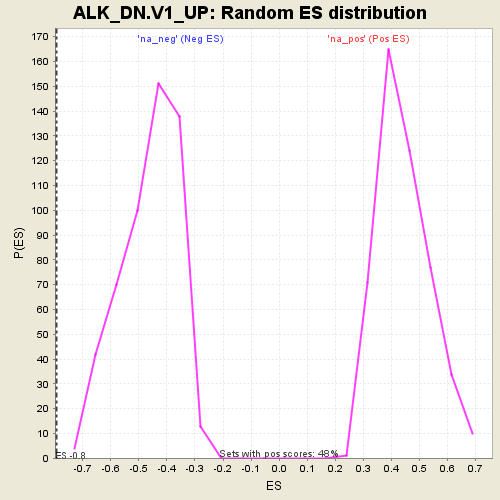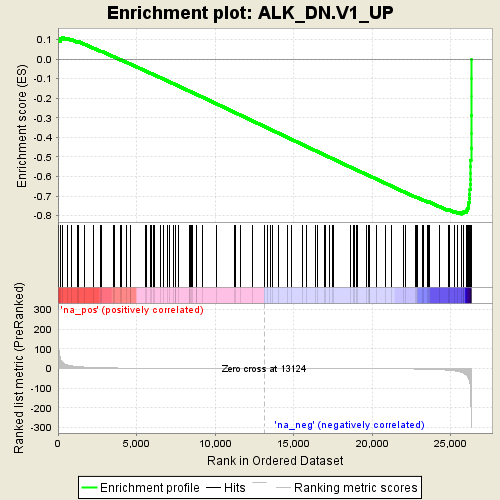
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | NS50832_padj |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_neg |
| GeneSet | ALK_DN.V1_UP |
| Enrichment Score (ES) | -0.79185736 |
| Normalized Enrichment Score (NES) | -1.7193749 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.110682644 |
| FWER p-Value | 0.817 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | SPP1 | 0 | 299.207 | 0.0968 | No | ||
| 2 | NNAT | 167 | 47.370 | 0.1058 | No | ||
| 3 | GDF15 | 292 | 30.340 | 0.1109 | No | ||
| 4 | OLFML3 | 590 | 18.443 | 0.1055 | No | ||
| 5 | CLSPN | 837 | 14.160 | 0.1007 | No | ||
| 6 | LIN28A | 1207 | 10.373 | 0.0900 | No | ||
| 7 | CA2 | 1305 | 9.624 | 0.0894 | No | ||
| 8 | PYGB | 1708 | 7.220 | 0.0764 | No | ||
| 9 | ENPP2 | 2284 | 5.147 | 0.0561 | No | ||
| 10 | GRPR | 2716 | 4.151 | 0.0410 | No | ||
| 11 | FJX1 | 2785 | 4.014 | 0.0397 | No | ||
| 12 | SCN1A | 3501 | 2.869 | 0.0134 | No | ||
| 13 | PRDM1 | 3563 | 2.787 | 0.0120 | No | ||
| 14 | UNC13A | 4004 | 2.315 | -0.0041 | No | ||
| 15 | ADAM12 | 4049 | 2.274 | -0.0050 | No | ||
| 16 | SLC12A8 | 4334 | 1.985 | -0.0152 | No | ||
| 17 | DECR2 | 4616 | 1.765 | -0.0254 | No | ||
| 18 | CFTR | 4617 | 1.764 | -0.0248 | No | ||
| 19 | AIF1 | 5597 | 1.121 | -0.0618 | No | ||
| 20 | IL6 | 5641 | 1.099 | -0.0631 | No | ||
| 21 | CBS | 5888 | 0.974 | -0.0721 | No | ||
| 22 | FBXW4P1 | 5951 | 0.948 | -0.0742 | No | ||
| 23 | ACTL6B | 6062 | 0.909 | -0.0781 | No | ||
| 24 | ENTPD3 | 6150 | 0.877 | -0.0811 | No | ||
| 25 | CCNK | 6524 | 0.754 | -0.0951 | No | ||
| 26 | CCR7 | 6684 | 0.692 | -0.1010 | No | ||
| 27 | CES1P1 | 6972 | 0.612 | -0.1117 | No | ||
| 28 | EPAS1 | 7104 | 0.575 | -0.1165 | No | ||
| 29 | AQP7 | 7323 | 0.523 | -0.1247 | No | ||
| 30 | GALK1 | 7464 | 0.494 | -0.1299 | No | ||
| 31 | NTRK1 | 7506 | 0.485 | -0.1313 | No | ||
| 32 | ANGPTL4 | 7672 | 0.452 | -0.1374 | No | ||
| 33 | PDZD7 | 8368 | 0.343 | -0.1638 | No | ||
| 34 | FMO1 | 8393 | 0.339 | -0.1646 | No | ||
| 35 | SAA1 | 8415 | 0.335 | -0.1653 | No | ||
| 36 | MRAS | 8468 | 0.326 | -0.1672 | No | ||
| 37 | ANGPT2 | 8531 | 0.318 | -0.1695 | No | ||
| 38 | IL1R2 | 8793 | 0.286 | -0.1793 | No | ||
| 39 | CATSPERG | 8808 | 0.285 | -0.1798 | No | ||
| 40 | MLC1 | 9213 | 0.237 | -0.1951 | No | ||
| 41 | RIBC2 | 10098 | 0.154 | -0.2288 | No | ||
| 42 | SMPX | 11228 | 0.081 | -0.2718 | No | ||
| 43 | BEGAIN | 11288 | 0.077 | -0.2741 | No | ||
| 44 | FBXL8 | 11634 | 0.058 | -0.2872 | No | ||
| 45 | B3GALT4 | 12393 | 0.027 | -0.3161 | No | ||
| 46 | MPO | 13130 | -0.000 | -0.3442 | No | ||
| 47 | DLX6 | 13359 | -0.007 | -0.3529 | No | ||
| 48 | CSTA | 13512 | -0.013 | -0.3587 | No | ||
| 49 | GDF5 | 13637 | -0.017 | -0.3634 | No | ||
| 50 | P2RX3 | 14035 | -0.033 | -0.3786 | No | ||
| 51 | EREG | 14604 | -0.054 | -0.4002 | No | ||
| 52 | RBM17 | 14845 | -0.063 | -0.4094 | No | ||
