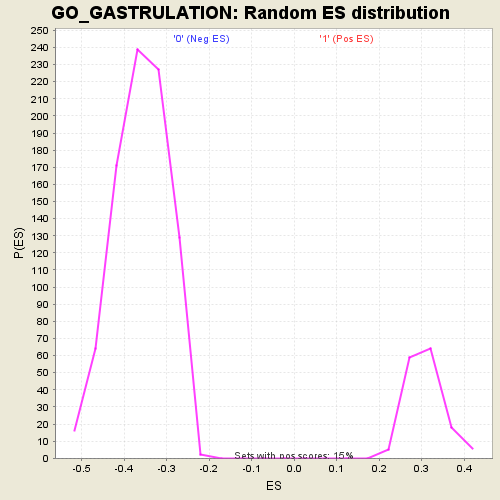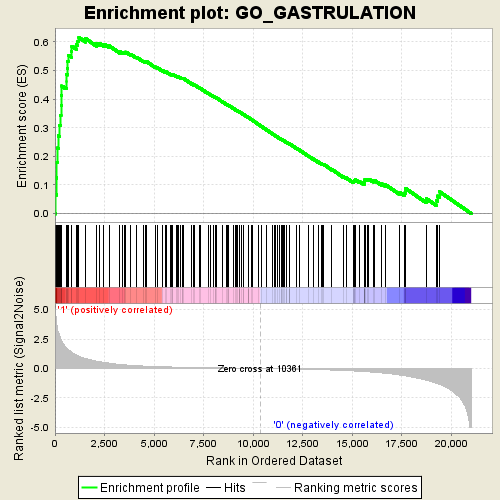
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | NS50832_collapsed_to_symbols.NS50832.cls #558AP2_versus_S78.NS50832.cls #558AP2_versus_S78_repos |
| Phenotype | NS50832.cls#558AP2_versus_S78_repos |
| Upregulated in class | 1 |
| GeneSet | GO_GASTRULATION |
| Enrichment Score (ES) | 0.6155582 |
| Normalized Enrichment Score (NES) | 1.9998205 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.0110979965 |
| FWER p-Value | 0.027 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | EOMES | EOMES Entrez, Source | eomesodermin homolog (Xenopus laevis) | 33 | 4.630 | 0.0659 | Yes |
| 2 | HAND1 | HAND1 Entrez, Source | heart and neural crest derivatives expressed 1 | 63 | 4.144 | 0.1249 | Yes |
| 3 | APLNR | 86 | 3.762 | 0.1787 | Yes | ||
| 4 | CER1 | CER1 Entrez, Source | cerberus 1, cysteine knot superfamily, homolog (Xenopus laevis) | 106 | 3.534 | 0.2294 | Yes |
| 5 | SIX2 | SIX2 Entrez, Source | sine oculis homeobox homolog 2 (Drosophila) | 161 | 3.101 | 0.2720 | Yes |
| 6 | GATA6 | GATA6 Entrez, Source | GATA binding protein 6 | 237 | 2.741 | 0.3083 | Yes |
| 7 | FGF8 | FGF8 Entrez, Source | fibroblast growth factor 8 (androgen-induced) | 277 | 2.560 | 0.3438 | Yes |
| 8 | COL8A1 | COL8A1 Entrez, Source | collagen, type VIII, alpha 1 | 302 | 2.477 | 0.3787 | Yes |
| 9 | SOX17 | SOX17 Entrez, Source | SRY (sex determining region Y)-box 17 | 314 | 2.414 | 0.4134 | Yes |
| 10 | CHRD | CHRD Entrez, Source | chordin | 343 | 2.329 | 0.4460 | Yes |
| 11 | ETV2 | ETV2 Entrez, Source | ets variant gene 2 | 563 | 1.763 | 0.4612 | Yes |
| 12 | MESP1 | MESP1 Entrez, Source | mesoderm posterior 1 homolog (mouse) | 565 | 1.763 | 0.4869 | Yes |
| 13 | BMP4 | BMP4 Entrez, Source | bone morphogenetic protein 4 | 620 | 1.680 | 0.5088 | Yes |
| 14 | T | T Entrez, Source | T, brachyury homolog (mouse) | 641 | 1.634 | 0.5316 | Yes |
| 15 | INHBA | INHBA Entrez, Source | inhibin, beta A (activin A, activin AB alpha polypeptide) | 661 | 1.602 | 0.5541 | Yes |
| 16 | LAMA3 | LAMA3 Entrez, Source | laminin, alpha 3 | 829 | 1.378 | 0.5662 | Yes |
| 17 | HHEX | HHEX Entrez, Source | homeobox, hematopoietically expressed | 846 | 1.359 | 0.5852 | Yes |
| 18 | VTN | VTN Entrez, Source | vitronectin | 1075 | 1.131 | 0.5908 | Yes |
| 19 | DKK1 | DKK1 Entrez, Source | dickkopf homolog 1 (Xenopus laevis) | 1149 | 1.071 | 0.6029 | Yes |
| 20 | GDF3 | GDF3 Entrez, Source | growth differentiation factor 3 | 1200 | 1.035 | 0.6156 | Yes |
