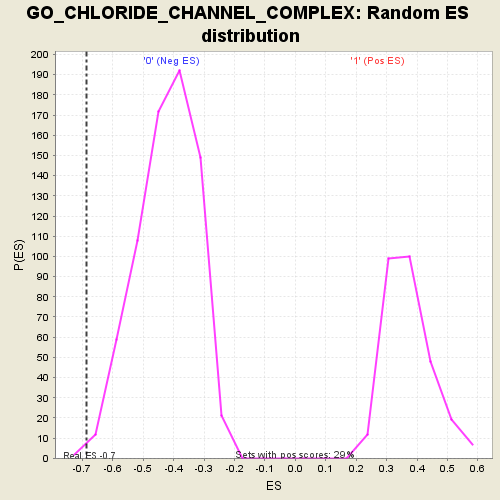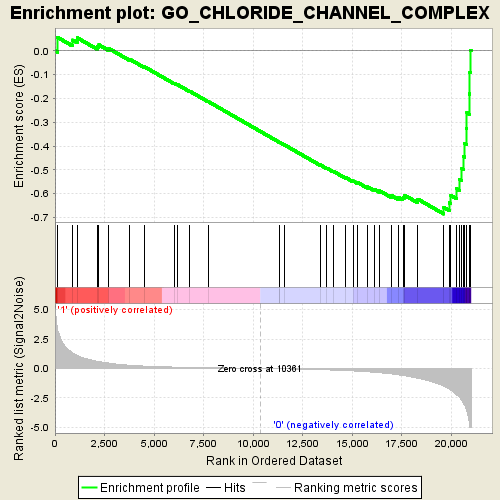
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | NS50832_collapsed_to_symbols.NS50832.cls #558AP2_versus_S78.NS50832.cls #558AP2_versus_S78_repos |
| Phenotype | NS50832.cls#558AP2_versus_S78_repos |
| Upregulated in class | 0 |
| GeneSet | GO_CHLORIDE_CHANNEL_COMPLEX |
| Enrichment Score (ES) | -0.6852404 |
| Normalized Enrichment Score (NES) | -1.6221493 |
| Nominal p-value | 0.002797203 |
| FDR q-value | 1.0 |
| FWER p-Value | 1.0 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | GABRD | GABRD Entrez, Source | gamma-aminobutyric acid (GABA) A receptor, delta | 133 | 3.320 | 0.0558 | No |
| 2 | GLRA4 | GLRA4 Entrez, Source | glycine receptor, alpha 4 | 875 | 1.324 | 0.0452 | No |
| 3 | FXYD3 | FXYD3 Entrez, Source | FXYD domain containing ion transport regulator 3 | 1115 | 1.106 | 0.0545 | No |
| 4 | CFTR | CFTR Entrez, Source | cystic fibrosis transmembrane conductance regulator (ATP-binding cassette sub-family C, member 7) | 2143 | 0.607 | 0.0169 | No |
| 5 | CLCNKA | CLCNKA Entrez, Source | chloride channel Ka | 2212 | 0.591 | 0.0247 | No |
| 6 | GLRB | GLRB Entrez, Source | glycine receptor, beta | 2718 | 0.461 | 0.0092 | No |
| 7 | TTYH1 | TTYH1 Entrez, Source | tweety homolog 1 (Drosophila) | 3746 | 0.278 | -0.0346 | No |
| 8 | GABRQ | GABRQ Entrez, Source | gamma-aminobutyric acid (GABA) receptor, theta | 4502 | 0.205 | -0.0668 | No |
| 9 | SLC26A6 | SLC26A6 Entrez, Source | solute carrier family 26, member 6 | 6015 | 0.121 | -0.1367 | No |
| 10 | GABRG3 | GABRG3 Entrez, Source | gamma-aminobutyric acid (GABA) A receptor, gamma 3 | 6163 | 0.115 | -0.1416 | No |
| 11 | ANO1 | 6775 | 0.094 | -0.1690 | No | ||
| 12 | CLIC1 | CLIC1 Entrez, Source | chloride intracellular channel 1 | 7736 | 0.066 | -0.2136 | No |
| 13 | TTYH3 | TTYH3 Entrez, Source | tweety homolog 3 (Drosophila) | 11310 | -0.025 | -0.3837 | No |
| 14 | CLCC1 | CLCC1 Entrez, Source | chloride channel CLIC-like 1 | 11323 | -0.026 | -0.3838 | No |
| 15 | GABRB3 | GABRB3 Entrez, Source | gamma-aminobutyric acid (GABA) A receptor, beta 3 | 11579 | -0.034 | -0.3953 | No |
| 16 | CLIC4 | CLIC4 Entrez, Source | chloride intracellular channel 4 | 13367 | -0.100 | -0.4787 | No |
| 17 | CLCN2 | CLCN2 Entrez, Source | chloride channel 2 | 13701 | -0.116 | -0.4924 | No |
| 18 | BEST2 | 14023 | -0.133 | -0.5053 | No | ||
| 19 | GABRA3 | GABRA3 Entrez, Source | gamma-aminobutyric acid (GABA) A receptor, alpha 3 | 14671 | -0.179 | -0.5328 | No |