| 53 | ADAM28 | 15574 | -0.107 | -0.4371 | No | ||
| 54 | LTC4S | 15850 | -0.123 | -0.4475 | No | ||
| 55 | PYY2 | 16410 | -0.162 | -0.4688 | No | ||
| 56 | CHI3L1 | 16514 | -0.172 | -0.4727 | No | ||
| 57 | AMELY | 16936 | -0.211 | -0.4887 | No | ||
| 58 | C8orf4 | 17032 | -0.220 | -0.4922 | No | ||
| 59 | HLA-DMB | 17306 | -0.250 | -0.5026 | No | ||
| 60 | KYNU | 17504 | -0.275 | -0.5100 | No | ||
| 61 | LYL1 | 17518 | -0.277 | -0.5104 | No | ||
| 62 | BST1 | 18643 | -0.442 | -0.5532 | No | ||
| 63 | CA3 | 18822 | -0.476 | -0.5598 | No | ||
| 64 | BMPR1B | 18895 | -0.493 | -0.5624 | No | ||
| 65 | NOD2 | 18995 | -0.512 | -0.5660 | No | ||
| 66 | TBL1Y | 19096 | -0.533 | -0.5696 | No | ||
| 67 | LMX1B | 19662 | -0.678 | -0.5910 | No | ||
| 68 | DMP1 | 19790 | -0.711 | -0.5956 | No | ||
| 69 | THAP3 | 19805 | -0.717 | -0.5959 | No | ||
| 70 | STEAP4 | 19863 | -0.738 | -0.5978 | No | ||
| 71 | XBP1 | 20283 | -0.879 | -0.6135 | No | ||
| 72 | CHST8 | 20844 | -1.116 | -0.6345 | No | ||
| 73 | TTLL2 | 21261 | -1.342 | -0.6500 | No | ||
| 74 | CARD9 | 22020 | -1.883 | -0.6783 | No | ||
| 75 | SCN9A | 22097 | -1.941 | -0.6806 | No | ||
| 76 | VEGFC | 22780 | -2.666 | -0.7057 | No | ||
| 77 | RRAS | 22841 | -2.760 | -0.7071 | No | ||
| 78 | GNA15 | 22901 | -2.824 | -0.7085 | No | ||
| 79 | BDKRB2 | 23220 | -3.273 | -0.7195 | No | ||
| 80 | MT1H | 23299 | -3.419 | -0.7214 | No | ||
| 81 | FOXP3 | 23543 | -3.872 | -0.7294 | No | ||
| 82 | ZNF639 | 23610 | -4.022 | -0.7306 | No | ||
| 83 | SLC7A5 | 23621 | -4.053 | -0.7297 | No | ||
| 84 | VSTM4 | 24290 | -5.955 | -0.7533 | No | ||
| 85 | ETV1 | 24832 | -8.406 | -0.7712 | No | ||
| 86 | NFKBIL1 | 24900 | -8.881 | -0.7709 | No | ||
| 87 | ARHGAP4 | 25250 | -12.026 | -0.7803 | No | ||
| 88 | CH25H | 25431 | -14.376 | -0.7825 | No | ||
| 89 | STMN2 | 25677 | -19.359 | -0.7856 | Yes | ||
| 90 | GRM1 | 25789 | -22.515 | -0.7825 | Yes | ||
| 91 | SLC1A3 | 25845 | -24.869 | -0.7766 | Yes | ||
| 92 | SLC6A9 | 25989 | -33.245 | -0.7713 | Yes | ||
| 93 | SLC1A5 | 26086 | -42.020 | -0.7614 | Yes | ||
| 94 | SLC7A11 | 26126 | -47.089 | -0.7476 | Yes | ||
| 95 | FLVCR2 | 26164 | -53.891 | -0.7316 | Yes | ||
| 96 | FGF21 | 26188 | -59.810 | -0.7131 | Yes | ||
| 97 | CEBPG | 26210 | -66.827 | -0.6923 | Yes | ||
| 98 | PSAT1 | 26230 | -76.008 | -0.6684 | Yes | ||
| 99 | ASNS | 26241 | -83.978 | -0.6416 | Yes | ||
| 100 | VEGFA | 26251 | -89.987 | -0.6129 | Yes | ||
| 101 | MTHFD2 | 26257 | -95.222 | -0.5822 | Yes | ||
| 102 | STC2 | 26260 | -97.527 | -0.5508 | Yes | ||
| 103 | CTH | 26265 | -99.001 | -0.5189 | Yes | ||
| 104 | PCK2 | 26298 | -198.705 | -0.4558 | Yes | ||
| 105 | TRIB3 | 26304 | -235.323 | -0.3799 | Yes | ||
| 106 | CXCL5 | 26306 | -277.394 | -0.2901 | Yes | ||
| 107 | DDIT4 | 26311 | -299.207 | -0.1935 | Yes | ||
| 108 | INHBE | 26312 | -299.207 | -0.0967 | Yes | ||
| 109 | CHAC1 | 26314 | -299.207 | 0.0001 | Yes |