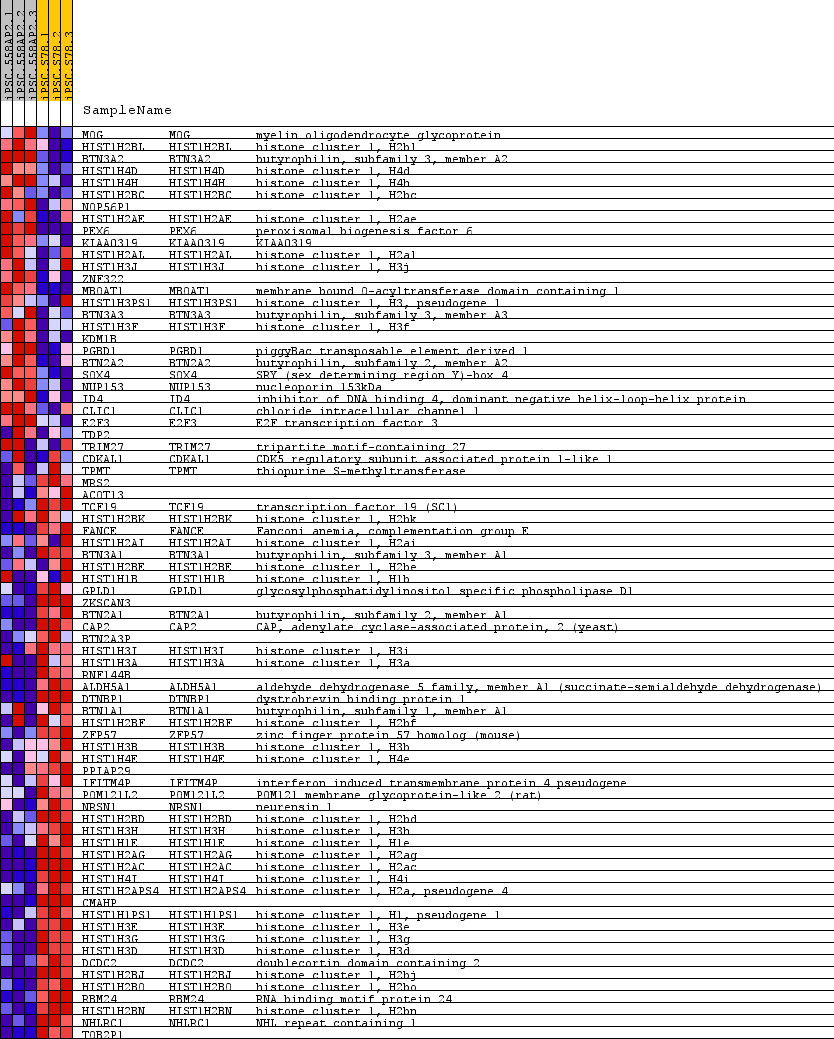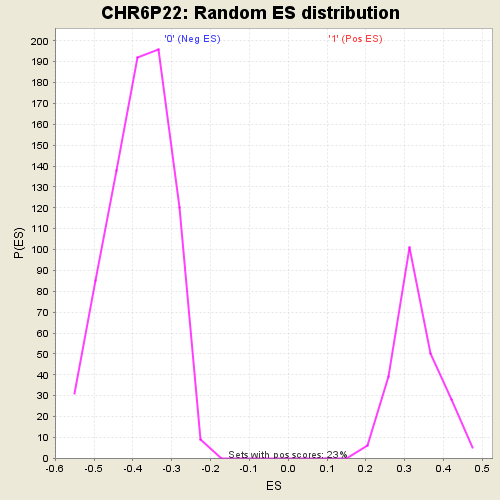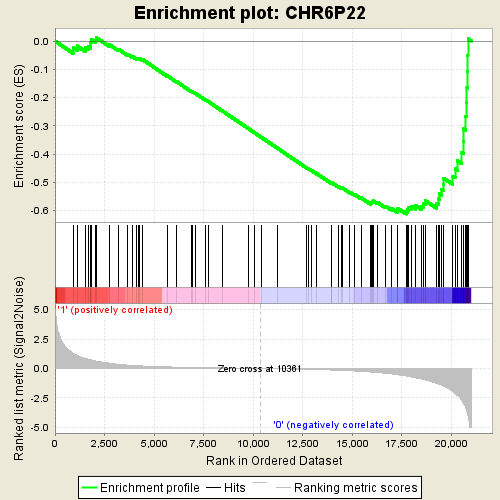
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | NS50832_collapsed_to_symbols.NS50832.cls #558AP2_versus_S78.NS50832.cls #558AP2_versus_S78_repos |
| Phenotype | NS50832.cls#558AP2_versus_S78_repos |
| Upregulated in class | 0 |
| GeneSet | CHR6P22 |
| Enrichment Score (ES) | -0.6119422 |
| Normalized Enrichment Score (NES) | -1.5946054 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.8619165 |
| FWER p-Value | 1.0 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | MOG | MOG Entrez, Source | myelin oligodendrocyte glycoprotein | 910 | 1.292 | -0.0243 | No |
| 2 | HIST1H2BL | HIST1H2BL Entrez, Source | histone cluster 1, H2bl | 1112 | 1.110 | -0.0174 | No |
| 3 | BTN3A2 | BTN3A2 Entrez, Source | butyrophilin, subfamily 3, member A2 | 1517 | 0.851 | -0.0241 | No |
| 4 | HIST1H4D | HIST1H4D Entrez, Source | histone cluster 1, H4d | 1674 | 0.784 | -0.0199 | No |
| 5 | HIST1H4H | HIST1H4H Entrez, Source | histone cluster 1, H4h | 1800 | 0.733 | -0.0150 | No |
| 6 | HIST1H2BC | HIST1H2BC Entrez, Source | histone cluster 1, H2bc | 1804 | 0.731 | -0.0042 | No |
| 7 | NOP56P1 | 1825 | 0.719 | 0.0055 | No | ||
| 8 | HIST1H2AE | HIST1H2AE Entrez, Source | histone cluster 1, H2ae | 2062 | 0.630 | 0.0036 | No |
| 9 | PEX6 | PEX6 Entrez, Source | peroxisomal biogenesis factor 6 | 2071 | 0.628 | 0.0125 | No |
| 10 | KIAA0319 | KIAA0319 Entrez, Source | KIAA0319 | 2740 | 0.455 | -0.0126 | No |
| 11 | HIST1H2AL | HIST1H2AL Entrez, Source | histone cluster 1, H2al | 3206 | 0.359 | -0.0295 | No |
| 12 | HIST1H3J | HIST1H3J Entrez, Source | histone cluster 1, H3j | 3655 | 0.289 | -0.0466 | No |
| 13 | ZNF322 | 3927 | 0.254 | -0.0558 | No | ||
| 14 | MBOAT1 | MBOAT1 Entrez, Source | membrane bound O-acyltransferase domain containing 1 | 4102 | 0.238 | -0.0606 | No |
| 15 | HIST1H3PS1 | HIST1H3PS1 Entrez, Source | histone cluster 1, H3, pseudogene 1 | 4212 | 0.228 | -0.0624 | No |
