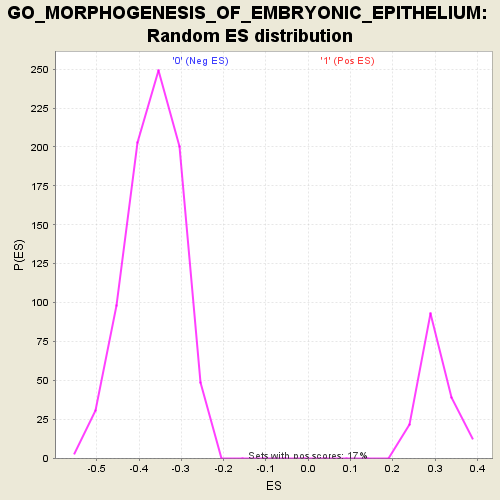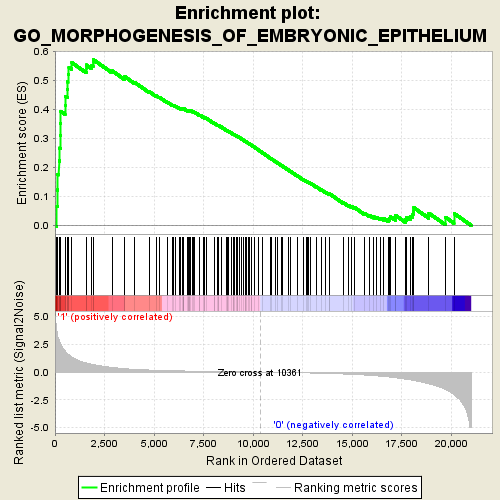
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | NS50832_collapsed_to_symbols.NS50832.cls #558AP2_versus_S78.NS50832.cls #558AP2_versus_S78_repos |
| Phenotype | NS50832.cls#558AP2_versus_S78_repos |
| Upregulated in class | 1 |
| GeneSet | GO_MORPHOGENESIS_OF_EMBRYONIC_EPITHELIUM |
| Enrichment Score (ES) | 0.57148993 |
| Normalized Enrichment Score (NES) | 1.89851 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.054105885 |
| FWER p-Value | 0.284 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | HAND1 | HAND1 Entrez, Source | heart and neural crest derivatives expressed 1 | 63 | 4.144 | 0.0663 | Yes |
| 2 | SIX1 | SIX1 Entrez, Source | sine oculis homeobox homolog 1 (Drosophila) | 113 | 3.493 | 0.1224 | Yes |
| 3 | IRX1 | IRX1 Entrez, Source | iroquois homeobox protein 1 | 134 | 3.309 | 0.1768 | Yes |
| 4 | GRHL3 | GRHL3 Entrez, Source | grainyhead-like 3 (Drosophila) | 198 | 2.948 | 0.2230 | Yes |
| 5 | ALX1 | 239 | 2.734 | 0.2668 | Yes | ||
| 6 | TFAP2A | TFAP2A Entrez, Source | transcription factor AP-2 alpha (activating enhancer binding protein 2 alpha) | 264 | 2.598 | 0.3092 | Yes |
| 7 | FGF8 | FGF8 Entrez, Source | fibroblast growth factor 8 (androgen-induced) | 277 | 2.560 | 0.3514 | Yes |
| 8 | WNT7B | WNT7B Entrez, Source | wingless-type MMTV integration site family, member 7B | 290 | 2.527 | 0.3931 | Yes |
| 9 | GATA3 | GATA3 Entrez, Source | GATA binding protein 3 | 505 | 1.890 | 0.4144 | Yes |
| 10 | IRX3 | IRX3 Entrez, Source | iroquois homeobox protein 3 | 515 | 1.867 | 0.4452 | Yes |
| 11 | BMP4 | BMP4 Entrez, Source | bone morphogenetic protein 4 | 620 | 1.680 | 0.4683 | Yes |
| 12 | T | T Entrez, Source | T, brachyury homolog (mouse) | 641 | 1.634 | 0.4947 | Yes |
| 13 | TWIST1 | TWIST1 Entrez, Source | twist homolog 1 (acrocephalosyndactyly 3; Saethre-Chotzen syndrome) (Drosophila) | 653 | 1.619 | 0.5213 | Yes |
| 14 | GDNF | GDNF Entrez, Source | glial cell derived neurotrophic factor | 699 | 1.554 | 0.5451 | Yes |
| 15 | HHEX | HHEX Entrez, Source | homeobox, hematopoietically expressed | 846 | 1.359 | 0.5608 | Yes |
| 16 | GREM1 | GREM1 Entrez, Source | gremlin 1, cysteine knot superfamily, homolog (Xenopus laevis) | 1577 | 0.822 | 0.5396 | Yes |
| 17 | PTCH1 | PTCH1 Entrez, Source | patched homolog 1 (Drosophila) | 1582 | 0.820 | 0.5531 | Yes |
| 18 | PCDH8 | PCDH8 Entrez, Source | protocadherin 8 | 1853 | 0.708 | 0.5520 | Yes |
| 19 | RET | RET Entrez, Source | ret proto-oncogene (multiple endocrine neoplasia and medullary thyroid carcinoma 1, Hirschsprung disease) | 1919 | 0.680 | 0.5603 | Yes |
| 20 | FZD1 | FZD1 Entrez, Source | frizzled homolog 1 (Drosophila) | 1923 | 0.678 | 0.5715 | Yes |
