase) | 3743 | 0.128 | -0.1557 | No |
| 5 | CENPT | CENPT Entrez, Source | centromere protein T | 4193 | 0.105 | -0.1756 | No |
| 6 | PAPD7 | 4410 | 0.096 | -0.1832 | No | ||
| 7 | CENPI | CENPI Entrez, Source | centromere protein I | 4427 | 0.096 | -0.1799 | No |
| 8 | BAHCC1 | BAHCC1 Entrez, Source | BAH domain and coiled-coil containing 1 | 4481 | 0.094 | -0.1787 | No |
| 9 | CENPW | 4559 | 0.091 | -0.1789 | No | ||
| 10 | HMGA2 | HMGA2 Entrez, Source | high mobility group AT-hook 2 | 4738 | 0.085 | -0.1849 | No |
| 11 | H3F3A | H3F3A Entrez, Source | H3 histone, family 3A | 4796 | 0.083 | -0.1843 | No |
| 12 | HMGB1 | HMGB1 Entrez, Source | high-mobility group box 1 | 4901 | 0.078 | -0.1866 | No |
| 13 | PHF13 | PHF13 Entrez, Source | PHD finger protein 13 | 5124 | 0.072 | -0.1955 | No |
| 14 | H2AFY2 | H2AFY2 Entrez, Source | H2A histone family, member Y2 | 5352 | 0.066 | -0.2050 | No |
| 15 | TSPYL2 | TSPYL2 Entrez, Source | TSPY-like 2 | 5567 | 0.061 | -0.2140 | No |
| 16 | SHPRH | SHPRH Entrez, Source | SNF2 histone linker PHD RING helicase | 5572 | 0.061 | -0.2115 | No |
| 17 | UBN1 | UBN1 Entrez, Source | ubinuclein 1 | 5613 | 0.060 | -0.2111 | No |
| 18 | KAT6A | 5657 | 0.060 | -0.2108 | No | ||
| 19 | CENPN | CENPN Entrez, Source | centromere protein N | 5773 | 0.057 | -0.2146 | No |
| 20 | CDK1 | 5810 | 0.056 | -0.2141 | No | ||
| 21 | NASP | NASP Entrez, Source | nuclear autoantigenic sperm protein (histone-binding) | 5820 | 0.056 | -0.2121 | No |
| 22 | CENPV | 5884 | 0.054 | -0.2132 | No | ||
| 23 | RBBP4 | RBBP4 Entrez, Source | retinoblastoma binding protein 4 | 5982 | 0.052 | -0.2162 | No |
| 24 | PAF1 | PAF1 Entrez, Source | Paf1, RNA polymerase II associated factor, homolog (S. cerevisiae) | 6144 | 0.049 | -0.2229 | No |
| 25 | CCNB1 | CCNB1 Entrez, Source | cyclin B1 | 6149 | 0.048 | -0.2210 | No |
| 26 | ASF1B | ASF1B Entrez, Source | ASF1 anti-silencing function 1 homolog B (S. cerevisiae) | 6218 | 0.047 | -0.2226 | No |
| 27 | ASH1L | ASH1L Entrez, Source | ash1 (absent, small, or homeotic)-like (Drosophila) | 6266 | 0.046 | -0.2232 | No |
| 28 | ITGB3BP | ITGB3BP Entrez, Source | integrin beta 3 binding protein (beta3-endonexin) | 6501 | 0.042 | -0.2341 | No |
| 29 | CABIN1 | CABIN1 Entrez, Source | - | 6510 | 0.042 | -0.2327 | No |
| 30 | CHAF1B | CHAF1B Entrez, Source | chromatin assembly factor 1, subunit B (p60) | 6516 | 0.041 | -0.2312 | No |
| 31 | TOP2A | TOP2A Entrez, Source | topoisomerase (DNA) II alpha 170kDa | 6685 | 0.038 | -0.2386 | No |
| 32 | TSPYL1 | TSPYL1 Entrez, Source | TSPY-like 1 | 6840 | 0.035 | -0.2455 | No |