| 16 | BTN3A3 | BTN3A3 Entrez, Source | butyrophilin, subfamily 3, member A3 | 4262 | 0.224 | -0.0614 | No |
| 17 | HIST1H3F | HIST1H3F Entrez, Source | histone cluster 1, H3f | 4421 | 0.211 | -0.0659 | No |
| 18 | KDM1B | 5651 | 0.138 | -0.1226 | No | ||
| 19 | PGBD1 | PGBD1 Entrez, Source | piggyBac transposable element derived 1 | 6138 | 0.116 | -0.1441 | No |
| 20 | BTN2A2 | BTN2A2 Entrez, Source | butyrophilin, subfamily 2, member A2 | 6870 | 0.090 | -0.1777 | No |
| 21 | SOX4 | SOX4 Entrez, Source | SRY (sex determining region Y)-box 4 | 6940 | 0.088 | -0.1797 | No |
| 22 | NUP153 | NUP153 Entrez, Source | nucleoporin 153kDa | 7061 | 0.084 | -0.1842 | No |
| 23 | ID4 | ID4 Entrez, Source | inhibitor of DNA binding 4, dominant negative helix-loop-helix protein | 7605 | 0.069 | -0.2091 | No |
| 24 | CLIC1 | CLIC1 Entrez, Source | chloride intracellular channel 1 | 7736 | 0.066 | -0.2143 | No |
| 25 | E2F3 | E2F3 Entrez, Source | E2F transcription factor 3 | 8447 | 0.047 | -0.2476 | No |
| 26 | TDP2 | 9766 | 0.015 | -0.3104 | No | ||
| 27 | TRIM27 | TRIM27 Entrez, Source | tripartite motif-containing 27 | 10041 | 0.008 | -0.3233 | No |
| 28 | CDKAL1 | CDKAL1 Entrez, Source | CDK5 regulatory subunit associated protein 1-like 1 | 10389 | -0.001 | -0.3399 | No |
| 29 | TPMT | TPMT Entrez, Source | thiopurine S-methyltransferase | 11195 | -0.023 | -0.3781 | No |
| 30 | MRS2 | 12693 | -0.071 | -0.4486 | No | ||
| 31 | ACOT13 | 12804 | -0.077 | -0.4527 | No | ||
| 32 | TCF19 | TCF19 Entrez, Source | transcription factor 19 (SC1) | 12928 | -0.081 | -0.4574 | No |
| 33 | HIST1H2BK | HIST1H2BK Entrez, Source | histone cluster 1, H2bk | 13205 | -0.093 | -0.4692 | No |
| 34 | FANCE | FANCE Entrez, Source | Fanconi anemia, complementation group E | 13954 | -0.130 | -0.5030 | No |
| 35 | HIST1H2AI | HIST1H2AI Entrez, Source | histone cluster 1, H2ai | 13966 | -0.131 | -0.5016 | No |
| 36 | BTN3A1 | BTN3A1 Entrez, Source | butyrophilin, subfamily 3, member A1 | 14319 | -0.153 | -0.5162 | No |