| 21 | IRX2 | IRX2 Entrez, Source | iroquois homeobox protein 2 | 2882 | 0.421 | 0.5326 | No |
| 22 | TULP3 | TULP3 Entrez, Source | tubby like protein 3 | 3496 | 0.310 | 0.5084 | No |
| 23 | NODAL | NODAL Entrez, Source | nodal homolog (mouse) | 3518 | 0.308 | 0.5126 | No |
| 24 | IFT172 | IFT172 Entrez, Source | intraflagellar transport 172 homolog (Chlamydomonas) | 4017 | 0.246 | 0.4928 | No |
| 25 | SFRP2 | SFRP2 Entrez, Source | secreted frizzled-related protein 2 | 4747 | 0.188 | 0.4610 | No |
| 26 | APAF1 | APAF1 Entrez, Source | apoptotic peptidase activating factor | 5094 | 0.168 | 0.4472 | No |
| 27 | IFT122 | IFT122 Entrez, Source | intraflagellar transport 122 homolog (Chlamydomonas) | 5255 | 0.158 | 0.4422 | No |
| 28 | TRIM71 | TRIM71 Entrez, Source | tripartite motif-containing 71 | 5656 | 0.137 | 0.4253 | No |
| 29 | NOG | NOG Entrez, Source | noggin | 5925 | 0.125 | 0.4146 | No |
| 30 | KAT2A | 5987 | 0.123 | 0.4137 | No | ||
| 31 | JAG2 | JAG2 Entrez, Source | jagged 2 | 6098 | 0.118 | 0.4104 | No |
| 32 | SEMA4C | SEMA4C Entrez, Source | sema domain, immunoglobulin domain (Ig), transmembrane domain (TM) and short cytoplasmic domain, (semaphorin) 4C | 6273 | 0.111 | 0.4039 | No |
| 33 | LAMA5 | LAMA5 Entrez, Source | laminin, alpha 5 | 6338 | 0.108 | 0.4027 | No |
| 34 | GLI2 | GLI2 Entrez, Source | GLI-Kruppel family member GLI2 | 6412 | 0.105 | 0.4010 | No |
| 35 | LRP6 | LRP6 Entrez, Source | low density lipoprotein receptor-related protein 6 | 6462 | 0.104 | 0.4003 | No |
| 36 | CELSR1 | CELSR1 Entrez, Source | cadherin, EGF LAG seven-pass G-type receptor 1 (flamingo homolog, Drosophila) | 6486 | 0.103 | 0.4009 | No |
| 37 | SUFU | SUFU Entrez, Source | suppressor of fused homolog (Drosophila) | 6496 | 0.102 | 0.4022 | No |
| 38 | SKI | SKI Entrez, Source | v-ski sarcoma viral oncogene homolog (avian) | 6684 | 0.096 | 0.3949 | No |
| 39 | SDC4 | SDC4 Entrez, Source | syndecan 4 (amphiglycan, ryudocan) | 6725 | 0.095 | 0.3946 | No |
| 40 | SCRIB | SCRIB Entrez, Source | scribbled homolog (Drosophila) | 6749 | 0.094 | 0.3950 | No |
| 41 | KIF20B | 6774 | 0.094 | 0.3954 | No | ||
| 42 | DEAF1 | DEAF1 Entrez, Source | deformed epidermal autoregulatory factor 1 (Drosophila) | 6779 | 0.093 | 0.3968 | No |
| 43 | SEC24B | SEC24B Entrez, Source | SEC24 related gene family, member B (S. cerevisiae) | 6819 | 0.092 | 0.3965 | No |
| 44 | SOX4 | SOX4 Entrez, Source | SRY (sex determining region Y)-box 4 | 6940 | 0.088 | 0.3922 | No |
| 45 | PLXNB2 | PLXNB2 Entrez, Source | plexin B2 | 6964 | 0.087 | 0.3926 | No |
| 46 | SHROOM3 | SHROOM3 Entrez, Source | shroom family member 3 | 7041 | 0.085 | 0.3903 | No |
