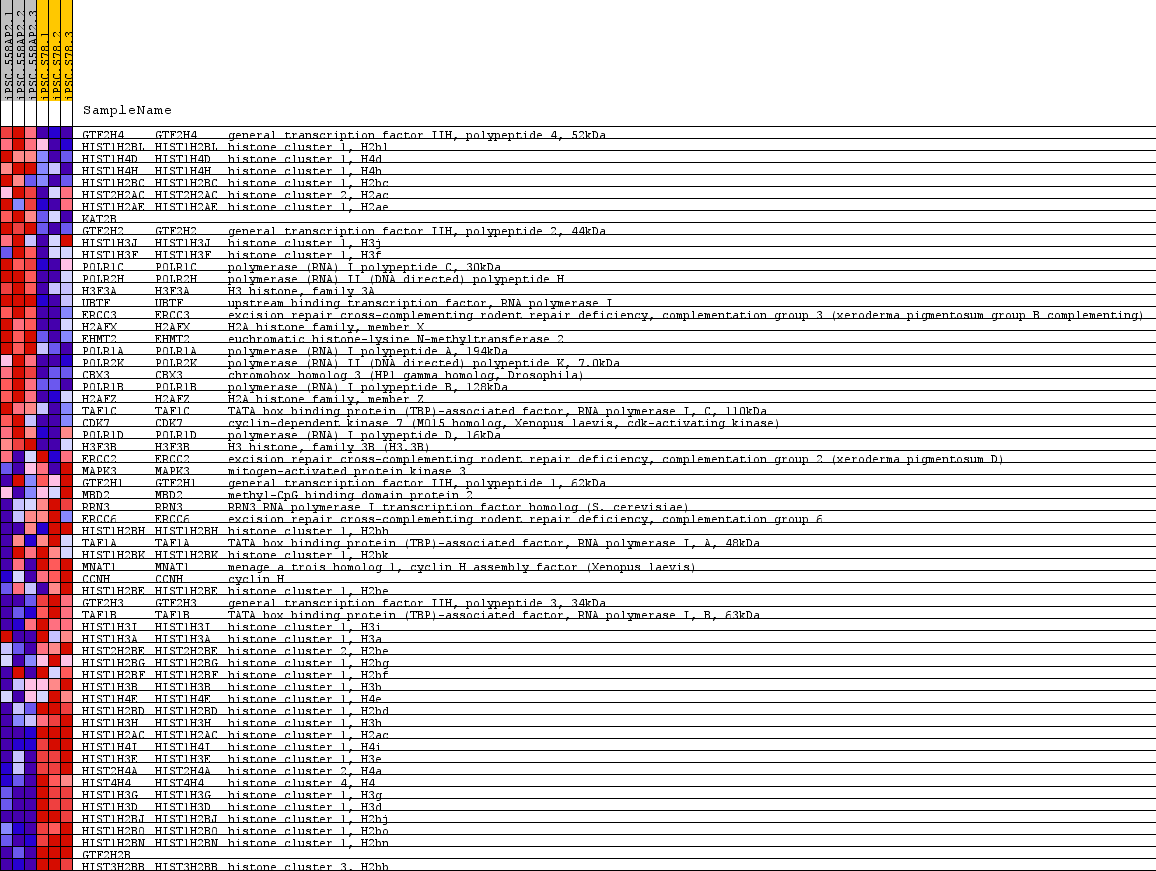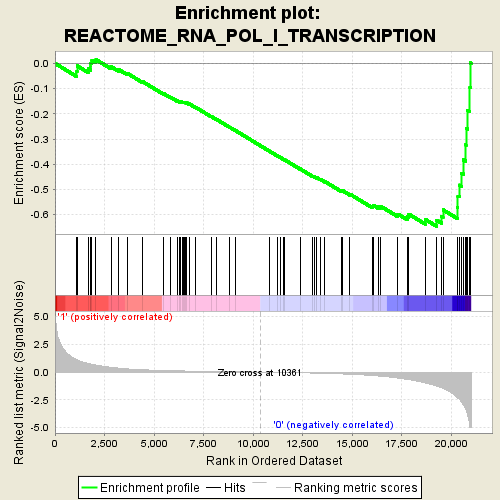
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | NS50832_collapsed_to_symbols.NS50832.cls #558AP2_versus_S78.NS50832.cls #558AP2_versus_S78_repos |
| Phenotype | NS50832.cls#558AP2_versus_S78_repos |
| Upregulated in class | 0 |
| GeneSet | REACTOME_RNA_POL_I_TRANSCRIPTION |
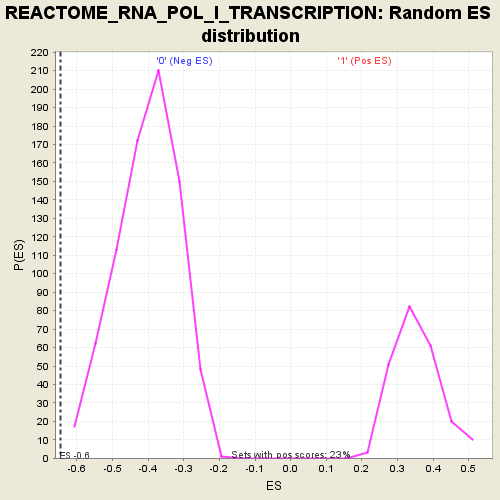
| Enrichment Score (ES) | -0.6457944 |
| Normalized Enrichment Score (NES) | -1.6112123 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.8367749 |
| FWER p-Value | 1.0 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | GTF2H4 | GTF2H4 Entrez, Source | general transcription factor IIH, polypeptide 4, 52kDa | 1097 | 1.119 | -0.0306 | No |
| 2 | HIST1H2BL | HIST1H2BL Entrez, Source | histone cluster 1, H2bl | 1112 | 1.110 | -0.0097 | No |
| 3 | HIST1H4D | HIST1H4D Entrez, Source | histone cluster 1, H4d | 1674 | 0.784 | -0.0212 | No |
| 4 | HIST1H4H | HIST1H4H Entrez, Source | histone cluster 1, H4h | 1800 | 0.733 | -0.0129 | No |
| 5 | HIST1H2BC | HIST1H2BC Entrez, Source | histone cluster 1, H2bc | 1804 | 0.731 | 0.0012 | No |
| 6 | HIST2H2AC | HIST2H2AC Entrez, Source | histone cluster 2, H2ac | 1857 | 0.707 | 0.0125 | No |
| 7 | HIST1H2AE | HIST1H2AE Entrez, Source | histone cluster 1, H2ae | 2062 | 0.630 | 0.0150 | No |
| 8 | KAT2B | 2826 | 0.433 | -0.0130 | No | ||
| 9 | GTF2H2 | GTF2H2 Entrez, Source | general transcription factor IIH, polypeptide 2, 44kDa | 3213 | 0.357 | -0.0245 | No |
| 10 | HIST1H3J | HIST1H3J Entrez, Source | histone cluster 1, H3j | 3655 | 0.289 | -0.0399 | No |
| 11 | HIST1H3F | HIST1H3F Entrez, Source | histone cluster 1, H3f | 4421 | 0.211 | -0.0724 | No |
| 12 | POLR1C | POLR1C Entrez, Source | polymerase (RNA) I polypeptide C, 30kDa | 5463 | 0.147 | -0.1193 | No |
