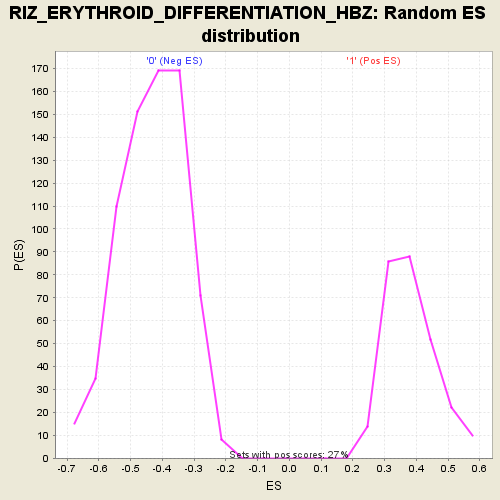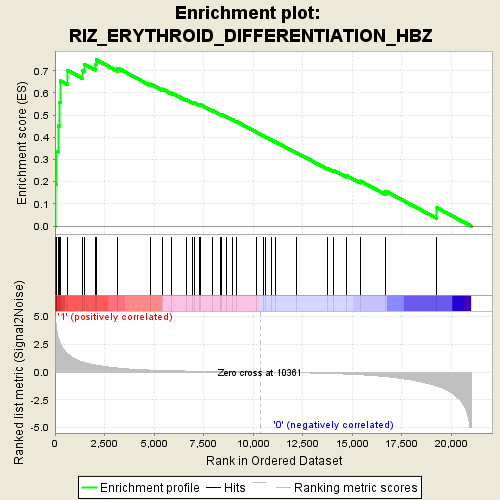
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | NS50832_collapsed_to_symbols.NS50832.cls #558AP2_versus_S78.NS50832.cls #558AP2_versus_S78_repos |
| Phenotype | NS50832.cls#558AP2_versus_S78_repos |
| Upregulated in class | 1 |
| GeneSet | RIZ_ERYTHROID_DIFFERENTIATION_HBZ |
| Enrichment Score (ES) | 0.7501938 |
| Normalized Enrichment Score (NES) | 1.9708331 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.014434453 |
| FWER p-Value | 0.045 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | NR4A2 | NR4A2 Entrez, Source | nuclear receptor subfamily 4, group A, member 2 | 18 | 5.000 | 0.1905 | Yes |
| 2 | POU3F2 | POU3F2 Entrez, Source | POU domain, class 3, transcription factor 2 | 74 | 3.921 | 0.3380 | Yes |
| 3 | SIX2 | SIX2 Entrez, Source | sine oculis homeobox homolog 2 (Drosophila) | 161 | 3.101 | 0.4526 | Yes |
| 4 | PITX2 | PITX2 Entrez, Source | paired-like homeodomain transcription factor 2 | 223 | 2.814 | 0.5574 | Yes |
| 5 | SOX1 | SOX1 Entrez, Source | SRY (sex determining region Y)-box 1 | 268 | 2.586 | 0.6542 | Yes |
| 6 | POU2F3 | POU2F3 Entrez, Source | POU domain, class 2, transcription factor 3 | 613 | 1.689 | 0.7025 | Yes |
| 7 | NKX2-5 | NKX2-5 Entrez, Source | NK2 transcription factor related, locus 5 (Drosophila) | 1370 | 0.926 | 0.7018 | Yes |
| 8 | POU3F3 | POU3F3 Entrez, Source | POU domain, class 3, transcription factor 3 | 1496 | 0.863 | 0.7289 | Yes |
| 9 | MPL | MPL Entrez, Source | myeloproliferative leukemia virus oncogene | 2033 | 0.640 | 0.7278 | Yes |
| 10 | GATA5 | GATA5 Entrez, Source | GATA binding protein 5 | 2069 | 0.628 | 0.7502 | Yes |
| 11 | FOSB | FOSB Entrez, Source | FBJ murine osteosarcoma viral oncogene homolog B | 3157 | 0.367 | 0.7124 | No |
| 12 | HMGB1 | HMGB1 Entrez, Source | high-mobility group box 1 | 4814 | 0.184 | 0.6404 | No |
| 13 | TFAP2C | TFAP2C Entrez, Source | transcription factor AP-2 gamma (activating enhancer binding protein 2 gamma) | 5407 | 0.150 | 0.6179 | No |
| 14 | ZNF436 | ZNF436 Entrez, Source | zinc finger protein 436 | 5887 | 0.127 | 0.5999 | No |
| 15 | CBX3 | CBX3 Entrez, Source | chromobox homolog 3 (HP1 gamma homolog, Drosophila) | 6625 | 0.099 | 0.5685 | No |