| 21 | CRB2 | CRB2 Entrez, Source | crumbs homolog 2 (Drosophila) | 1536 | 0.841 | 0.6118 | No |
| 22 | TRIM15 | TRIM15 Entrez, Source | tripartite motif-containing 15 | 2096 | 0.621 | 0.5940 | No |
| 23 | MIXL1 | MIXL1 Entrez, Source | Mix1 homeobox-like 1 (Xenopus laevis) | 2253 | 0.577 | 0.5949 | No |
| 24 | FOXC1 | FOXC1 Entrez, Source | forkhead box C1 | 2465 | 0.519 | 0.5924 | No |
| 25 | ITGB4 | ITGB4 Entrez, Source | integrin, beta 4 | 2725 | 0.459 | 0.5866 | No |
| 26 | NPHP3 | NPHP3 Entrez, Source | nephronophthisis 3 (adolescent) | 3270 | 0.347 | 0.5656 | No |
| 27 | LAMB1 | LAMB1 Entrez, Source | laminin, beta 1 | 3409 | 0.326 | 0.5638 | No |
| 28 | NODAL | NODAL Entrez, Source | nodal homolog (mouse) | 3518 | 0.308 | 0.5631 | No |
| 29 | MMP2 | MMP2 Entrez, Source | matrix metallopeptidase 2 (gelatinase A, 72kDa gelatinase, 72kDa type IV collagenase) | 3574 | 0.299 | 0.5648 | No |
| 30 | TGFBR2 | TGFBR2 Entrez, Source | transforming growth factor, beta receptor II (70/80kDa) | 3829 | 0.267 | 0.5565 | No |
| 31 | COL7A1 | COL7A1 Entrez, Source | collagen, type VII, alpha 1 (epidermolysis bullosa, dystrophic, dominant and recessive) | 4096 | 0.239 | 0.5472 | No |
| 32 | FRS2 | FRS2 Entrez, Source | fibroblast growth factor receptor substrate 2 | 4456 | 0.208 | 0.5331 | No |
| 33 | COL4A2 | COL4A2 Entrez, Source | collagen, type IV, alpha 2 | 4555 | 0.201 | 0.5313 | No |
| 34 | EYA1 | EYA1 Entrez, Source | eyes absent homolog 1 (Drosophila) | 4596 | 0.198 | 0.5323 | No |
| 35 | SNAI1 | SNAI1 Entrez, Source | snail homolog 1 (Drosophila) | 5060 | 0.169 | 0.5125 | No |
| 36 | PAF1 | PAF1 Entrez, Source | Paf1, RNA polymerase II associated factor, homolog (S. cerevisiae) | 5153 | 0.163 | 0.5105 | No |
| 37 | DUSP4 | DUSP4 Entrez, Source | dual specificity phosphatase 4 | 5416 | 0.149 | 0.5001 | No |
| 38 | TWSG1 | TWSG1 Entrez, Source | twisted gastrulation homolog 1 (Drosophila) | 5553 | 0.142 | 0.4957 | No |
| 39 | LRP5 | LRP5 Entrez, Source | low density lipoprotein receptor-related protein 5 | 5603 | 0.140 | 0.4953 | No |
| 40 | MMP14 | MMP14 Entrez, Source | matrix metallopeptidase 14 (membrane-inserted) | 5833 | 0.129 | 0.4862 | No |
| 41 | AMOT | AMOT Entrez, Source | angiomotin | 5877 | 0.127 | 0.4860 | No |
| 42 | NOG | NOG Entrez, Source | noggin | 5925 | 0.125 | 0.4856 | No |
| 43 | EXT1 | EXT1 Entrez, Source | exostoses (multiple) 1 | 5942 | 0.124 | 0.4866 | No |
