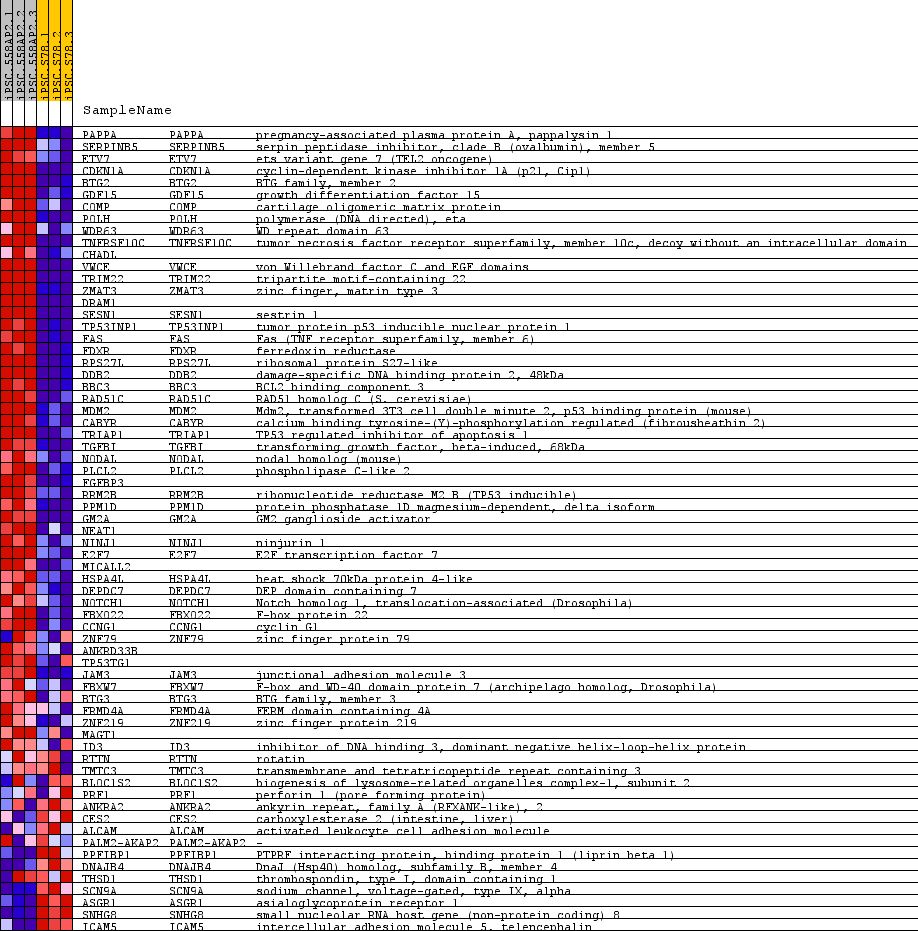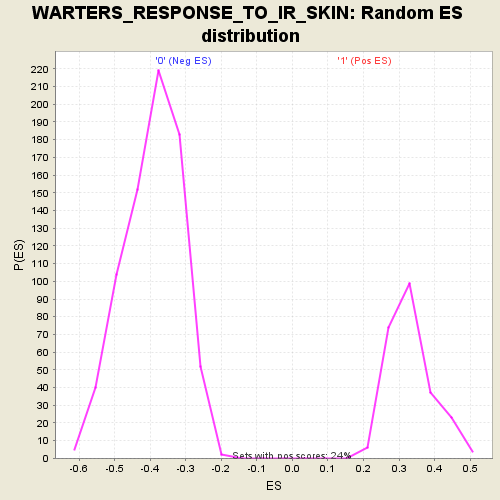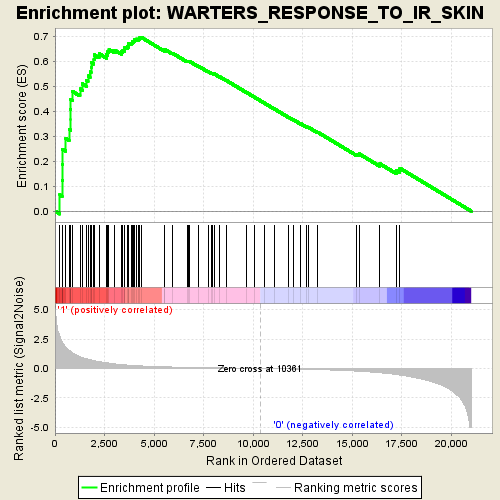
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | NS50832_collapsed_to_symbols.NS50832.cls #558AP2_versus_S78.NS50832.cls #558AP2_versus_S78_repos |
| Phenotype | NS50832.cls#558AP2_versus_S78_repos |
| Upregulated in class | 1 |
| GeneSet | WARTERS_RESPONSE_TO_IR_SKIN |
| Enrichment Score (ES) | 0.69644755 |
| Normalized Enrichment Score (NES) | 2.097812 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.0019980841 |
| FWER p-Value | 0.001 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | PAPPA | PAPPA Entrez, Source | pregnancy-associated plasma protein A, pappalysin 1 | 242 | 2.715 | 0.0651 | Yes |
| 2 | SERPINB5 | SERPINB5 Entrez, Source | serpin peptidase inhibitor, clade B (ovalbumin), member 5 | 354 | 2.302 | 0.1248 | Yes |
| 3 | ETV7 | ETV7 Entrez, Source | ets variant gene 7 (TEL2 oncogene) | 374 | 2.223 | 0.1866 | Yes |
| 4 | CDKN1A | CDKN1A Entrez, Source | cyclin-dependent kinase inhibitor 1A (p21, Cip1) | 383 | 2.199 | 0.2483 | Yes |
| 5 | BTG2 | BTG2 Entrez, Source | BTG family, member 2 | 543 | 1.815 | 0.2919 | Yes |
| 6 | GDF15 | GDF15 Entrez, Source | growth differentiation factor 15 | 707 | 1.545 | 0.3277 | Yes |
| 7 | COMP | COMP Entrez, Source | cartilage oligomeric matrix protein | 773 | 1.463 | 0.3659 | Yes |
| 8 | POLH | POLH Entrez, Source | polymerase (DNA directed), eta | 775 | 1.461 | 0.4071 | Yes |
| 9 | WDR63 | WDR63 Entrez, Source | WD repeat domain 63 | 792 | 1.441 | 0.4471 | Yes |
| 10 | TNFRSF10C | TNFRSF10C Entrez, Source | tumor necrosis factor receptor superfamily, member 10c, decoy without an intracellular domain | 889 | 1.311 | 0.4795 | Yes |
| 11 | CHADL | 1258 | 1.000 | 0.4901 | Yes | ||
| 12 | VWCE | VWCE Entrez, Source | von Willebrand factor C and EGF domains | 1377 | 0.921 | 0.5105 | Yes |
| 13 | TRIM22 | TRIM22 Entrez, Source | tripartite motif-containing 22 | 1594 | 0.815 | 0.5232 | Yes |
| 14 | ZMAT3 | ZMAT3 Entrez, Source | zinc finger, matrin type 3 | 1662 | 0.789 | 0.5422 | Yes |
| 15 | DRAM1 | 1765 | 0.743 | 0.5583 | Yes | ||
| 16 | SESN1 | SESN1 Entrez, Source | sestrin 1 | 1816 | 0.723 | 0.5764 | Yes |
| 17 | TP53INP1 | TP53INP1 Entrez, Source | tumor protein p53 inducible nuclear protein 1 | 1829 | 0.717 | 0.5960 | Yes |
| 18 | FAS | FAS Entrez, Source | Fas (TNF receptor superfamily, member 6) | 1961 | 0.665 | 0.6086 | Yes |