| 47 | TSC2 | TSC2 Entrez, Source | tuberous sclerosis 2 | 7292 | 0.078 | 0.3797 | No |
| 48 | STK4 | STK4 Entrez, Source | serine/threonine kinase 4 | 7309 | 0.078 | 0.3802 | No |
| 49 | KDM2B | 7469 | 0.073 | 0.3738 | No | ||
| 50 | PHACTR4 | PHACTR4 Entrez, Source | phosphatase and actin regulator 4 | 7555 | 0.070 | 0.3709 | No |
| 51 | LHX2 | LHX2 Entrez, Source | LIM homeobox 2 | 7562 | 0.070 | 0.3718 | No |
| 52 | MIB1 | MIB1 Entrez, Source | mindbomb homolog 1 (Drosophila) | 7658 | 0.068 | 0.3684 | No |
| 53 | COBL | COBL Entrez, Source | cordon-bleu homolog (mouse) | 8034 | 0.057 | 0.3514 | No |
| 54 | TMED2 | TMED2 Entrez, Source | transmembrane emp24 domain trafficking protein 2 | 8061 | 0.057 | 0.3511 | No |
| 55 | FUZ | FUZ Entrez, Source | fuzzy homolog (Drosophila) | 8211 | 0.053 | 0.3448 | No |
| 56 | STK3 | STK3 Entrez, Source | serine/threonine kinase 3 (STE20 homolog, yeast) | 8227 | 0.052 | 0.3450 | No |
| 57 | IFT52 | IFT52 Entrez, Source | intraflagellar transport 52 homolog (Chlamydomonas) | 8262 | 0.052 | 0.3442 | No |
| 58 | GDF7 | GDF7 Entrez, Source | growth differentiation factor 7 | 8374 | 0.049 | 0.3397 | No |
| 59 | ARID1A | ARID1A Entrez, Source | AT rich interactive domain 1A (SWI- like) | 8667 | 0.042 | 0.3264 | No |
| 60 | TSC1 | TSC1 Entrez, Source | tuberous sclerosis 1 | 8697 | 0.042 | 0.3257 | No |
| 61 | LUZP1 | LUZP1 Entrez, Source | leucine zipper protein 1 | 8698 | 0.042 | 0.3264 | No |
| 62 | PFN1 | PFN1 Entrez, Source | profilin 1 | 8761 | 0.040 | 0.3241 | No |
| 63 | FZD3 | FZD3 Entrez, Source | frizzled homolog 3 (Drosophila) | 8890 | 0.036 | 0.3186 | No |
| 64 | SALL4 | SALL4 Entrez, Source | sal-like 4 (Drosophila) | 8987 | 0.034 | 0.3146 | No |
| 65 | FGFR2 | FGFR2 Entrez, Source | ffer syndrome, Jackson-Weiss syndrome) | 9005 | 0.033 | 0.3143 | No |
| 66 | MTHFD1L | MTHFD1L Entrez, Source | methylenetetrahydrofolate dehydrogenase (NADP+ dependent) 1-like | 9034 | 0.033 | 0.3135 | No |
| 67 | ARHGAP35 | 9152 | 0.030 | 0.3084 | No | ||
| 68 | RPS7 | RPS7 Entrez, Source | ribosomal protein S7 | 9184 | 0.029 | 0.3074 | No |
| 69 | DVL1 | DVL1 Entrez, Source | dishevelled, dsh homolog 1 (Drosophila) | 9223 | 0.028 | 0.3060 | No |
| 70 | DVL2 | DVL2 Entrez, Source | dishevelled, dsh homolog 2 (Drosophila) | 9296 | 0.026 | 0.3030 | No |
| 71 | VANGL2 | VANGL2 Entrez, Source | vang-like 2 (van gogh, Drosophila) | 9402 | 0.023 | 0.2984 | No |
| 72 | EPB41L5 | EPB41L5 Entrez, Source | erythrocyte membrane protein band 4.1 like 5 | 9515 | 0.021 | 0.2934 | No |
| 73 | SOX11 | SOX11 Entrez, Source | SRY (sex determining region Y)-box 11 | 9608 | 0.019 | 0.2893 | No |
| 74 | PRKACB | PRKACB Entrez, Source | protein kinase, cAMP-dependent, catalytic, beta | 9612 | 0.018 | 0.2894 | No |