| 44 | BMPR2 | BMPR2 Entrez, Source | bone morphogenetic protein receptor, type II (serine/threonine kinase) | 6127 | 0.117 | 0.4795 | No |
| 45 | ITGA7 | ITGA7 Entrez, Source | integrin, alpha 7 | 6198 | 0.114 | 0.4778 | No |
| 46 | NF2 | NF2 Entrez, Source | neurofibromin 2 (bilateral acoustic neuroma) | 6239 | 0.113 | 0.4776 | No |
| 47 | DUSP2 | DUSP2 Entrez, Source | dual specificity phosphatase 2 | 6337 | 0.108 | 0.4745 | No |
| 48 | COL11A1 | COL11A1 Entrez, Source | collagen, type XI, alpha 1 | 6410 | 0.105 | 0.4726 | No |
| 49 | EPHA2 | EPHA2 Entrez, Source | EPH receptor A2 | 6431 | 0.105 | 0.4731 | No |
| 50 | LRP6 | LRP6 Entrez, Source | low density lipoprotein receptor-related protein 6 | 6462 | 0.104 | 0.4732 | No |
| 51 | OTX2 | OTX2 Entrez, Source | orthodenticle homolog 2 (Drosophila) | 6899 | 0.090 | 0.4536 | No |
| 52 | HMGA2 | HMGA2 Entrez, Source | high mobility group AT-hook 2 | 6995 | 0.086 | 0.4503 | No |
| 53 | RIC8A | RIC8A Entrez, Source | resistance to inhibitors of cholinesterase 8 homolog A (C. elegans) | 7050 | 0.085 | 0.4490 | No |
| 54 | SOX2 | SOX2 Entrez, Source | SRY (sex determining region Y)-box 2 | 7278 | 0.079 | 0.4392 | No |
| 55 | POFUT2 | POFUT2 Entrez, Source | protein O-fucosyltransferase 2 | 7324 | 0.077 | 0.4382 | No |
| 56 | ACVR2B | ACVR2B Entrez, Source | activin A receptor, type IIB | 7740 | 0.066 | 0.4193 | No |
| 57 | GPI | GPI Entrez, Source | glucose phosphate isomerase | 7816 | 0.063 | 0.4166 | No |
| 58 | BMP7 | BMP7 Entrez, Source | bone morphogenetic protein 7 (osteogenic protein 1) | 7994 | 0.059 | 0.4089 | No |
| 59 | RTF1 | RTF1 Entrez, Source | Rtf1, Paf1/RNA polymerase II complex component, homolog (S. cerevisiae) | 8093 | 0.056 | 0.4051 | No |
| 60 | SMAD3 | SMAD3 Entrez, Source | SMAD, mothers against DPP homolog 3 (Drosophila) | 8112 | 0.055 | 0.4050 | No |
| 61 | LEF1 | LEF1 Entrez, Source | lymphoid enhancer-binding factor 1 | 8157 | 0.054 | 0.4037 | No |
| 62 | EXT2 | EXT2 Entrez, Source | exostoses (multiple) 2 | 8424 | 0.048 | 0.3916 | No |
| 63 | HSBP1 | HSBP1 Entrez, Source | heat shock factor binding protein 1 | 8632 | 0.043 | 0.3823 | No |
| 64 | ATOH8 | ATOH8 Entrez, Source | atonal homolog 8 (Drosophila) | 8690 | 0.042 | 0.3802 | No |
| 65 | SYF2 | SYF2 Entrez, Source | SYF2 homolog, RNA splicing factor (S. cerevisiae) | 8730 | 0.041 | 0.3789 | No |
| 66 | LDB1 | LDB1 Entrez, Source | LIM domain binding 1 | 8736 | 0.041 | 0.3793 | No |
| 67 | FGFR2 | FGFR2 Entrez, Source | ffer syndrome, Jackson-Weiss syndrome) | 9005 | 0.033 | 0.3669 | No |
