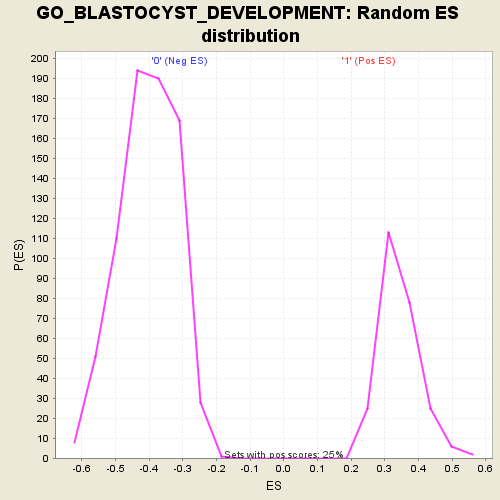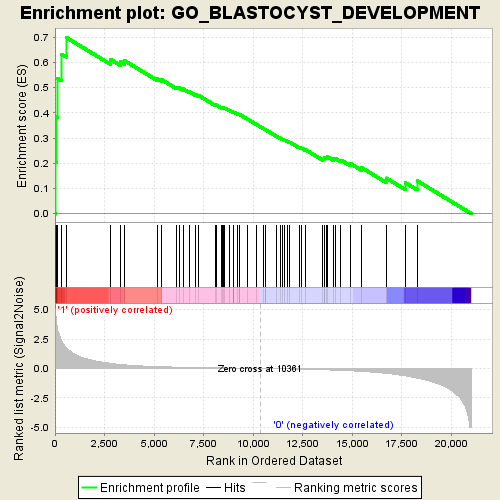
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | NS50832_collapsed_to_symbols.NS50832.cls #558AP2_versus_S78.NS50832.cls #558AP2_versus_S78_repos |
| Phenotype | NS50832.cls#558AP2_versus_S78_repos |
| Upregulated in class | 1 |
| GeneSet | GO_BLASTOCYST_DEVELOPMENT |
| Enrichment Score (ES) | 0.6993448 |
| Normalized Enrichment Score (NES) | 2.0277207 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.010260118 |
| FWER p-Value | 0.015 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | EOMES | EOMES Entrez, Source | eomesodermin homolog (Xenopus laevis) | 33 | 4.630 | 0.2024 | Yes |
| 2 | HAND1 | HAND1 Entrez, Source | heart and neural crest derivatives expressed 1 | 63 | 4.144 | 0.3837 | Yes |
| 3 | ZP3 | ZP3 Entrez, Source | zona pellucida glycoprotein 3 (sperm receptor) | 108 | 3.527 | 0.5370 | Yes |
| 4 | SOX17 | SOX17 Entrez, Source | SRY (sex determining region Y)-box 17 | 314 | 2.414 | 0.6335 | Yes |
| 5 | ETV2 | ETV2 Entrez, Source | ets variant gene 2 | 563 | 1.763 | 0.6993 | Yes |
| 6 | PRDM14 | PRDM14 Entrez, Source | PR domain containing 14 | 2779 | 0.444 | 0.6131 | No |
| 7 | GINS1 | GINS1 Entrez, Source | GINS complex subunit 1 (Psf1 homolog) | 3307 | 0.342 | 0.6030 | No |
| 8 | NODAL | NODAL Entrez, Source | nodal homolog (mouse) | 3518 | 0.308 | 0.6065 | No |
| 9 | DAD1 | DAD1 Entrez, Source | defender against cell death 1 | 5144 | 0.165 | 0.5362 | No |
| 10 | NASP | NASP Entrez, Source | nuclear autoantigenic sperm protein (histone-binding) | 5392 | 0.151 | 0.5310 | No |
| 11 | MED21 | 6119 | 0.117 | 0.5015 | No | ||
| 12 | RPL7L1 | RPL7L1 Entrez, Source | ribosomal protein L7-like 1 | 6262 | 0.112 | 0.4996 | No |
| 13 | NLE1 | NLE1 Entrez, Source | notchless homolog 1 (Drosophila) | 6454 | 0.104 | 0.4951 | No |
| 14 | TGFBR1 | TGFBR1 Entrez, Source | transforming growth factor, beta receptor I (activin A receptor type II-like kinase, 53kDa) | 6771 | 0.094 | 0.4841 | No |
| 15 | JUNB | JUNB Entrez, Source | jun B proto-oncogene | 7063 | 0.084 | 0.4739 | No |
| 16 | TET1 | 7233 | 0.080 | 0.4694 | No | ||
| 17 | RTF1 | RTF1 Entrez, Source | Rtf1, Paf1/RNA polymerase II complex component, homolog (S. cerevisiae) | 8093 | 0.056 | 0.4308 | No |
| 18 | PSMC4 | PSMC4 Entrez, Source | proteasome (prosome, macropain) 26S subunit, ATPase, 4 | 8132 | 0.055 | 0.4314 | No |
| 19 | GINS4 | GINS4 Entrez, Source | GINS complex subunit 4 (Sld5 homolog) | 8397 | 0.048 | 0.4209 | No |
| 20 | XAB2 | XAB2 Entrez, Source | XPA binding protein 2 | 8442 | 0.047 | 0.4209 | No |
| 21 | PRPF19 | PRPF19 Entrez, Source | PRP19/PSO4 pre-mRNA processing factor 19 homolog (S. cerevisiae) | 8513 | 0.046 | 0.4196 | No |
| 22 | RRP7A | 8567 | 0.044 | 0.4190 | No | ||
| 23 | ZNF830 | 8793 | 0.039 | 0.4100 | No | ||
| 24 | SALL4 | SALL4 Entrez, Source | sal-like 4 (Drosophila) | 8987 | 0.034 | 0.4023 | No |
| 25 | PSMC3 | PSMC3 Entrez, Source | proteasome (prosome, macropain) 26S subunit, ATPase, 3 | 9017 | 0.033 | 0.4023 | No |
| 26 | TAF8 | 9025 | 0.033 | 0.4035 | No | ||
