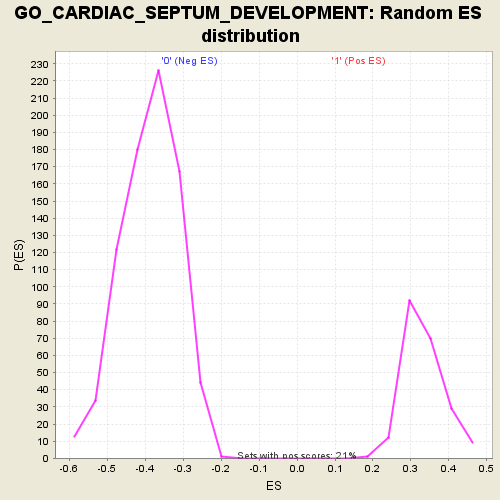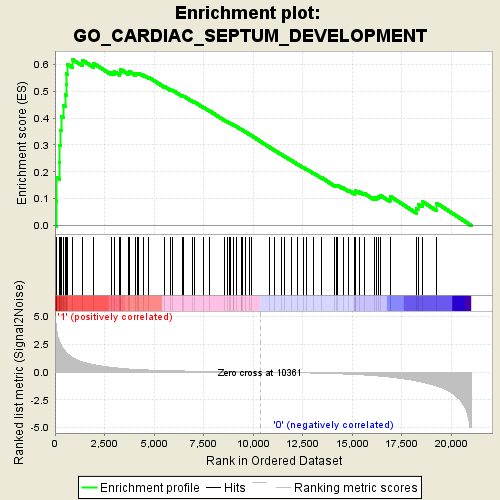
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | NS50832_collapsed_to_symbols.NS50832.cls #558AP2_versus_S78.NS50832.cls #558AP2_versus_S78_repos |
| Phenotype | NS50832.cls#558AP2_versus_S78_repos |
| Upregulated in class | 1 |
| GeneSet | GO_CARDIAC_SEPTUM_DEVELOPMENT |
| Enrichment Score (ES) | 0.6181427 |
| Normalized Enrichment Score (NES) | 1.8674704 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.07632094 |
| FWER p-Value | 0.466 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | HAND1 | HAND1 Entrez, Source | heart and neural crest derivatives expressed 1 | 63 | 4.144 | 0.0915 | Yes |
| 2 | MSX2 | MSX2 Entrez, Source | msh homeobox homolog 2 (Drosophila) | 79 | 3.875 | 0.1791 | Yes |
| 3 | PITX2 | PITX2 Entrez, Source | paired-like homeodomain transcription factor 2 | 223 | 2.814 | 0.2364 | Yes |
| 4 | GATA6 | GATA6 Entrez, Source | GATA binding protein 6 | 237 | 2.741 | 0.2982 | Yes |
| 5 | FGF8 | FGF8 Entrez, Source | fibroblast growth factor 8 (androgen-induced) | 277 | 2.560 | 0.3547 | Yes |
| 6 | PRDM1 | PRDM1 Entrez, Source | PR domain containing 1, with ZNF domain | 326 | 2.379 | 0.4067 | Yes |
| 7 | GJA5 | GJA5 Entrez, Source | gap junction protein, alpha 5, 40kDa (connexin 40) | 433 | 2.053 | 0.4484 | Yes |
| 8 | GATA3 | GATA3 Entrez, Source | GATA binding protein 3 | 505 | 1.890 | 0.4881 | Yes |
| 9 | ISL1 | ISL1 Entrez, Source | ISL1 transcription factor, LIM/homeodomain, (islet-1) | 558 | 1.772 | 0.5260 | Yes |
| 10 | TBX3 | TBX3 Entrez, Source | T-box 3 (ulnar mammary syndrome) | 569 | 1.753 | 0.5655 | Yes |
| 11 | BMP4 | BMP4 Entrez, Source | bone morphogenetic protein 4 | 620 | 1.680 | 0.6014 | Yes |
| 12 | TBX1 | TBX1 Entrez, Source | T-box 1 | 892 | 1.305 | 0.6181 | Yes |
| 13 | NKX2-5 | NKX2-5 Entrez, Source | NK2 transcription factor related, locus 5 (Drosophila) | 1370 | 0.926 | 0.6165 | No |
| 14 | FZD1 | FZD1 Entrez, Source | frizzled homolog 1 (Drosophila) | 1923 | 0.678 | 0.6055 | No |
| 15 | LRP2 | LRP2 Entrez, Source | low density lipoprotein-related protein 2 | 2840 | 0.429 | 0.5715 | No |
| 16 | MDM2 | MDM2 Entrez, Source | Mdm2, transformed 3T3 cell double minute 2, p53 binding protein (mouse) | 3005 | 0.396 | 0.5727 | No |
| 17 | NPHP3 | NPHP3 Entrez, Source | nephronophthisis 3 (adolescent) | 3270 | 0.347 | 0.5680 | No |
| 18 | SMAD6 | SMAD6 Entrez, Source | SMAD, mothers against DPP homolog 6 (Drosophila) | 3284 | 0.345 | 0.5752 | No |
| 19 | CITED2 | CITED2 Entrez, Source | Cbp/p300-interacting transactivator, with Glu/Asp-rich carboxy-terminal domain, 2 | 3316 | 0.341 | 0.5815 | No |
| 20 | FGFRL1 | FGFRL1 Entrez, Source | fibroblast growth factor receptor-like 1 | 3700 | 0.284 | 0.5697 | No |
| 21 | DHRS3 | DHRS3 Entrez, Source | dehydrogenase/reductase (SDR family) member 3 | 3752 | 0.277 | 0.5735 | No |