| 68 | RPS6 | RPS6 Entrez, Source | ribosomal protein S6 | 9126 | 0.030 | 0.3616 | No |
| 69 | SMAD4 | SMAD4 Entrez, Source | SMAD, mothers against DPP homolog 4 (Drosophila) | 9149 | 0.030 | 0.3610 | No |
| 70 | DVL1 | DVL1 Entrez, Source | dishevelled, dsh homolog 1 (Drosophila) | 9223 | 0.028 | 0.3579 | No |
| 71 | AXIN1 | AXIN1 Entrez, Source | axin 1 | 9278 | 0.026 | 0.3557 | No |
| 72 | CTR9 | CTR9 Entrez, Source | Ctr9, Paf1/RNA polymerase II complex component, homolog (S. cerevisiae) | 9289 | 0.026 | 0.3556 | No |
| 73 | DVL2 | DVL2 Entrez, Source | dishevelled, dsh homolog 2 (Drosophila) | 9296 | 0.026 | 0.3557 | No |
| 74 | VANGL2 | VANGL2 Entrez, Source | vang-like 2 (van gogh, Drosophila) | 9402 | 0.023 | 0.3510 | No |
| 75 | GSC | GSC Entrez, Source | goosecoid | 9498 | 0.021 | 0.3467 | No |
| 76 | EPB41L5 | EPB41L5 Entrez, Source | erythrocyte membrane protein band 4.1 like 5 | 9515 | 0.021 | 0.3463 | No |
| 77 | TXNRD1 | TXNRD1 Entrez, Source | thioredoxin reductase 1 | 9767 | 0.015 | 0.3345 | No |
| 78 | MEGF8 | MEGF8 Entrez, Source | multiple EGF-like-domains 8 | 9768 | 0.015 | 0.3347 | No |
| 79 | SFRP1 | SFRP1 Entrez, Source | secreted frizzled-related protein 1 | 9775 | 0.014 | 0.3346 | No |
| 80 | CTNNB1 | CTNNB1 Entrez, Source | catenin (cadherin-associated protein), beta 1, 88kDa | 9928 | 0.011 | 0.3275 | No |
| 81 | KIF16B | 9979 | 0.009 | 0.3252 | No | ||
| 82 | ITGB5 | ITGB5 Entrez, Source | integrin, beta 5 | 10236 | 0.003 | 0.3130 | No |
| 83 | ETS2 | ETS2 Entrez, Source | v-ets erythroblastosis virus E26 oncogene homolog 2 (avian) | 10434 | -0.002 | 0.3036 | No |
| 84 | ITGB1 | ITGB1 Entrez, Source | integrin, beta 1 (fibronectin receptor, beta polypeptide, antigen CD29 includes MDF2, MSK12) | 10671 | -0.009 | 0.2924 | No |
| 85 | HIRA | HIRA Entrez, Source | HIR histone cell cycle regulation defective homolog A (S. cerevisiae) | 10943 | -0.016 | 0.2796 | No |
| 86 | LEO1 | LEO1 Entrez, Source | Leo1, Paf1/RNA polymerase II complex component, homolog (S. cerevisiae) | 11059 | -0.019 | 0.2744 | No |
| 87 | EXOC4 | EXOC4 Entrez, Source | exocyst complex component 4 | 11103 | -0.020 | 0.2726 | No |
| 88 | UGDH | UGDH Entrez, Source | UDP-glucose dehydrogenase | 11204 | -0.023 | 0.2681 | No |
| 89 | ITGA2 | ITGA2 Entrez, Source | integrin, alpha 2 (CD49B, alpha 2 subunit of VLA-2 receptor) | 11322 | -0.026 | 0.2629 | No |
| 90 | FN1 | FN1 Entrez, Source | fibronectin 1 | 11419 | -0.028 | 0.2587 | No |