| 75 | MTHFD1 | MTHFD1 Entrez, Source | methylenetetrahydrofolate dehydrogenase (NADP+ dependent) 1, methenyltetrahydrofolate cyclohydrolase, formyltetrahydrofolate synthetase | 9669 | 0.017 | 0.2870 | No |
| 76 | SFRP1 | SFRP1 Entrez, Source | secreted frizzled-related protein 1 | 9775 | 0.014 | 0.2822 | No |
| 77 | PTK7 | PTK7 Entrez, Source | PTK7 protein tyrosine kinase 7 | 9831 | 0.013 | 0.2798 | No |
| 78 | CTNNB1 | CTNNB1 Entrez, Source | catenin (cadherin-associated protein), beta 1, 88kDa | 9928 | 0.011 | 0.2754 | No |
| 79 | VASP | VASP Entrez, Source | vasodilator-stimulated phosphoprotein | 9932 | 0.011 | 0.2754 | No |
| 80 | ALDH1A3 | ALDH1A3 Entrez, Source | aldehyde dehydrogenase 1 family, member A3 | 10051 | 0.007 | 0.2699 | No |
| 81 | ST14 | ST14 Entrez, Source | suppression of tumorigenicity 14 (colon carcinoma) | 10279 | 0.002 | 0.2590 | No |
| 82 | SETD2 | SETD2 Entrez, Source | SET domain containing 2 | 10457 | -0.002 | 0.2506 | No |
| 83 | GRSF1 | GRSF1 Entrez, Source | G-rich RNA sequence binding factor 1 | 10853 | -0.013 | 0.2319 | No |
| 84 | RARG | RARG Entrez, Source | retinoic acid receptor, gamma | 10913 | -0.015 | 0.2293 | No |
| 85 | SPINT1 | SPINT1 Entrez, Source | serine peptidase inhibitor, Kunitz type 1 | 11139 | -0.021 | 0.2189 | No |
| 86 | IFT57 | IFT57 Entrez, Source | intraflagellar transport 57 homolog (Chlamydomonas) | 11198 | -0.023 | 0.2165 | No |
| 87 | TRAF6 | TRAF6 Entrez, Source | TNF receptor-associated factor 6 | 11434 | -0.029 | 0.2057 | No |
| 88 | STIL | STIL Entrez, Source | SCL/TAL1 interrupting locus | 11469 | -0.030 | 0.2046 | No |
| 89 | TCTN1 | 11756 | -0.039 | 0.1915 | No | ||
| 90 | HIF1A | HIF1A Entrez, Source | hypoxia-inducible factor 1, alpha subunit (basic helix-loop-helix transcription factor) | 11883 | -0.043 | 0.1862 | No |
| 91 | MKS1 | MKS1 Entrez, Source | Meckel syndrome, type 1 | 12212 | -0.054 | 0.1714 | No |
| 92 | BCL10 | BCL10 Entrez, Source | B-cell CLL/lymphoma 10 | 12551 | -0.066 | 0.1563 | No |
| 93 | DVL3 | DVL3 Entrez, Source | dishevelled, dsh homolog 3 (Drosophila) | 12687 | -0.071 | 0.1510 | No |
| 94 | BBS4 | BBS4 Entrez, Source | Bardet-Biedl syndrome 4 | 12758 | -0.074 | 0.1489 | No |
| 95 | PRKACA | PRKACA Entrez, Source | protein kinase, cAMP-dependent, catalytic, alpha | 12805 | -0.077 | 0.1480 | No |
| 96 | CC2D2A | 12876 | -0.079 | 0.1459 | No | ||
| 97 | OSR1 | OSR1 Entrez, Source | odd-skipped related 1 (Drosophila) | 13164 | -0.091 | 0.1337 | No |
| 98 | RALA | RALA Entrez, Source | v-ral simian leukemia viral oncogene homolog A (ras related) | 13422 | -0.103 | 0.1231 | No |
| 99 | SOX8 | SOX8 Entrez, Source | SRY (sex determining region Y)-box 8 | 13649 | -0.113 | 0.1142 | No |