| 22 | HEG1 | HEG1 Entrez, Source | HEG homolog 1 (zebrafish) | 4054 | 0.242 | 0.5647 | No |
| 23 | ZFPM1 | ZFPM1 Entrez, Source | zinc finger protein, multitype 1 | 4136 | 0.235 | 0.5662 | No |
| 24 | RBM15 | RBM15 Entrez, Source | RNA binding motif protein 15 | 4219 | 0.227 | 0.5674 | No |
| 25 | FRS2 | FRS2 Entrez, Source | fibroblast growth factor receptor substrate 2 | 4456 | 0.208 | 0.5609 | No |
| 26 | GATA4 | GATA4 Entrez, Source | GATA binding protein 4 | 4707 | 0.191 | 0.5533 | No |
| 27 | NOTCH1 | NOTCH1 Entrez, Source | Notch homolog 1, translocation-associated (Drosophila) | 5526 | 0.144 | 0.5174 | No |
| 28 | NDST1 | NDST1 Entrez, Source | N-deacetylase/N-sulfotransferase (heparan glucosaminyl) 1 | 5841 | 0.129 | 0.5054 | No |
| 29 | SMO | SMO Entrez, Source | smoothened homolog (Drosophila) | 5916 | 0.125 | 0.5047 | No |
| 30 | NOTCH2 | NOTCH2 Entrez, Source | Notch homolog 2 (Drosophila) | 6422 | 0.105 | 0.4829 | No |
| 31 | SUFU | SUFU Entrez, Source | suppressor of fused homolog (Drosophila) | 6496 | 0.102 | 0.4818 | No |
| 32 | SOX4 | SOX4 Entrez, Source | SRY (sex determining region Y)-box 4 | 6940 | 0.088 | 0.4626 | No |
| 33 | PROX1 | PROX1 Entrez, Source | prospero-related homeobox 1 | 7029 | 0.085 | 0.4603 | No |
| 34 | NPRL3 | 7496 | 0.072 | 0.4397 | No | ||
| 35 | MYH10 | MYH10 Entrez, Source | myosin, heavy chain 10, non-muscle | 7808 | 0.064 | 0.4263 | No |
| 36 | PARVA | PARVA Entrez, Source | parvin, alpha | 8541 | 0.045 | 0.3923 | No |
| 37 | LUZP1 | LUZP1 Entrez, Source | leucine zipper protein 1 | 8698 | 0.042 | 0.3858 | No |
| 38 | CYR61 | CYR61 Entrez, Source | cysteine-rich, angiogenic inducer, 61 | 8787 | 0.039 | 0.3825 | No |
| 39 | DCTN5 | DCTN5 Entrez, Source | dynactin 5 (p25) | 8831 | 0.038 | 0.3813 | No |
| 40 | SALL4 | SALL4 Entrez, Source | sal-like 4 (Drosophila) | 8987 | 0.034 | 0.3746 | No |
| 41 | FGFR2 | FGFR2 Entrez, Source | ffer syndrome, Jackson-Weiss syndrome) | 9005 | 0.033 | 0.3746 | No |
| 42 | SMAD4 | SMAD4 Entrez, Source | SMAD, mothers against DPP homolog 4 (Drosophila) | 9149 | 0.030 | 0.3684 | No |
| 43 | TAB1 | 9389 | 0.024 | 0.3575 | No | ||
| 44 | VANGL2 | VANGL2 Entrez, Source | vang-like 2 (van gogh, Drosophila) | 9402 | 0.023 | 0.3575 | No |
| 45 | SMAD7 | SMAD7 Entrez, Source | SMAD, mothers against DPP homolog 7 (Drosophila) | 9450 | 0.022 | 0.3558 | No |
| 46 | SOX11 | SOX11 Entrez, Source | SRY (sex determining region Y)-box 11 | 9608 | 0.019 | 0.3487 | No |
| 47 | PTK7 | PTK7 Entrez, Source | PTK7 protein tyrosine kinase 7 | 9831 | 0.013 | 0.3383 | No |
| 48 | AP2B1 | AP2B1 Entrez, Source | adaptor-related protein complex 2, beta 1 subunit | 9897 | 0.011 | 0.3355 | No |
| 49 | PLXND1 | PLXND1 Entrez, Source | plexin D1 | 10793 | -0.012 | 0.2930 | No |
| 50 | SNX17 | SNX17 Entrez, Source | sorting nexin 17 | 11090 | -0.020 | 0.2793 | No |
| 51 | DNM2 | DNM2 Entrez, Source | dynamin 2 | 11432 | -0.029 | 0.2636 | No |