| 19 | FDXR | FDXR Entrez, Source | ferredoxin reductase | 1974 | 0.659 | 0.6266 | Yes |
| 20 | RPS27L | RPS27L Entrez, Source | ribosomal protein S27-like | 2219 | 0.587 | 0.6315 | Yes |
| 21 | DDB2 | DDB2 Entrez, Source | damage-specific DNA binding protein 2, 48kDa | 2601 | 0.487 | 0.6271 | Yes |
| 22 | BBC3 | BBC3 Entrez, Source | BCL2 binding component 3 | 2645 | 0.478 | 0.6385 | Yes |
| 23 | RAD51C | RAD51C Entrez, Source | RAD51 homolog C (S. cerevisiae) | 2719 | 0.460 | 0.6480 | Yes |
| 24 | MDM2 | MDM2 Entrez, Source | Mdm2, transformed 3T3 cell double minute 2, p53 binding protein (mouse) | 3005 | 0.396 | 0.6456 | Yes |
| 25 | CABYR | CABYR Entrez, Source | calcium binding tyrosine-(Y)-phosphorylation regulated (fibrousheathin 2) | 3344 | 0.336 | 0.6389 | Yes |
| 26 | TRIAP1 | TRIAP1 Entrez, Source | TP53 regulated inhibitor of apoptosis 1 | 3416 | 0.325 | 0.6447 | Yes |
| 27 | TGFBI | TGFBI Entrez, Source | transforming growth factor, beta-induced, 68kDa | 3501 | 0.310 | 0.6494 | Yes |
| 28 | NODAL | NODAL Entrez, Source | nodal homolog (mouse) | 3518 | 0.308 | 0.6573 | Yes |
| 29 | PLCL2 | PLCL2 Entrez, Source | phospholipase C-like 2 | 3642 | 0.291 | 0.6597 | Yes |
| 30 | FGFBP3 | 3690 | 0.286 | 0.6655 | Yes | ||
| 31 | RRM2B | RRM2B Entrez, Source | ribonucleotide reductase M2 B (TP53 inducible) | 3693 | 0.285 | 0.6734 | Yes |
| 32 | PPM1D | PPM1D Entrez, Source | protein phosphatase 1D magnesium-dependent, delta isoform | 3838 | 0.266 | 0.6740 | Yes |
| 33 | GM2A | GM2A Entrez, Source | GM2 ganglioside activator | 3889 | 0.259 | 0.6790 | Yes |
| 34 | NEAT1 | 3974 | 0.250 | 0.6820 | Yes | ||
| 35 | NINJ1 | NINJ1 Entrez, Source | ninjurin 1 | 4000 | 0.247 | 0.6878 | Yes |
| 36 | E2F7 | E2F7 Entrez, Source | E2F transcription factor 7 | 4089 | 0.239 | 0.6903 | Yes |
| 37 | MICALL2 | 4224 | 0.227 | 0.6903 | Yes | ||
| 38 | HSPA4L | HSPA4L Entrez, Source | heat shock 70kDa protein 4-like | 4267 | 0.224 | 0.6946 | Yes |
| 39 | DEPDC7 | DEPDC7 Entrez, Source | DEP domain containing 7 | 4358 | 0.216 | 0.6964 | Yes |
| 40 | NOTCH1 | NOTCH1 Entrez, Source | Notch homolog 1, translocation-associated (Drosophila) | 5526 | 0.144 | 0.6447 | No |
| 41 | FBXO22 | FBXO22 Entrez, Source | F-box protein 22 | 5536 | 0.143 | 0.6483 | No |
| 42 | CCNG1 | CCNG1 Entrez, Source | cyclin G1 | 5914 | 0.125 | 0.6338 | No |
| 43 | ZNF79 | ZNF79 Entrez, Source | zinc finger protein 79 | 6655 | 0.097 | 0.6012 | No |
| 44 | ANKRD33B | 6715 | 0.095 | 0.6011 | No | ||
