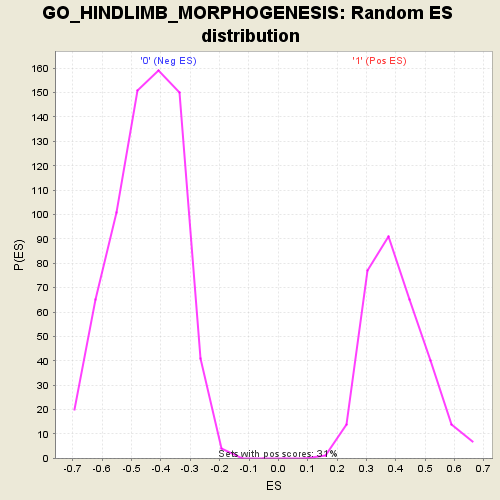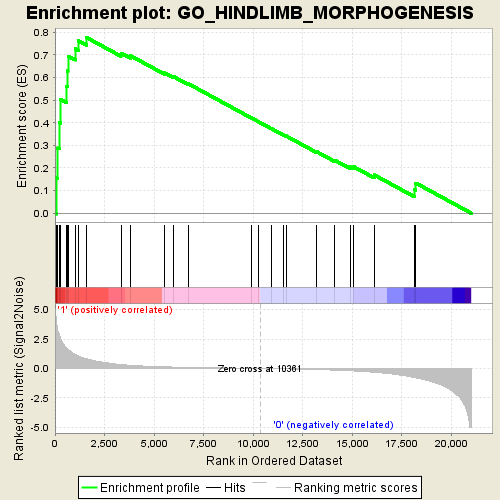
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | NS50832_collapsed_to_symbols.NS50832.cls #558AP2_versus_S78.NS50832.cls #558AP2_versus_S78_repos |
| Phenotype | NS50832.cls#558AP2_versus_S78_repos |
| Upregulated in class | 1 |
| GeneSet | GO_HINDLIMB_MORPHOGENESIS |
| Enrichment Score (ES) | 0.7772909 |
| Normalized Enrichment Score (NES) | 1.9441807 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.024362847 |
| FWER p-Value | 0.108 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | MSX2 | MSX2 Entrez, Source | msh homeobox homolog 2 (Drosophila) | 79 | 3.875 | 0.1564 | Yes |
| 2 | OSR2 | OSR2 Entrez, Source | odd-skipped related 2 (Drosophila) | 138 | 3.259 | 0.2883 | Yes |
| 3 | PITX2 | PITX2 Entrez, Source | paired-like homeodomain transcription factor 2 | 223 | 2.814 | 0.4006 | Yes |
| 4 | FGF8 | FGF8 Entrez, Source | fibroblast growth factor 8 (androgen-induced) | 277 | 2.560 | 0.5039 | Yes |
| 5 | TBX3 | TBX3 Entrez, Source | T-box 3 (ulnar mammary syndrome) | 569 | 1.753 | 0.5625 | Yes |
| 6 | BMP4 | BMP4 Entrez, Source | bone morphogenetic protein 4 | 620 | 1.680 | 0.6295 | Yes |
| 7 | TWIST1 | TWIST1 Entrez, Source | twist homolog 1 (acrocephalosyndactyly 3; Saethre-Chotzen syndrome) (Drosophila) | 653 | 1.619 | 0.6949 | Yes |
| 8 | MSX1 | MSX1 Entrez, Source | msh homeobox homolog 1 (Drosophila) | 1017 | 1.192 | 0.7269 | Yes |
| 9 | FGF4 | FGF4 Entrez, Source | fibroblast growth factor 4 (heparin secretory transforming protein 1, Kaposi sarcoma oncogene) | 1178 | 1.049 | 0.7626 | Yes |
| 10 | PTCH1 | PTCH1 Entrez, Source | patched homolog 1 (Drosophila) | 1582 | 0.820 | 0.7773 | Yes |
| 11 | RSPO2 | RSPO2 Entrez, Source | R-spondin 2 homolog (Xenopus laevis) | 3326 | 0.340 | 0.7082 | No |
| 12 | RPGRIP1L | 3815 | 0.269 | 0.6961 | No | ||
| 13 | NOTCH1 | NOTCH1 Entrez, Source | Notch homolog 1, translocation-associated (Drosophila) | 5526 | 0.144 | 0.6204 | No |
| 14 | GPC3 | GPC3 Entrez, Source | glypican 3 | 5978 | 0.123 | 0.6040 | No |
| 15 | GNAS | GNAS Entrez, Source | GNAS complex locus | 6705 | 0.096 | 0.5733 | No |
| 16 | CTNNB1 | CTNNB1 Entrez, Source | catenin (cadherin-associated protein), beta 1, 88kDa | 9928 | 0.011 | 0.4201 | No |
| 17 | MED1 | 10237 | 0.003 | 0.4055 | No | ||
| 18 | RARG | RARG Entrez, Source | retinoic acid receptor, gamma | 10913 | -0.015 | 0.3739 | No |
| 19 | WNT3 | WNT3 Entrez, Source | wingless-type MMTV integration site family, member 3 | 11541 | -0.032 | 0.3454 | No |
| 20 | BMPR1A | BMPR1A Entrez, Source | bone morphogenetic protein receptor, type IA | 11652 | -0.036 | 0.3416 | No |
| 21 | OSR1 | OSR1 Entrez, Source | odd-skipped related 1 (Drosophila) | 13164 | -0.091 | 0.2733 | No |
| 22 | AFF3 | AFF3 Entrez, Source | AF4/FMR2 family, member 3 | 14105 | -0.139 | 0.2342 | No |
| 23 | CHD7 | CHD7 Entrez, Source | chromodomain helicase DNA binding protein 7 | 14900 | -0.196 | 0.2045 | No |
| 24 | SALL3 | SALL3 Entrez, Source | sal-like 3 (Drosophila) | 15070 | -0.209 | 0.2051 | No |
| 25 | RARB | RARB Entrez, Source | retinoic acid receptor, beta | 16104 | -0.320 | 0.1690 | No |
| 26 | FMN1 | FMN1 Entrez, Source | formin 1 | 18127 | -0.763 | 0.1041 | No |
| 27 | ZBTB16 | ZBTB16 Entrez, Source | zinc finger and BTB domain containing 16 | 18199 | -0.786 | 0.1332 | No |