| 52 | ANK2 | ANK2 Entrez, Source | ankyrin 2, neuronal | 11580 | -0.034 | 0.2574 | No |
| 53 | SAV1 | SAV1 Entrez, Source | salvador homolog 1 (Drosophila) | 11928 | -0.045 | 0.2418 | No |
| 54 | MAML1 | MAML1 Entrez, Source | mastermind-like 1 (Drosophila) | 12246 | -0.055 | 0.2279 | No |
| 55 | PCSK5 | PCSK5 Entrez, Source | proprotein convertase subtilisin/kexin type 5 | 12530 | -0.065 | 0.2158 | No |
| 56 | DVL3 | DVL3 Entrez, Source | dishevelled, dsh homolog 3 (Drosophila) | 12687 | -0.071 | 0.2100 | No |
| 57 | SALL1 | SALL1 Entrez, Source | sal-like 1 (Drosophila) | 13043 | -0.086 | 0.1950 | No |
| 58 | WNT11 | WNT11 Entrez, Source | wingless-type MMTV integration site family, member 11 | 13442 | -0.104 | 0.1783 | No |
| 59 | ID2 | ID2 Entrez, Source | inhibitor of DNA binding 2, dominant negative helix-loop-helix protein | 14109 | -0.139 | 0.1496 | No |
| 60 | TRIP11 | TRIP11 Entrez, Source | thyroid hormone receptor interactor 11 | 14171 | -0.143 | 0.1500 | No |
| 61 | HEYL | HEYL Entrez, Source | hairy/enhancer-of-split related with YRPW motif-like | 14258 | -0.148 | 0.1492 | No |
| 62 | WNT5A | WNT5A Entrez, Source | wingless-type MMTV integration site family, member 5A | 14532 | -0.168 | 0.1400 | No |
| 63 | LMO4 | LMO4 Entrez, Source | LIM domain only 4 | 14794 | -0.190 | 0.1319 | No |
| 64 | EGLN1 | EGLN1 Entrez, Source | egl nine homolog 1 (C. elegans) | 15114 | -0.212 | 0.1215 | No |
| 65 | ACVR1 | ACVR1 Entrez, Source | activin A receptor, type I | 15133 | -0.214 | 0.1255 | No |
| 66 | HES1 | HES1 Entrez, Source | hairy and enhancer of split 1, (Drosophila) | 15160 | -0.217 | 0.1292 | No |
| 67 | LTBP1 | LTBP1 Entrez, Source | latent transforming growth factor beta binding protein 1 | 15360 | -0.236 | 0.1250 | No |
| 68 | JAG1 | JAG1 Entrez, Source | jagged 1 (Alagille syndrome) | 15590 | -0.257 | 0.1199 | No |
| 69 | RARB | RARB Entrez, Source | retinoic acid receptor, beta | 16104 | -0.320 | 0.1027 | No |
| 70 | RARA | RARA Entrez, Source | retinoic acid receptor, alpha | 16221 | -0.336 | 0.1048 | No |
| 71 | HEY2 | HEY2 Entrez, Source | hairy/enhancer-of-split related with YRPW motif 2 | 16302 | -0.348 | 0.1089 | No |
| 72 | CRELD1 | CRELD1 Entrez, Source | cysteine-rich with EGF-like domains 1 | 16420 | -0.365 | 0.1116 | No |
| 73 | FZD2 | FZD2 Entrez, Source | frizzled homolog 2 (Drosophila) | 16897 | -0.441 | 0.0989 | No |
| 74 | ZFPM2 | ZFPM2 Entrez, Source | zinc finger protein, multitype 2 | 16914 | -0.444 | 0.1083 | No |
| 75 | DAND5 | DAND5 Entrez, Source | DAN domain family, member 5 | 18241 | -0.803 | 0.0632 | No |
| 76 | ADAMTS6 | ADAMTS6 Entrez, Source | ADAM metallopeptidase with thrombospondin type 1 motif, 6 | 18310 | -0.831 | 0.0789 | No |
| 77 | HEY1 | HEY1 Entrez, Source | hairy/enhancer-of-split related with YRPW motif 1 | 18518 | -0.902 | 0.0895 | No |
| 78 | STRA6 | STRA6 Entrez, Source | stimulated by retinoic acid gene 6 homolog (mouse) | 19257 | -1.257 | 0.0829 | No |