| 33 | HP1BP3 | HP1BP3 Entrez, Source | heterochromatin protein 1, binding protein 3 | 6963 | 0.033 | -0.2507 | No |
| 34 | SART3 | SART3 Entrez, Source | squamous cell carcinoma antigen recognised by T cells 3 | 6994 | 0.033 | -0.2509 | No |
| 35 | BAHD1 | BAHD1 Entrez, Source | bromo adjacent homology domain containing 1 | 7103 | 0.030 | -0.2555 | No |
| 36 | H2AFY | H2AFY Entrez, Source | H2A histone family, member Y | 7239 | 0.028 | -0.2616 | No |
| 37 | NCAPH | NCAPH Entrez, Source | non-SMC condensin I complex, subunit H | 7320 | 0.027 | -0.2648 | No |
| 38 | MIS18BP1 | 7344 | 0.027 | -0.2649 | No | ||
| 39 | ATRX | ATRX Entrez, Source | alpha thalassemia/mental retardation syndrome X-linked (RAD54 homolog, S. cerevisiae) | 7398 | 0.026 | -0.2667 | No |
| 40 | NUSAP1 | NUSAP1 Entrez, Source | nucleolar and spindle associated protein 1 | 7458 | 0.025 | -0.2688 | No |
| 41 | HAT1 | HAT1 Entrez, Source | histone acetyltransferase 1 | 7675 | 0.021 | -0.2796 | No |
| 42 | NCAPG | NCAPG Entrez, Source | non-SMC condensin I complex, subunit G | 7753 | 0.020 | -0.2830 | No |
| 43 | DAXX | DAXX Entrez, Source | death-associated protein 6 | 7886 | 0.018 | -0.2894 | No |
| 44 | SPTY2D1 | SPTY2D1 Entrez, Source | SPT2, Suppressor of Ty, domain containing 1 (S. cerevisiae) | 7955 | 0.017 | -0.2923 | No |
| 45 | CENPL | CENPL Entrez, Source | centromere protein L | 8036 | 0.016 | -0.2960 | No |
| 46 | HMGB2 | HMGB2 Entrez, Source | high-mobility group box 2 | 8044 | 0.016 | -0.2957 | No |
| 47 | CDCA5 | CDCA5 Entrez, Source | cell division cycle associated 5 | 8058 | 0.015 | -0.2957 | No |
| 48 | CENPO | CENPO Entrez, Source | centromere protein O | 8167 | 0.014 | -0.3010 | No |
| 49 | NCAPD2 | NCAPD2 Entrez, Source | non-SMC condensin I complex, subunit D2 | 8212 | 0.013 | -0.3028 | No |
| 50 | AKAP8L | AKAP8L Entrez, Source | A kinase (PRKA) anchor protein 8-like | 8217 | 0.013 | -0.3025 | No |
| 51 | AKAP8 | AKAP8 Entrez, Source | A kinase (PRKA) anchor protein 8 | 8265 | 0.012 | -0.3045 | No |
| 52 | H1FX | H1FX Entrez, Source | H1 histone family, member X | 8340 | 0.011 | -0.3081 | No |
| 53 | CENPH | CENPH Entrez, Source | centromere protein H | 8385 | 0.010 | -0.3100 | No |
| 54 | NAP1L1 | NAP1L1 Entrez, Source | nucleosome assembly protein 1-like 1 | 8480 | 0.009 | -0.3147 | No |
| 55 | PSME4 | PSME4 Entrez, Source | proteasome (prosome, macropain) activator subunit 4 | 8559 | 0.008 | -0.3186 | No |
| 56 | BRD2 | BRD2 Entrez, Source | bromodomain containing 2 | 8663 | 0.007 | -0.3240 | No |
| 57 | H3F3B | H3F3B Entrez, Source | H3 histone, family 3B (H3.3B) | 8669 | 0.007 | -0.3239 | No |