| 13 | POLR2H | POLR2H Entrez, Source | polymerase (RNA) II (DNA directed) polypeptide H | 5820 | 0.130 | -0.1337 | No |
| 14 | H3F3A | H3F3A Entrez, Source | H3 histone, family 3A | 6169 | 0.115 | -0.1481 | No |
| 15 | UBTF | UBTF Entrez, Source | upstream binding transcription factor, RNA polymerase I | 6288 | 0.111 | -0.1516 | No |
| 16 | ERCC3 | ERCC3 Entrez, Source | excision repair cross-complementing rodent repair deficiency, complementation group 3 (xeroderma pigmentosum group B complementing) | 6310 | 0.109 | -0.1505 | No |
| 17 | H2AFX | H2AFX Entrez, Source | H2A histone family, member X | 6444 | 0.104 | -0.1548 | No |
| 18 | EHMT2 | EHMT2 Entrez, Source | euchromatic histone-lysine N-methyltransferase 2 | 6478 | 0.103 | -0.1544 | No |
| 19 | POLR1A | POLR1A Entrez, Source | polymerase (RNA) I polypeptide A, 194kDa | 6516 | 0.102 | -0.1542 | No |
| 20 | POLR2K | POLR2K Entrez, Source | polymerase (RNA) II (DNA directed) polypeptide K, 7.0kDa | 6579 | 0.100 | -0.1552 | No |
| 21 | CBX3 | CBX3 Entrez, Source | chromobox homolog 3 (HP1 gamma homolog, Drosophila) | 6625 | 0.099 | -0.1554 | No |
| 22 | POLR1B | POLR1B Entrez, Source | polymerase (RNA) I polypeptide B, 128kDa | 6770 | 0.094 | -0.1605 | No |
| 23 | H2AFZ | H2AFZ Entrez, Source | H2A histone family, member Z | 7072 | 0.084 | -0.1732 | No |
| 24 | TAF1C | TAF1C Entrez, Source | TATA box binding protein (TBP)-associated factor, RNA polymerase I, C, 110kDa | 7868 | 0.062 | -0.2100 | No |
| 25 | CDK7 | CDK7 Entrez, Source | cyclin-dependent kinase 7 (MO15 homolog, Xenopus laevis, cdk-activating kinase) | 8163 | 0.054 | -0.2230 | No |
| 26 | POLR1D | POLR1D Entrez, Source | polymerase (RNA) I polypeptide D, 16kDa | 8811 | 0.038 | -0.2531 | No |
| 27 | H3F3B | H3F3B Entrez, Source | H3 histone, family 3B (H3.3B) | 9123 | 0.030 | -0.2674 | No |
| 28 | ERCC2 | ERCC2 Entrez, Source | excision repair cross-complementing rodent repair deficiency, complementation group 2 (xeroderma pigmentosum D) | 10820 | -0.012 | -0.3482 | No |