| 45 | TP53TG1 | 6757 | 0.094 | 0.6018 | No | ||
| 46 | JAM3 | JAM3 Entrez, Source | junctional adhesion molecule 3 | 7235 | 0.080 | 0.5813 | No |
| 47 | FBXW7 | FBXW7 Entrez, Source | F-box and WD-40 domain protein 7 (archipelago homolog, Drosophila) | 7720 | 0.066 | 0.5600 | No |
| 48 | BTG3 | BTG3 Entrez, Source | BTG family, member 3 | 7888 | 0.062 | 0.5538 | No |
| 49 | FRMD4A | FRMD4A Entrez, Source | FERM domain containing 4A | 7929 | 0.061 | 0.5536 | No |
| 50 | ZNF219 | ZNF219 Entrez, Source | zinc finger protein 219 | 8024 | 0.058 | 0.5507 | No |
| 51 | MAGT1 | 8318 | 0.050 | 0.5381 | No | ||
| 52 | ID3 | ID3 Entrez, Source | inhibitor of DNA binding 3, dominant negative helix-loop-helix protein | 8646 | 0.043 | 0.5237 | No |
| 53 | RTTN | RTTN Entrez, Source | rotatin | 9663 | 0.017 | 0.4756 | No |
| 54 | TMTC3 | TMTC3 Entrez, Source | transmembrane and tetratricopeptide repeat containing 3 | 10058 | 0.007 | 0.4570 | No |
| 55 | BLOC1S2 | BLOC1S2 Entrez, Source | biogenesis of lysosome-related organelles complex-1, subunit 2 | 10555 | -0.006 | 0.4335 | No |
| 56 | PRF1 | PRF1 Entrez, Source | perforin 1 (pore forming protein) | 11091 | -0.020 | 0.4084 | No |
| 57 | ANKRA2 | ANKRA2 Entrez, Source | ankyrin repeat, family A (RFXANK-like), 2 | 11764 | -0.040 | 0.3774 | No |
| 58 | CES2 | CES2 Entrez, Source | carboxylesterase 2 (intestine, liver) | 12008 | -0.047 | 0.3672 | No |
| 59 | ALCAM | ALCAM Entrez, Source | activated leukocyte cell adhesion molecule | 12403 | -0.060 | 0.3501 | No |
| 60 | PALM2-AKAP2 | PALM2-AKAP2 Entrez, Source | - | 12683 | -0.071 | 0.3387 | No |
| 61 | PPFIBP1 | PPFIBP1 Entrez, Source | PTPRF interacting protein, binding protein 1 (liprin beta 1) | 12781 | -0.075 | 0.3362 | No |
| 62 | DNAJB4 | DNAJB4 Entrez, Source | DnaJ (Hsp40) homolog, subfamily B, member 4 | 13234 | -0.094 | 0.3173 | No |
| 63 | THSD1 | THSD1 Entrez, Source | thrombospondin, type I, domain containing 1 | 15218 | -0.223 | 0.2288 | No |
| 64 | SCN9A | SCN9A Entrez, Source | sodium channel, voltage-gated, type IX, alpha | 15333 | -0.234 | 0.2299 | No |
| 65 | ASGR1 | ASGR1 Entrez, Source | asialoglycoprotein receptor 1 | 16387 | -0.360 | 0.1898 | No |
| 66 | SNHG8 | SNHG8 Entrez, Source | small nucleolar RNA host gene (non-protein coding) 8 | 17221 | -0.515 | 0.1645 | No |
| 67 | ICAM5 | ICAM5 Entrez, Source | intercellular adhesion molecule 5, telencephalin | 17395 | -0.553 | 0.1719 | No |