| 91 | MMP15 | MMP15 Entrez, Source | matrix metallopeptidase 15 (membrane-inserted) | 11444 | -0.029 | 0.2580 | No |
| 92 | ITGAV | ITGAV Entrez, Source | integrin, alpha V (vitronectin receptor, alpha polypeptide, antigen CD51) | 11482 | -0.030 | 0.2567 | No |
| 93 | WNT3 | WNT3 Entrez, Source | wingless-type MMTV integration site family, member 3 | 11541 | -0.032 | 0.2543 | No |
| 94 | SRF | SRF Entrez, Source | serum response factor (c-fos serum response element-binding transcription factor) | 11583 | -0.034 | 0.2529 | No |
| 95 | BMPR1A | BMPR1A Entrez, Source | bone morphogenetic protein receptor, type IA | 11652 | -0.036 | 0.2501 | No |
| 96 | SMAD1 | SMAD1 Entrez, Source | SMAD, mothers against DPP homolog 1 (Drosophila) | 11684 | -0.037 | 0.2492 | No |
| 97 | COL6A1 | COL6A1 Entrez, Source | collagen, type VI, alpha 1 | 11808 | -0.041 | 0.2439 | No |
| 98 | SMAD2 | SMAD2 Entrez, Source | SMAD, mothers against DPP homolog 2 (Drosophila) | 11831 | -0.042 | 0.2434 | No |
| 99 | CUL3 | CUL3 Entrez, Source | cullin 3 | 11847 | -0.042 | 0.2433 | No |
| 100 | DLD | DLD Entrez, Source | dihydrolipoamide dehydrogenase | 12200 | -0.053 | 0.2272 | No |
| 101 | ZBTB17 | ZBTB17 Entrez, Source | zinc finger and BTB domain containing 17 | 12307 | -0.057 | 0.2230 | No |
| 102 | PRKACA | PRKACA Entrez, Source | protein kinase, cAMP-dependent, catalytic, alpha | 12805 | -0.077 | 0.2003 | No |
| 103 | ITGA3 | ITGA3 Entrez, Source | integrin, alpha 3 (antigen CD49C, alpha 3 subunit of VLA-3 receptor) | 13024 | -0.086 | 0.1911 | No |
| 104 | CDC73 | CDC73 Entrez, Source | cell division cycle 73, Paf1/RNA polymerase II complex component, homolog (S. cerevisiae) | 13272 | -0.096 | 0.1806 | No |
| 105 | WNT11 | WNT11 Entrez, Source | wingless-type MMTV integration site family, member 11 | 13442 | -0.104 | 0.1740 | No |
| 106 | POU5F1 | POU5F1 Entrez, Source | POU domain, class 5, transcription factor 1 | 13506 | -0.106 | 0.1726 | No |
| 107 | PRKAR1A | PRKAR1A Entrez, Source | protein kinase, cAMP-dependent, regulatory, type I, alpha (tissue specific extinguisher 1) | 13557 | -0.109 | 0.1717 | No |
| 108 | MKKS | MKKS Entrez, Source | McKusick-Kaufman syndrome | 13969 | -0.131 | 0.1539 | No |
| 109 | WNT5A | WNT5A Entrez, Source | wingless-type MMTV integration site family, member 5A | 14532 | -0.168 | 0.1295 | No |
| 110 | LHX1 | LHX1 Entrez, Source | LIM homeobox 1 | 14679 | -0.181 | 0.1251 | No |
| 111 | COL12A1 | COL12A1 Entrez, Source | collagen, type XII, alpha 1 | 15039 | -0.207 | 0.1109 | No |