| 58 | ACIN1 | ACIN1 Entrez, Source | apoptotic chromatin condensation inducer 1 | 8696 | 0.006 | -0.3251 | No |
| 59 | NCAPH2 | NCAPH2 Entrez, Source | non-SMC condensin II complex, subunit H2 | 8767 | 0.005 | -0.3287 | No |
| 60 | NCAPG2 | NCAPG2 Entrez, Source | non-SMC condensin II complex, subunit G2 | 8819 | 0.005 | -0.3312 | No |
| 61 | CDAN1 | CDAN1 Entrez, Source | congenital dyserythropoietic anemia, type I | 8846 | 0.004 | -0.3325 | No |
| 62 | NCAPD3 | NCAPD3 Entrez, Source | non-SMC condensin II complex, subunit D3 | 8956 | 0.003 | -0.3383 | No |
| 63 | SET | SET Entrez, Source | SET translocation (myeloid leukemia-associated) | 9022 | 0.002 | -0.3418 | No |
| 64 | CTCF | CTCF Entrez, Source | CCCTC-binding factor (zinc finger protein) | 9087 | 0.001 | -0.3452 | No |
| 65 | SMARCA5 | SMARCA5 Entrez, Source | SWI/SNF related, matrix associated, actin dependent regulator of chromatin, subfamily a, member 5 | 9153 | -0.000 | -0.3488 | No |
| 66 | SMC4 | SMC4 Entrez, Source | structural maintenance of chromosomes 4 | 9300 | -0.002 | -0.3566 | No |
| 67 | AIFM1 | 9331 | -0.003 | -0.3582 | No | ||
| 68 | NAP1L4 | NAP1L4 Entrez, Source | nucleosome assembly protein 1-like 4 | 9352 | -0.003 | -0.3591 | No |
| 69 | HIST1H1D | HIST1H1D Entrez, Source | histone cluster 1, H1d | 9462 | -0.005 | -0.3648 | No |
| 70 | HJURP | 9474 | -0.005 | -0.3652 | No | ||
| 71 | CENPQ | CENPQ Entrez, Source | centromere protein Q | 9534 | -0.006 | -0.3682 | No |
| 72 | CHAF1A | CHAF1A Entrez, Source | chromatin assembly factor 1, subunit A (p150) | 9647 | -0.007 | -0.3740 | No |
| 73 | HMGA1 | HMGA1 Entrez, Source | high mobility group AT-hook 1 | 9651 | -0.008 | -0.3738 | No |
| 74 | RUVBL1 | RUVBL1 Entrez, Source | RuvB-like 1 (E. coli) | 9892 | -0.011 | -0.3864 | No |
| 75 | OIP5 | OIP5 Entrez, Source | Opa interacting protein 5 | 10058 | -0.013 | -0.3948 | No |
| 76 | RBBP7 | RBBP7 Entrez, Source | retinoblastoma binding protein 7 | 10101 | -0.014 | -0.3965 | No |
| 77 | MCM2 | MCM2 Entrez, Source | MCM2 minichromosome maintenance deficient 2, mitotin (S. cerevisiae) | 10199 | -0.016 | -0.4011 | No |
| 78 | NPM1 | NPM1 Entrez, Source | nucleophosmin (nucleolar phosphoprotein B23, numatrin) | 10367 | -0.018 | -0.4094 | No |
| 79 | H2AFX | H2AFX Entrez, Source | H2A histone family, member X | 10371 | -0.018 | -0.4088 | No |
| 80 | NOC2L | NOC2L Entrez, Source | nucleolar complex associated 2 homolog (S. cerevisiae) | 10391 | -0.019 | -0.4090 | No |
| 81 | NAP1L3 | NAP1L3 Entrez, Source | nucleosome assembly protein 1-like 3 | 10455 | -0.019 | -0.4116 | No |
| 82 | CENPP | CENPP Entrez, Source | centromere protein P | 10486 | -0.020 | -0.4124 | No |