| 27 | MFN2 | MFN2 Entrez, Source | mitofusin 2 | 9211 | 0.028 | 0.3958 | No |
| 28 | CTR9 | CTR9 Entrez, Source | Ctr9, Paf1/RNA polymerase II complex component, homolog (S. cerevisiae) | 9289 | 0.026 | 0.3933 | No |
| 29 | BYSL | BYSL Entrez, Source | bystin-like | 9316 | 0.025 | 0.3932 | No |
| 30 | SMARCB1 | SMARCB1 Entrez, Source | SWI/SNF related, matrix associated, actin dependent regulator of chromatin, subfamily b, member 1 | 9703 | 0.016 | 0.3755 | No |
| 31 | ELF3 | ELF3 Entrez, Source | E74-like factor 3 (ets domain transcription factor, epithelial-specific ) | 10166 | 0.005 | 0.3536 | No |
| 32 | CNOT2 | CNOT2 Entrez, Source | CCR4-NOT transcription complex, subunit 2 | 10490 | -0.003 | 0.3383 | No |
| 33 | SP3 | SP3 Entrez, Source | Sp3 transcription factor | 10596 | -0.007 | 0.3336 | No |
| 34 | NCAPG2 | NCAPG2 Entrez, Source | non-SMC condensin II complex, subunit G2 | 10627 | -0.008 | 0.3325 | No |
| 35 | RAD51B | 11184 | -0.022 | 0.3069 | No | ||
| 36 | LATS2 | LATS2 Entrez, Source | LATS, large tumor suppressor, homolog 2 (Drosophila) | 11346 | -0.026 | 0.3004 | No |
| 37 | CNOT3 | CNOT3 Entrez, Source | CCR4-NOT transcription complex, subunit 3 | 11481 | -0.030 | 0.2953 | No |
| 38 | SRF | SRF Entrez, Source | serum response factor (c-fos serum response element-binding transcription factor) | 11583 | -0.034 | 0.2920 | No |
| 39 | NDEL1 | NDEL1 Entrez, Source | nudE nuclear distribution gene E homolog like 1 (A. nidulans) | 11714 | -0.038 | 0.2875 | No |
| 40 | CUL3 | CUL3 Entrez, Source | cullin 3 | 11847 | -0.042 | 0.2830 | No |
| 41 | TJP1 | TJP1 Entrez, Source | tight junction protein 1 (zona occludens 1) | 12308 | -0.057 | 0.2636 | No |
| 42 | NEK2 | NEK2 Entrez, Source | NIMA (never in mitosis gene a)-related kinase 2 | 12428 | -0.061 | 0.2606 | No |
| 43 | BRCA2 | BRCA2 Entrez, Source | breast cancer 2, early onset | 12620 | -0.068 | 0.2545 | No |
| 44 | POU5F1 | POU5F1 Entrez, Source | POU domain, class 5, transcription factor 1 | 13506 | -0.106 | 0.2169 | No |
| 45 | PALB2 | PALB2 Entrez, Source | - | 13570 | -0.110 | 0.2187 | No |
| 46 | LATS1 | LATS1 Entrez, Source | LATS, large tumor suppressor, homolog 1 (Drosophila) | 13584 | -0.110 | 0.2229 | No |
| 47 | CDX2 | CDX2 Entrez, Source | caudal type homeobox transcription factor 2 | 13696 | -0.116 | 0.2227 | No |
| 48 | RBBP8 | RBBP8 Entrez, Source | retinoblastoma binding protein 8 | 13740 | -0.118 | 0.2259 | No |
| 49 | CCNB1IP1 | CCNB1IP1 Entrez, Source | cyclin B1 interacting protein 1 | 14041 | -0.135 | 0.2175 | No |
| 50 | ADA | ADA Entrez, Source | adenosine deaminase | 14124 | -0.140 | 0.2198 | No |
| 51 | NBN | NBN Entrez, Source | nibrin | 14422 | -0.161 | 0.2127 | No |
| 52 | SKIL | SKIL Entrez, Source | SKI-like | 14896 | -0.196 | 0.1987 | No |
| 53 | WDR74 | WDR74 Entrez, Source | WD repeat domain 74 | 15459 | -0.244 | 0.1826 | No |
| 54 | SBDS | SBDS Entrez, Source | Shwachman-Bodian-Diamond syndrome | 16698 | -0.407 | 0.1414 | No |
| 55 | HNF1B | 17674 | -0.626 | 0.1225 | No | ||
| 56 | CAPN2 | CAPN2 Entrez, Source | calpain 2, (m/II) large subunit | 18272 | -0.815 | 0.1299 | No |