| 112 | SOX7 | SOX7 Entrez, Source | SRY (sex determining region Y)-box 7 | 15067 | -0.209 | 0.1126 | No |
| 113 | RNF2 | RNF2 Entrez, Source | ring finger protein 2 | 15097 | -0.211 | 0.1143 | No |
| 114 | ACVR2A | ACVR2A Entrez, Source | activin A receptor, type IIA | 15105 | -0.212 | 0.1171 | No |
| 115 | ACVR1 | ACVR1 Entrez, Source | activin A receptor, type I | 15133 | -0.214 | 0.1189 | No |
| 116 | TBX6 | TBX6 Entrez, Source | T-box 6 | 15374 | -0.237 | 0.1109 | No |
| 117 | KDM6B | 15593 | -0.258 | 0.1042 | No | ||
| 118 | KDM6A | 15605 | -0.259 | 0.1074 | No | ||
| 119 | DUSP1 | DUSP1 Entrez, Source | dual specificity phosphatase 1 | 15610 | -0.259 | 0.1110 | No |
| 120 | ITGB2 | ITGB2 Entrez, Source | integrin, beta 2 (complement component 3 receptor 3 and 4 subunit) | 15620 | -0.260 | 0.1143 | No |
| 121 | LAMB3 | LAMB3 Entrez, Source | laminin, beta 3 | 15629 | -0.261 | 0.1178 | No |
| 122 | FOXC2 | FOXC2 Entrez, Source | forkhead box C2 (MFH-1, mesenchyme forkhead 1) | 15651 | -0.264 | 0.1206 | No |
| 123 | DUSP5 | DUSP5 Entrez, Source | dual specificity phosphatase 5 | 15744 | -0.275 | 0.1202 | No |
| 124 | ARFRP1 | ARFRP1 Entrez, Source | ADP-ribosylation factor related protein 1 | 15834 | -0.286 | 0.1201 | No |
| 125 | MMP9 | MMP9 Entrez, Source | matrix metallopeptidase 9 (gelatinase B, 92kDa gelatinase, 92kDa type IV collagenase) | 16044 | -0.311 | 0.1146 | No |
| 126 | NANOG | NANOG Entrez, Source | Nanog homeobox | 16112 | -0.321 | 0.1161 | No |
| 127 | ITGA5 | ITGA5 Entrez, Source | integrin, alpha 5 (fibronectin receptor, alpha polypeptide) | 16484 | -0.377 | 0.1038 | No |
| 128 | EYA2 | EYA2 Entrez, Source | eyes absent homolog 2 (Drosophila) | 16678 | -0.405 | 0.1004 | No |
| 129 | NR4A3 | NR4A3 Entrez, Source | nuclear receptor subfamily 4, group A, member 3 | 17377 | -0.549 | 0.0750 | No |
| 130 | TAL1 | TAL1 Entrez, Source | T-cell acute lymphocytic leukemia 1 | 17629 | -0.614 | 0.0719 | No |
| 131 | HNF1B | 17674 | -0.626 | 0.0789 | No | ||
| 132 | ITGA4 | ITGA4 Entrez, Source | integrin, alpha 4 (antigen CD49D, alpha 4 subunit of VLA-4 receptor) | 17683 | -0.628 | 0.0877 | No |
| 133 | WLS | 18731 | -0.990 | 0.0519 | No | ||
| 134 | TLX2 | TLX2 Entrez, Source | T-cell leukemia homeobox 2 | 19232 | -1.243 | 0.0461 | No |
| 135 | WNT8B | WNT8B Entrez, Source | wingless-type MMTV integration site family, member 8B | 19299 | -1.275 | 0.0615 | No |
| 136 | KLF4 | KLF4 Entrez, Source | Kruppel-like factor 4 (gut) | 19398 | -1.343 | 0.0764 | No |