| 83 | MIS18A | 10713 | -0.024 | -0.4236 | No | ||
| 84 | HIRA | HIRA Entrez, Source | HIR histone cell cycle regulation defective homolog A (S. cerevisiae) | 10955 | -0.029 | -0.4355 | No |
| 85 | HELLS | HELLS Entrez, Source | helicase, lymphoid-specific | 10964 | -0.029 | -0.4347 | No |
| 86 | HIST1H4E | HIST1H4E Entrez, Source | histone cluster 1, H4e | 11068 | -0.031 | -0.4390 | No |
| 87 | SRPK1 | SRPK1 Entrez, Source | SFRS protein kinase 1 | 11134 | -0.032 | -0.4411 | No |
| 88 | CENPM | CENPM Entrez, Source | centromere protein M | 11198 | -0.033 | -0.4431 | No |
| 89 | SMYD3 | SMYD3 Entrez, Source | SET and MYND domain containing 3 | 11209 | -0.033 | -0.4422 | No |
| 90 | CHMP1A | 11305 | -0.035 | -0.4459 | No | ||
| 91 | CENPA | CENPA Entrez, Source | centromere protein A | 11482 | -0.038 | -0.4538 | No |
| 92 | NAP1L2 | NAP1L2 Entrez, Source | nucleosome assembly protein 1-like 2 | 11735 | -0.043 | -0.4657 | No |
| 93 | KAT6B | 11783 | -0.044 | -0.4663 | No | ||
| 94 | RSF1 | RSF1 Entrez, Source | remodeling and spacing factor 1 | 11853 | -0.045 | -0.4681 | No |
| 95 | TSPYL4 | TSPYL4 Entrez, Source | TSPY-like 4 | 11862 | -0.046 | -0.4665 | No |
| 96 | NAA10 | 12099 | -0.051 | -0.4772 | No | ||
| 97 | NAA60 | 12379 | -0.058 | -0.4899 | No | ||
| 98 | ERN2 | ERN2 Entrez, Source | endoplasmic reticulum to nucleus signalling 2 | 12611 | -0.064 | -0.4996 | No |
| 99 | SMC2 | SMC2 Entrez, Source | structural maintenance of chromosomes 2 | 12638 | -0.065 | -0.4982 | No |
| 100 | HIST1H4H | HIST1H4H Entrez, Source | histone cluster 1, H4h | 12747 | -0.068 | -0.5011 | No |
| 101 | TNRC18 | TNRC18 Entrez, Source | trinucleotide repeat containing 18 | 12814 | -0.070 | -0.5016 | No |
| 102 | TP53 | TP53 Entrez, Source | tumor protein p53 (Li-Fraumeni syndrome) | 12831 | -0.070 | -0.4994 | No |
| 103 | ANP32B | ANP32B Entrez, Source | acidic (leucine-rich) nuclear phosphoprotein 32 family, member B | 13424 | -0.090 | -0.5278 | No |
| 104 | NAP1L6 | NAP1L6 Entrez, Source | nucleosome assembly protein 1-like 6 | 13471 | -0.091 | -0.5263 | No |
| 105 | HIST1H2BH | HIST1H2BH Entrez, Source | histone cluster 1, H2bh | 13513 | -0.093 | -0.5245 | No |
| 106 | CENPK | CENPK Entrez, Source | centromere protein K | 13793 | -0.106 | -0.5350 | No |
| 107 | CHD5 | CHD5 Entrez, Source | chromodomain helicase DNA binding protein 5 | 13878 | -0.110 | -0.5348 | No |
| 108 | ASF1A | ASF1A Entrez, Source | ASF1 anti-silencing function 1 homolog A (S. cerevisiae) | 14450 | -0.142 | -0.5597 | No |
| 109 | NAP1L5 | NAP1L5 Entrez, Source | nucleosome assembly protein 1-like 5 | 14791 | -0.164 | -0.5711 | No |