| 16 | SOX4 | SOX4 Entrez, Source | SRY (sex determining region Y)-box 4 | 6940 | 0.088 | 0.5568 | No |
| 17 | PROX1 | PROX1 Entrez, Source | prospero-related homeobox 1 | 7029 | 0.085 | 0.5559 | No |
| 18 | MCM4 | MCM4 Entrez, Source | MCM4 minichromosome maintenance deficient 4 (S. cerevisiae) | 7304 | 0.078 | 0.5458 | No |
| 19 | KDM5A | 7360 | 0.076 | 0.5461 | No | ||
| 20 | MCM7 | MCM7 Entrez, Source | MCM7 minichromosome maintenance deficient 7 (S. cerevisiae) | 7964 | 0.059 | 0.5196 | No |
| 21 | CUX1 | 8365 | 0.049 | 0.5024 | No | ||
| 22 | AFF1 | AFF1 Entrez, Source | AF4/FMR2 family, member 1 | 8417 | 0.048 | 0.5018 | No |
| 23 | E2F8 | E2F8 Entrez, Source | E2F transcription factor 8 | 8661 | 0.042 | 0.4918 | No |
| 24 | BASP1 | BASP1 Entrez, Source | brain abundant, membrane attached signal protein 1 | 8942 | 0.035 | 0.4798 | No |
| 25 | PAPOLA | PAPOLA Entrez, Source | poly(A) polymerase alpha | 9173 | 0.029 | 0.4699 | No |
| 26 | TMPO | TMPO Entrez, Source | thymopoietin | 10168 | 0.005 | 0.4227 | No |
| 27 | SMARCD2 | SMARCD2 Entrez, Source | SWI/SNF related, matrix associated, actin dependent regulator of chromatin, subfamily d, member 2 | 10501 | -0.004 | 0.4070 | No |
| 28 | RB1 | RB1 Entrez, Source | retinoblastoma 1 (including osteosarcoma) | 10594 | -0.007 | 0.4029 | No |
| 29 | MCM5 | MCM5 Entrez, Source | MCM5 minichromosome maintenance deficient 5, cell division cycle 46 (S. cerevisiae) | 10895 | -0.014 | 0.3891 | No |
| 30 | CBX7 | CBX7 Entrez, Source | chromobox homolog 7 | 11124 | -0.020 | 0.3790 | No |
| 31 | RLIM | 12176 | -0.052 | 0.3309 | No | ||
| 32 | BARD1 | BARD1 Entrez, Source | BRCA1 associated RING domain 1 | 13766 | -0.120 | 0.2596 | No |
| 33 | SOX18 | SOX18 Entrez, Source | SRY (sex determining region Y)-box 18 | 14059 | -0.136 | 0.2509 | No |
| 34 | GCM2 | GCM2 Entrez, Source | glial cells missing homolog 2 (Drosophila) | 14683 | -0.181 | 0.2281 | No |
| 35 | NR5A1 | NR5A1 Entrez, Source | nuclear receptor subfamily 5, group A, member 1 | 15395 | -0.239 | 0.2033 | No |
| 36 | EYA2 | EYA2 Entrez, Source | eyes absent homolog 2 (Drosophila) | 16678 | -0.405 | 0.1576 | No |
| 37 | TSC22D3 | TSC22D3 Entrez, Source | TSC22 domain family, member 3 | 19263 | -1.259 | 0.0825 | No |