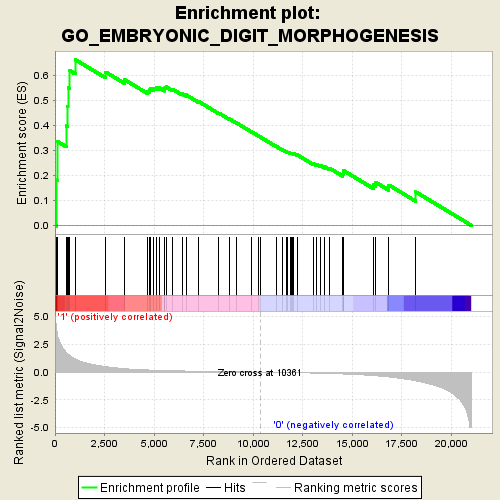
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | NS50832_collapsed_to_symbols.NS50832.cls #558AP2_versus_S78.NS50832.cls #558AP2_versus_S78_repos |
| Phenotype | NS50832.cls#558AP2_versus_S78_repos |
| Upregulated in class | 1 |
| GeneSet | GO_EMBRYONIC_DIGIT_MORPHOGENESIS |
| Enrichment Score (ES) | 0.66140276 |
| Normalized Enrichment Score (NES) | 1.8463506 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.092024155 |
| FWER p-Value | 0.602 |

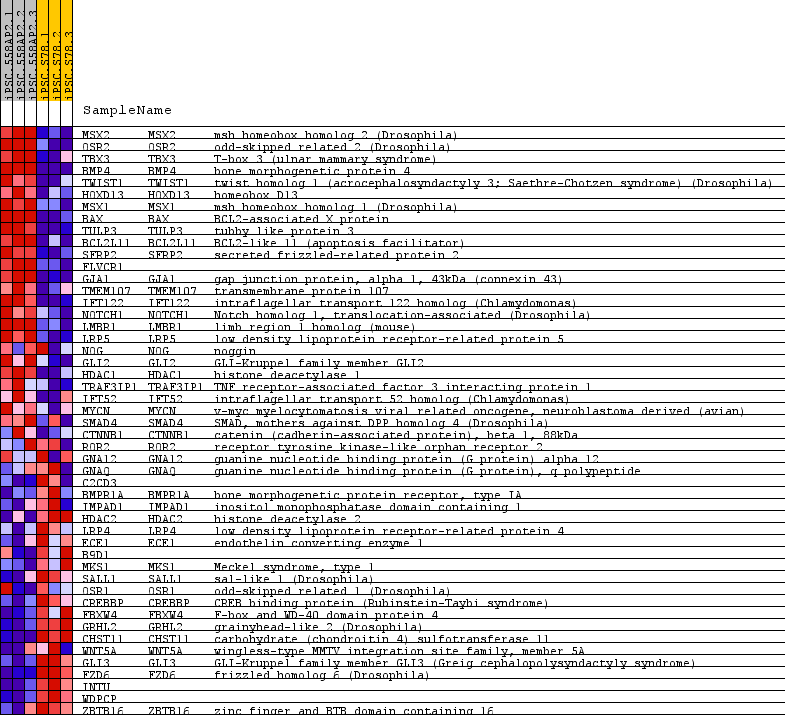
| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | MSX2 | MSX2 Entrez, Source | msh homeobox homolog 2 (Drosophila) | 79 | 3.875 | 0.1810 | Yes |
| 2 | OSR2 | OSR2 Entrez, Source | odd-skipped related 2 (Drosophila) | 138 | 3.259 | 0.3337 | Yes |
| 3 | TBX3 | TBX3 Entrez, Source | T-box 3 (ulnar mammary syndrome) | 569 | 1.753 | 0.3968 | Yes |
| 4 | BMP4 | BMP4 Entrez, Source | bone morphogenetic protein 4 | 620 | 1.680 | 0.4745 | Yes |
| 5 | TWIST1 | TWIST1 Entrez, Source | twist homolog 1 (acrocephalosyndactyly 3; Saethre-Chotzen syndrome) (Drosophila) | 653 | 1.619 | 0.5502 | Yes |
| 6 | HOXD13 | HOXD13 Entrez, Source | homeobox D13 | 745 | 1.501 | 0.6175 | Yes |
| 7 | MSX1 | MSX1 Entrez, Source | msh homeobox homolog 1 (Drosophila) | 1017 | 1.192 | 0.6614 | Yes |
| 8 | BAX | BAX Entrez, Source | BCL2-associated X protein | 2552 | 0.499 | 0.6120 | No |
| 9 | TULP3 | TULP3 Entrez, Source | tubby like protein 3 | 3496 | 0.310 | 0.5817 | No |
| 10 | BCL2L11 | BCL2L11 Entrez, Source | BCL2-like 11 (apoptosis facilitator) | 4661 | 0.194 | 0.5354 | No |
| 11 | SFRP2 | SFRP2 Entrez, Source | secreted frizzled-related protein 2 | 4747 | 0.188 | 0.5403 | No |
| 12 | FLVCR1 | 4807 | 0.185 | 0.5463 | No | ||
| 13 | GJA1 | GJA1 Entrez, Source | gap junction protein, alpha 1, 43kDa (connexin 43) | 4973 | 0.175 | 0.5468 | No |
| 14 | TMEM107 | TMEM107 Entrez, Source | transmembrane protein 107 | 5098 | 0.167 | 0.5488 | No |
| 15 | IFT122 | IFT122 Entrez, Source | intraflagellar transport 122 homolog (Chlamydomonas) | 5255 | 0.158 | 0.5489 | No |
| 16 | NOTCH1 | NOTCH1 Entrez, Source | Notch homolog 1, translocation-associated (Drosophila) | 5526 | 0.144 | 0.5429 | No |
| 17 | LMBR1 | LMBR1 Entrez, Source | limb region 1 homolog (mouse) | 5542 | 0.143 | 0.5489 | No |
| 18 | LRP5 | LRP5 Entrez, Source | low density lipoprotein receptor-related protein 5 | 5603 | 0.140 | 0.5527 | No |
| 19 | NOG | NOG Entrez, Source | noggin | 5925 | 0.125 | 0.5434 | No |
| 20 | GLI2 | GLI2 Entrez, Source | GLI-Kruppel family member GLI2 | 6412 | 0.105 | 0.5252 | No |
| 21 | HDAC1 | HDAC1 Entrez, Source | histone deacetylase 1 | 6612 | 0.099 | 0.5204 | No |
| 22 | TRAF3IP1 | TRAF3IP1 Entrez, Source | TNF receptor-associated factor 3 interacting protein 1 | 7251 | 0.079 | 0.4937 | No |