| 110 | TSSK6 | TSSK6 Entrez, Source | testis-specific serine kinase 6 | 15166 | -0.193 | -0.5830 | No |
| 111 | TSPY26P | 16015 | -0.278 | -0.6171 | Yes | ||
| 112 | HIST1H2BE | HIST1H2BE Entrez, Source | histone cluster 1, H2be | 16046 | -0.283 | -0.6063 | Yes |
| 113 | CDKN2A | CDKN2A Entrez, Source | cyclin-dependent kinase inhibitor 2A (melanoma, p16, inhibits CDK4) | 16088 | -0.288 | -0.5959 | Yes |
| 114 | HIST1H2BC | HIST1H2BC Entrez, Source | histone cluster 1, H2bc | 16155 | -0.298 | -0.5864 | Yes |
| 115 | HIST2H2BE | HIST2H2BE Entrez, Source | histone cluster 2, H2be | 16205 | -0.307 | -0.5757 | Yes |
| 116 | H1F0 | H1F0 Entrez, Source | H1 histone family, member 0 | 16514 | -0.352 | -0.5770 | Yes |
| 117 | SOX9 | SOX9 Entrez, Source | SRY (sex determining region Y)-box 9 (campomelic dysplasia, autosomal sex-reversal) | 16569 | -0.361 | -0.5641 | Yes |
| 118 | HIST1H1C | HIST1H1C Entrez, Source | histone cluster 1, H1c | 16788 | -0.397 | -0.5586 | Yes |
| 119 | TSPYL5 | TSPYL5 Entrez, Source | TSPY-like 5 | 16899 | -0.421 | -0.5461 | Yes |
| 120 | IPO4 | IPO4 Entrez, Source | importin 4 | 17360 | -0.537 | -0.5477 | Yes |
| 121 | SYCP1 | SYCP1 Entrez, Source | synaptonemal complex protein 1 | 17367 | -0.539 | -0.5244 | Yes |
| 122 | HIST1H1E | HIST1H1E Entrez, Source | histone cluster 1, H1e | 17659 | -0.633 | -0.5125 | Yes |
| 123 | HIST4H4 | HIST4H4 Entrez, Source | histone cluster 4, H4 | 17801 | -0.696 | -0.4896 | Yes |
| 124 | HIST1H2BK | HIST1H2BK Entrez, Source | histone cluster 1, H2bk | 17855 | -0.724 | -0.4608 | Yes |
| 125 | HIST1H2BG | HIST1H2BG Entrez, Source | histone cluster 1, H2bg | 17876 | -0.733 | -0.4297 | Yes |
| 126 | HIST1H2BD | HIST1H2BD Entrez, Source | histone cluster 1, H2bd | 18048 | -0.840 | -0.4022 | Yes |
| 127 | HIST1H3E | HIST1H3E Entrez, Source | histone cluster 1, H3e | 18150 | -0.927 | -0.3671 | Yes |
| 128 | HIST1H3H | HIST1H3H Entrez, Source | histone cluster 1, H3h | 18238 | -1.036 | -0.3264 | Yes |
| 129 | HIST1H2BO | HIST1H2BO Entrez, Source | histone cluster 1, H2bo | 18317 | -1.150 | -0.2802 | Yes |
| 130 | HIST3H2BB | HIST3H2BB Entrez, Source | histone cluster 3, H2bb | 18365 | -1.265 | -0.2273 | Yes |
| 131 | HIST1H2BN | HIST1H2BN Entrez, Source | histone cluster 1, H2bn | 18373 | -1.283 | -0.1714 | Yes |
| 132 | HIST1H4I | HIST1H4I Entrez, Source | histone cluster 1, H4i | 18380 | -1.300 | -0.1147 | Yes |
| 133 | HIST1H2BJ | HIST1H2BJ Entrez, Source | histone cluster 1, H2bj | 18393 | -1.337 | -0.0567 | Yes |
| 134 | HIST1H3G | HIST1H3G Entrez, Source | histone cluster 1, H3g | 18402 | -1.376 | 0.0032 | Yes |