| 37 | HIST1H2BE | HIST1H2BE Entrez, Source | histone cluster 1, H2be | 14463 | -0.164 | -0.5206 | No |
| 38 | HIST1H1B | HIST1H1B Entrez, Source | histone cluster 1, H1b | 14493 | -0.166 | -0.5195 | No |
| 39 | GPLD1 | GPLD1 Entrez, Source | glycosylphosphatidylinositol specific phospholipase D1 | 14874 | -0.195 | -0.5348 | No |
| 40 | ZKSCAN3 | 15102 | -0.211 | -0.5425 | No | ||
| 41 | BTN2A1 | BTN2A1 Entrez, Source | butyrophilin, subfamily 2, member A1 | 15435 | -0.242 | -0.5547 | No |
| 42 | CAP2 | CAP2 Entrez, Source | CAP, adenylate cyclase-associated protein, 2 (yeast) | 15903 | -0.295 | -0.5727 | No |
| 43 | BTN2A3P | 15983 | -0.303 | -0.5719 | No | ||
| 44 | HIST1H3I | HIST1H3I Entrez, Source | histone cluster 1, H3i | 16000 | -0.305 | -0.5682 | No |
| 45 | HIST1H3A | HIST1H3A Entrez, Source | histone cluster 1, H3a | 16043 | -0.311 | -0.5656 | No |
| 46 | RNF144B | 16242 | -0.339 | -0.5700 | No | ||
| 47 | ALDH5A1 | ALDH5A1 Entrez, Source | aldehyde dehydrogenase 5 family, member A1 (succinate-semialdehyde dehydrogenase) | 16671 | -0.404 | -0.5844 | No |
| 48 | DTNBP1 | DTNBP1 Entrez, Source | dystrobrevin binding protein 1 | 16952 | -0.451 | -0.5911 | No |
| 49 | BTN1A1 | BTN1A1 Entrez, Source | butyrophilin, subfamily 1, member A1 | 17253 | -0.522 | -0.5977 | No |
| 50 | HIST1H2BF | HIST1H2BF Entrez, Source | histone cluster 1, H2bf | 17290 | -0.531 | -0.5915 | No |
| 51 | ZFP57 | ZFP57 Entrez, Source | zinc finger protein 57 homolog (mouse) | 17718 | -0.640 | -0.6024 | Yes |
| 52 | HIST1H3B | HIST1H3B Entrez, Source | histone cluster 1, H3b | 17776 | -0.654 | -0.5954 | Yes |
| 53 | HIST1H4E | HIST1H4E Entrez, Source | histone cluster 1, H4e | 17843 | -0.676 | -0.5885 | Yes |
| 54 | PPIAP29 | 17990 | -0.720 | -0.5848 | Yes | ||
| 55 | IFITM4P | IFITM4P Entrez, Source | interferon induced transmembrane protein 4 pseudogene | 18200 | -0.786 | -0.5831 | Yes |
| 56 | POM121L2 | POM121L2 Entrez, Source | POM121 membrane glycoprotein-like 2 (rat) | 18492 | -0.891 | -0.5838 | Yes |