| 20 | ANO6 | 15041 | -0.207 | -0.5465 | No | ||
| 21 | GABRP | GABRP Entrez, Source | gamma-aminobutyric acid (GABA) A receptor, pi | 15262 | -0.226 | -0.5528 | No |
| 22 | TTYH2 | TTYH2 Entrez, Source | tweety homolog 2 (Drosophila) | 15756 | -0.277 | -0.5711 | No |
| 23 | GABRA5 | GABRA5 Entrez, Source | gamma-aminobutyric acid (GABA) A receptor, alpha 5 | 16114 | -0.321 | -0.5822 | No |
| 24 | GABRE | GABRE Entrez, Source | gamma-aminobutyric acid (GABA) A receptor, epsilon | 16374 | -0.358 | -0.5878 | No |
| 25 | CLIC2 | CLIC2 Entrez, Source | chloride intracellular channel 2 | 16982 | -0.457 | -0.6083 | No |
| 26 | GABRR1 | GABRR1 Entrez, Source | gamma-aminobutyric acid (GABA) receptor, rho 1 | 17321 | -0.537 | -0.6143 | No |
| 27 | BEST4 | 17551 | -0.596 | -0.6141 | No | ||
| 28 | CLCNKB | CLCNKB Entrez, Source | chloride channel Kb | 17640 | -0.618 | -0.6067 | No |
| 29 | ANO2 | 18299 | -0.827 | -0.6227 | No | ||
| 30 | GLRA3 | GLRA3 Entrez, Source | glycine receptor, alpha 3 | 19611 | -1.500 | -0.6572 | Yes |
| 31 | GABRG2 | GABRG2 Entrez, Source | gamma-aminobutyric acid (GABA) A receptor, gamma 2 | 19888 | -1.739 | -0.6378 | Yes |
| 32 | CLIC6 | CLIC6 Entrez, Source | chloride intracellular channel 6 | 19964 | -1.826 | -0.6072 | Yes |
| 33 | CLIC5 | CLIC5 Entrez, Source | chloride intracellular channel 5 | 20263 | -2.235 | -0.5796 | Yes |
| 34 | GABRB1 | GABRB1 Entrez, Source | gamma-aminobutyric acid (GABA) A receptor, beta 1 | 20398 | -2.457 | -0.5400 | Yes |
| 35 | CLCN1 | CLCN1 Entrez, Source | chloride channel 1, skeletal muscle (Thomsen disease, autosomal dominant) | 20519 | -2.716 | -0.4948 | Yes |
| 36 | GABRA1 | GABRA1 Entrez, Source | gamma-aminobutyric acid (GABA) A receptor, alpha 1 | 20590 | -2.905 | -0.4438 | Yes |
| 37 | GABRG1 | GABRG1 Entrez, Source | gamma-aminobutyric acid (GABA) A receptor, gamma 1 | 20662 | -3.113 | -0.3889 | Yes |
| 38 | GABRB2 | GABRB2 Entrez, Source | gamma-aminobutyric acid (GABA) A receptor, beta 2 | 20759 | -3.525 | -0.3275 | Yes |
| 39 | FXYD1 | FXYD1 Entrez, Source | FXYD domain containing ion transport regulator 1 (phospholemman) | 20773 | -3.615 | -0.2605 | Yes |
| 40 | GABRA4 | GABRA4 Entrez, Source | gamma-aminobutyric acid (GABA) A receptor, alpha 4 | 20891 | -4.536 | -0.1811 | Yes |
| 41 | BEST1 | 20921 | -4.925 | -0.0903 | Yes | ||
| 42 | GABRA2 | GABRA2 Entrez, Source | gamma-aminobutyric acid (GABA) A receptor, alpha 2 | 20966 | -5.000 | 0.0012 | Yes |