| 29 | MAPK3 | MAPK3 Entrez, Source | mitogen-activated protein kinase 3 | 11216 | -0.023 | -0.3666 | No |
| 30 | GTF2H1 | GTF2H1 Entrez, Source | general transcription factor IIH, polypeptide 1, 62kDa | 11355 | -0.026 | -0.3727 | No |
| 31 | MBD2 | MBD2 Entrez, Source | methyl-CpG binding domain protein 2 | 11517 | -0.031 | -0.3798 | No |
| 32 | RRN3 | RRN3 Entrez, Source | RRN3 RNA polymerase I transcription factor homolog (S. cerevisiae) | 11593 | -0.034 | -0.3827 | No |
| 33 | ERCC6 | ERCC6 Entrez, Source | excision repair cross-complementing rodent repair deficiency, complementation group 6 | 12359 | -0.059 | -0.4181 | No |
| 34 | HIST1H2BH | HIST1H2BH Entrez, Source | histone cluster 1, H2bh | 13000 | -0.085 | -0.4470 | No |
| 35 | TAF1A | TAF1A Entrez, Source | TATA box binding protein (TBP)-associated factor, RNA polymerase I, A, 48kDa | 13086 | -0.089 | -0.4494 | No |
| 36 | HIST1H2BK | HIST1H2BK Entrez, Source | histone cluster 1, H2bk | 13205 | -0.093 | -0.4532 | No |
| 37 | MNAT1 | MNAT1 Entrez, Source | menage a trois homolog 1, cyclin H assembly factor (Xenopus laevis) | 13376 | -0.101 | -0.4593 | No |
| 38 | CCNH | CCNH Entrez, Source | cyclin H | 13609 | -0.111 | -0.4683 | No |
| 39 | HIST1H2BE | HIST1H2BE Entrez, Source | histone cluster 1, H2be | 14463 | -0.164 | -0.5058 | No |
| 40 | GTF2H3 | GTF2H3 Entrez, Source | general transcription factor IIH, polypeptide 3, 34kDa | 14510 | -0.167 | -0.5048 | No |
| 41 | TAF1B | TAF1B Entrez, Source | TATA box binding protein (TBP)-associated factor, RNA polymerase I, B, 63kDa | 14877 | -0.195 | -0.5185 | No |
| 42 | HIST1H3I | HIST1H3I Entrez, Source | histone cluster 1, H3i | 16000 | -0.305 | -0.5661 | No |
| 43 | HIST1H3A | HIST1H3A Entrez, Source | histone cluster 1, H3a | 16043 | -0.311 | -0.5621 | No |
