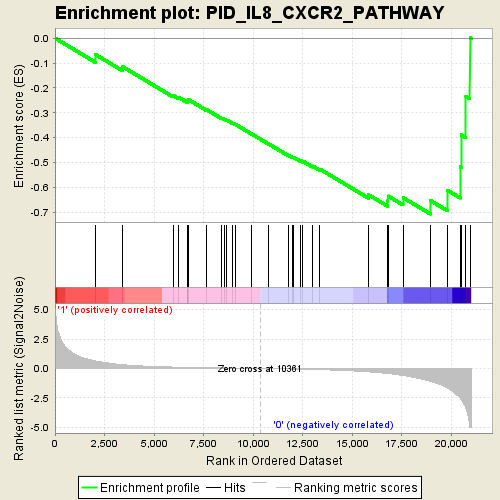
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | NS50832_collapsed_to_symbols.NS50832.cls #558AP2_versus_S78.NS50832.cls #558AP2_versus_S78_repos |
| Phenotype | NS50832.cls#558AP2_versus_S78_repos |
| Upregulated in class | 0 |
| GeneSet | PID_IL8_CXCR2_PATHWAY |
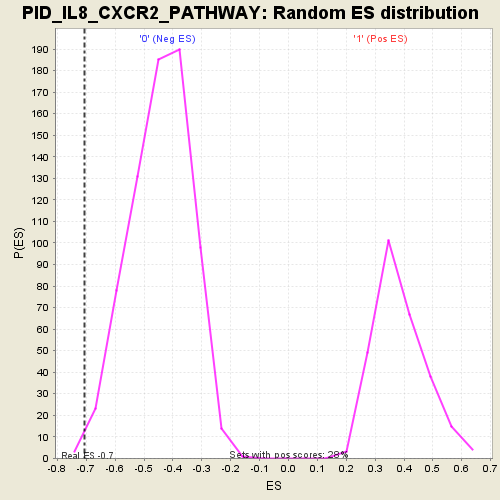
| Enrichment Score (ES) | -0.706935 |
| Normalized Enrichment Score (NES) | -1.5915701 |
| Nominal p-value | 0.004149378 |
| FDR q-value | 0.8120734 |
| FWER p-Value | 1.0 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | ARRB1 | ARRB1 Entrez, Source | arrestin, beta 1 | 2042 | 0.638 | -0.0659 | No |
| 2 | PRKCA | PRKCA Entrez, Source | protein kinase C, alpha | 3390 | 0.330 | -0.1139 | No |
| 3 | PLD2 | PLD2 Entrez, Source | phospholipase D2 | 5955 | 0.124 | -0.2301 | No |
| 4 | PPP2R1A | PPP2R1A Entrez, Source | protein phosphatase 2 (formerly 2A), regulatory subunit A (PR 65), alpha isoform | 6248 | 0.112 | -0.2385 | No |
| 5 | LYN | LYN Entrez, Source | v-yes-1 Yamaguchi sarcoma viral related oncogene homolog | 6659 | 0.097 | -0.2533 | No |
| 6 | DNM1 | DNM1 Entrez, Source | dynamin 1 | 6699 | 0.096 | -0.2504 | No |
| 7 | RAB11A | RAB11A Entrez, Source | RAB11A, member RAS oncogene family | 6724 | 0.095 | -0.2469 | No |
| 8 | RAB7A | 7631 | 0.069 | -0.2867 | No | ||
| 9 | DOCK2 | DOCK2 Entrez, Source | dedicator of cytokinesis 2 | 8416 | 0.048 | -0.3217 | No |
| 10 | RAC2 | RAC2 Entrez, Source | ras-related C3 botulinum toxin substrate 2 (rho family, small GTP binding protein Rac2) | 8554 | 0.045 | -0.3260 | No |
| 11 | ARRB2 | ARRB2 Entrez, Source | arrestin, beta 2 | 8665 | 0.042 | -0.3292 | No |
| 12 | GNAI2 | GNAI2 Entrez, Source | guanine nucleotide binding protein (G protein), alpha inhibiting activity polypeptide 2 | 8955 | 0.035 | -0.3413 | No |
| 13 | PLCB3 | PLCB3 Entrez, Source | phospholipase C, beta 3 (phosphatidylinositol-specific) | 9093 | 0.031 | -0.3463 | No |
| 14 | VASP | VASP Entrez, Source | vasodilator-stimulated phosphoprotein | 9932 | 0.011 | -0.3857 | No |
| 15 | PPP2CA | PPP2CA Entrez, Source | protein phosphatase 2 (formerly 2A), catalytic subunit, alpha isoform | 10775 | -0.011 | -0.4253 | No |
| 16 | AKT1 | AKT1 Entrez, Source | v-akt murine thymoma viral oncogene homolog 1 | 11777 | -0.040 | -0.4711 | No |
| 17 | GNG2 | GNG2 Entrez, Source | guanine nucleotide binding protein (G protein), gamma 2 | 11973 | -0.046 | -0.4781 | No |
| 18 | PDPK1 | PDPK1 Entrez, Source | 3-phosphoinositide dependent protein kinase-1 | 12022 | -0.048 | -0.4781 | No |
| 19 | PRKCG | PRKCG Entrez, Source | protein kinase C, gamma | 12387 | -0.060 | -0.4925 | No |
| 20 | GNB1 | GNB1 Entrez, Source | guanine nucleotide binding protein (G protein), beta polypeptide 1 | 12505 | -0.064 | -0.4949 | No |
| 21 | CBL | CBL Entrez, Source | Cas-Br-M (murine) ecotropic retroviral transforming sequence | 12990 | -0.084 | -0.5139 | No |
| 22 | RAB5A | RAB5A Entrez, Source | RAB5A, member RAS oncogene family | 13347 | -0.100 | -0.5259 | No |
| 23 | PLCB1 | PLCB1 Entrez, Source | phospholipase C, beta 1 (phosphoinositide-specific) | 15828 | -0.285 | -0.6302 | No |
| 24 | PLCB2 | PLCB2 Entrez, Source | phospholipase C, beta 2 | 16774 | -0.421 | -0.6545 | Yes |
| 25 | PRKCB | 16795 | -0.424 | -0.6345 | Yes | ||
| 26 | ELMO1 | ELMO1 Entrez, Source | engulfment and cell motility 1 | 17553 | -0.597 | -0.6411 | Yes |
| 27 | GNA14 | GNA14 Entrez, Source | guanine nucleotide binding protein (G protein), alpha 14 | 18934 | -1.095 | -0.6529 | Yes |
| 28 | FGR | FGR Entrez, Source | Gardner-Rasheed feline sarcoma viral (v-fgr) oncogene homolog | 19802 | -1.665 | -0.6120 | Yes |
| 29 | GNA15 | GNA15 Entrez, Source | guanine nucleotide binding protein (G protein), alpha 15 (Gq class) | 20445 | -2.544 | -0.5170 | Yes |
| 30 | PIK3R6 | 20484 | -2.635 | -0.3887 | Yes | ||
| 31 | CXCR2 | 20725 | -3.357 | -0.2343 | Yes | ||
| 32 | HCK | HCK Entrez, Source | hemopoietic cell kinase | 20931 | -5.000 | 0.0029 | Yes |