| 100 | GLMN | GLMN Entrez, Source | glomulin, FKBP associated protein | 13837 | -0.123 | 0.1073 | No |
| 101 | GRHL2 | GRHL2 Entrez, Source | grainyhead-like 2 (Drosophila) | 13840 | -0.124 | 0.1093 | No |
| 102 | WNT5A | WNT5A Entrez, Source | wingless-type MMTV integration site family, member 5A | 14532 | -0.168 | 0.0790 | No |
| 103 | LMO4 | LMO4 Entrez, Source | LIM domain only 4 | 14794 | -0.190 | 0.0696 | No |
| 104 | MED12 | MED12 Entrez, Source | mediator of RNA polymerase II transcription, subunit 12 homolog (S. cerevisiae) | 14973 | -0.202 | 0.0645 | No |
| 105 | RDH10 | RDH10 Entrez, Source | retinol dehydrogenase 10 (all-trans) | 15101 | -0.211 | 0.0619 | No |
| 106 | KDM6A | 15605 | -0.259 | 0.0421 | No | ||
| 107 | LIAS | LIAS Entrez, Source | lipoic acid synthetase | 15863 | -0.289 | 0.0347 | No |
| 108 | FZD6 | FZD6 Entrez, Source | frizzled homolog 6 (Drosophila) | 16057 | -0.313 | 0.0306 | No |
| 109 | RARA | RARA Entrez, Source | retinoic acid receptor, alpha | 16221 | -0.336 | 0.0285 | No |
| 110 | ALDH1A2 | ALDH1A2 Entrez, Source | aldehyde dehydrogenase 1 family, member A2 | 16424 | -0.366 | 0.0249 | No |
| 111 | IPMK | IPMK Entrez, Source | inositol polyphosphate multikinase | 16586 | -0.392 | 0.0237 | No |
| 112 | ADM | ADM Entrez, Source | adrenomedullin | 16823 | -0.430 | 0.0196 | No |
| 113 | WNT4 | WNT4 Entrez, Source | wingless-type MMTV integration site family, member 4 | 16859 | -0.435 | 0.0252 | No |
| 114 | FZD2 | FZD2 Entrez, Source | frizzled homolog 2 (Drosophila) | 16897 | -0.441 | 0.0308 | No |
| 115 | ZEB2 | 17166 | -0.501 | 0.0263 | No | ||
| 116 | DLC1 | DLC1 Entrez, Source | deleted in liver cancer 1 | 17169 | -0.501 | 0.0346 | No |
| 117 | HNF1B | 17674 | -0.626 | 0.0210 | No | ||
| 118 | OVOL2 | OVOL2 Entrez, Source | ovo-like 2 (Drosophila) | 17739 | -0.646 | 0.0287 | No |
| 119 | WNT16 | WNT16 Entrez, Source | wingless-type MMTV integration site family, member 16 | 17920 | -0.699 | 0.0318 | No |
| 120 | CTHRC1 | CTHRC1 Entrez, Source | collagen triple helix repeat containing 1 | 18048 | -0.741 | 0.0381 | No |
| 121 | SOX9 | SOX9 Entrez, Source | SRY (sex determining region Y)-box 9 (campomelic dysplasia, autosomal sex-reversal) | 18063 | -0.745 | 0.0499 | No |
| 122 | PRICKLE1 | PRICKLE1 Entrez, Source | prickle homolog 1 (Drosophila) | 18098 | -0.755 | 0.0609 | No |
| 123 | FOLR1 | FOLR1 Entrez, Source | folate receptor 1 (adult) | 18852 | -1.050 | 0.0423 | No |
| 124 | TGFB1I1 | TGFB1I1 Entrez, Source | transforming growth factor beta 1 induced transcript 1 | 19684 | -1.569 | 0.0288 | No |
| 125 | VEGFC | VEGFC Entrez, Source | vascular endothelial growth factor C | 20126 | -2.023 | 0.0415 | No |