| 23 | IFT52 | IFT52 Entrez, Source | intraflagellar transport 52 homolog (Chlamydomonas) | 8262 | 0.052 | 0.4480 | No |
| 24 | MYCN | MYCN Entrez, Source | v-myc myelocytomatosis viral related oncogene, neuroblastoma derived (avian) | 8781 | 0.039 | 0.4251 | No |
| 25 | SMAD4 | SMAD4 Entrez, Source | SMAD, mothers against DPP homolog 4 (Drosophila) | 9149 | 0.030 | 0.4090 | No |
| 26 | CTNNB1 | CTNNB1 Entrez, Source | catenin (cadherin-associated protein), beta 1, 88kDa | 9928 | 0.011 | 0.3724 | No |
| 27 | ROR2 | ROR2 Entrez, Source | receptor tyrosine kinase-like orphan receptor 2 | 10254 | 0.003 | 0.3570 | No |
| 28 | GNA12 | GNA12 Entrez, Source | guanine nucleotide binding protein (G protein) alpha 12 | 10372 | -0.000 | 0.3514 | No |
| 29 | GNAQ | GNAQ Entrez, Source | guanine nucleotide binding protein (G protein), q polypeptide | 11182 | -0.022 | 0.3138 | No |
| 30 | C2CD3 | 11456 | -0.030 | 0.3022 | No | ||
| 31 | BMPR1A | BMPR1A Entrez, Source | bone morphogenetic protein receptor, type IA | 11652 | -0.036 | 0.2946 | No |
| 32 | IMPAD1 | IMPAD1 Entrez, Source | inositol monophosphatase domain containing 1 | 11740 | -0.039 | 0.2923 | No |
| 33 | HDAC2 | HDAC2 Entrez, Source | histone deacetylase 2 | 11899 | -0.044 | 0.2869 | No |
| 34 | LRP4 | LRP4 Entrez, Source | low density lipoprotein receptor-related protein 4 | 11936 | -0.045 | 0.2873 | No |
| 35 | ECE1 | ECE1 Entrez, Source | endothelin converting enzyme 1 | 11992 | -0.047 | 0.2869 | No |
| 36 | B9D1 | 12025 | -0.048 | 0.2877 | No | ||
| 37 | MKS1 | MKS1 Entrez, Source | Meckel syndrome, type 1 | 12212 | -0.054 | 0.2813 | No |
| 38 | SALL1 | SALL1 Entrez, Source | sal-like 1 (Drosophila) | 13043 | -0.086 | 0.2458 | No |
| 39 | OSR1 | OSR1 Entrez, Source | odd-skipped related 1 (Drosophila) | 13164 | -0.091 | 0.2444 | No |
| 40 | CREBBP | CREBBP Entrez, Source | CREB binding protein (Rubinstein-Taybi syndrome) | 13396 | -0.102 | 0.2383 | No |
| 41 | FBXW4 | FBXW4 Entrez, Source | F-box and WD-40 domain protein 4 | 13598 | -0.111 | 0.2340 | No |
| 42 | GRHL2 | GRHL2 Entrez, Source | grainyhead-like 2 (Drosophila) | 13840 | -0.124 | 0.2284 | No |
| 43 | CHST11 | CHST11 Entrez, Source | carbohydrate (chondroitin 4) sulfotransferase 11 | 14514 | -0.167 | 0.2042 | No |
| 44 | WNT5A | WNT5A Entrez, Source | wingless-type MMTV integration site family, member 5A | 14532 | -0.168 | 0.2114 | No |
| 45 | GLI3 | GLI3 Entrez, Source | GLI-Kruppel family member GLI3 (Greig cephalopolysyndactyly syndrome) | 14554 | -0.170 | 0.2185 | No |
| 46 | FZD6 | FZD6 Entrez, Source | frizzled homolog 6 (Drosophila) | 16057 | -0.313 | 0.1617 | No |
| 47 | INTU | 16170 | -0.329 | 0.1721 | No | ||
| 48 | WDPCP | 16837 | -0.431 | 0.1608 | No | ||
| 49 | ZBTB16 | ZBTB16 Entrez, Source | zinc finger and BTB domain containing 16 | 18199 | -0.786 | 0.1333 | No |