| 57 | NRSN1 | NRSN1 Entrez, Source | neurensin 1 | 18573 | -0.927 | -0.5738 | Yes |
| 58 | HIST1H2BD | HIST1H2BD Entrez, Source | histone cluster 1, H2bd | 18676 | -0.971 | -0.5643 | Yes |
| 59 | HIST1H3H | HIST1H3H Entrez, Source | histone cluster 1, H3h | 19248 | -1.251 | -0.5730 | Yes |
| 60 | HIST1H1E | HIST1H1E Entrez, Source | histone cluster 1, H1e | 19354 | -1.311 | -0.5585 | Yes |
| 61 | HIST1H2AG | HIST1H2AG Entrez, Source | histone cluster 1, H2ag | 19384 | -1.334 | -0.5401 | Yes |
| 62 | HIST1H2AC | HIST1H2AC Entrez, Source | histone cluster 1, H2ac | 19503 | -1.417 | -0.5246 | Yes |
| 63 | HIST1H4I | HIST1H4I Entrez, Source | histone cluster 1, H4i | 19573 | -1.473 | -0.5060 | Yes |
| 64 | HIST1H2APS4 | HIST1H2APS4 Entrez, Source | histone cluster 1, H2a, pseudogene 4 | 19588 | -1.487 | -0.4846 | Yes |
| 65 | CMAHP | 20070 | -1.961 | -0.4784 | Yes | ||
| 66 | HIST1H1PS1 | HIST1H1PS1 Entrez, Source | histone cluster 1, H1, pseudogene 1 | 20179 | -2.113 | -0.4522 | Yes |
| 67 | HIST1H3E | HIST1H3E Entrez, Source | histone cluster 1, H3e | 20289 | -2.284 | -0.4234 | Yes |
| 68 | HIST1H3G | HIST1H3G Entrez, Source | histone cluster 1, H3g | 20514 | -2.709 | -0.3939 | Yes |
| 69 | HIST1H3D | HIST1H3D Entrez, Source | histone cluster 1, H3d | 20594 | -2.926 | -0.3541 | Yes |
| 70 | DCDC2 | DCDC2 Entrez, Source | doublecortin domain containing 2 | 20601 | -2.952 | -0.3105 | Yes |
| 71 | HIST1H2BJ | HIST1H2BJ Entrez, Source | histone cluster 1, H2bj | 20719 | -3.326 | -0.2667 | Yes |
| 72 | HIST1H2BO | HIST1H2BO Entrez, Source | histone cluster 1, H2bo | 20735 | -3.400 | -0.2169 | Yes |
| 73 | RBM24 | RBM24 Entrez, Source | RNA binding motif protein 24 | 20777 | -3.632 | -0.1648 | Yes |
| 74 | HIST1H2BN | HIST1H2BN Entrez, Source | histone cluster 1, H2bn | 20809 | -3.847 | -0.1091 | Yes |
| 75 | NHLRC1 | NHLRC1 Entrez, Source | NHL repeat containing 1 | 20819 | -3.910 | -0.0514 | Yes |
| 76 | TOB2P1 | 20831 | -4.007 | 0.0076 | Yes |