| 44 | HIST2H2BE | HIST2H2BE Entrez, Source | histone cluster 2, H2be | 16298 | -0.347 | -0.5674 | No |
| 45 | HIST1H2BG | HIST1H2BG Entrez, Source | histone cluster 1, H2bg | 16429 | -0.367 | -0.5665 | No |
| 46 | HIST1H2BF | HIST1H2BF Entrez, Source | histone cluster 1, H2bf | 17290 | -0.531 | -0.5973 | No |
| 47 | HIST1H3B | HIST1H3B Entrez, Source | histone cluster 1, H3b | 17776 | -0.654 | -0.6077 | No |
| 48 | HIST1H4E | HIST1H4E Entrez, Source | histone cluster 1, H4e | 17843 | -0.676 | -0.5977 | No |
| 49 | HIST1H2BD | HIST1H2BD Entrez, Source | histone cluster 1, H2bd | 18676 | -0.971 | -0.6185 | No |
| 50 | HIST1H3H | HIST1H3H Entrez, Source | histone cluster 1, H3h | 19248 | -1.251 | -0.6214 | Yes |
| 51 | HIST1H2AC | HIST1H2AC Entrez, Source | histone cluster 1, H2ac | 19503 | -1.417 | -0.6060 | Yes |
| 52 | HIST1H4I | HIST1H4I Entrez, Source | histone cluster 1, H4i | 19573 | -1.473 | -0.5806 | Yes |
| 53 | HIST1H3E | HIST1H3E Entrez, Source | histone cluster 1, H3e | 20289 | -2.284 | -0.5702 | Yes |
| 54 | HIST2H4A | HIST2H4A Entrez, Source | histone cluster 2, H4a | 20322 | -2.327 | -0.5264 | Yes |
| 55 | HIST4H4 | HIST4H4 Entrez, Source | histone cluster 4, H4 | 20390 | -2.440 | -0.4821 | Yes |
| 56 | HIST1H3G | HIST1H3G Entrez, Source | histone cluster 1, H3g | 20514 | -2.709 | -0.4352 | Yes |
| 57 | HIST1H3D | HIST1H3D Entrez, Source | histone cluster 1, H3d | 20594 | -2.926 | -0.3820 | Yes |
| 58 | HIST1H2BJ | HIST1H2BJ Entrez, Source | histone cluster 1, H2bj | 20719 | -3.326 | -0.3231 | Yes |
| 59 | HIST1H2BO | HIST1H2BO Entrez, Source | histone cluster 1, H2bo | 20735 | -3.400 | -0.2576 | Yes |
| 60 | HIST1H2BN | HIST1H2BN Entrez, Source | histone cluster 1, H2bn | 20809 | -3.847 | -0.1862 | Yes |
| 61 | GTF2H2B | 20927 | -5.000 | -0.0944 | Yes | ||
| 62 | HIST3H2BB | HIST3H2BB Entrez, Source | histone cluster 3, H2bb | 20947 | -5.000 | 0.0021 | Yes |