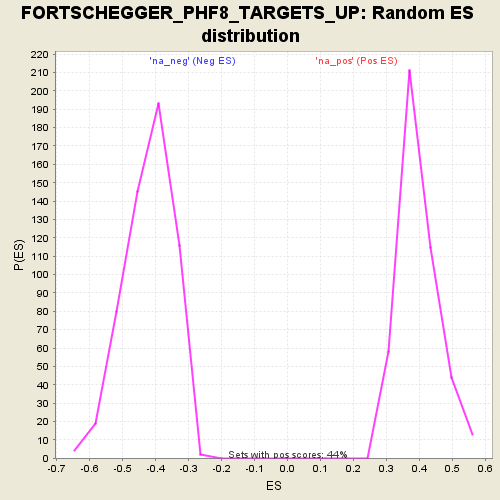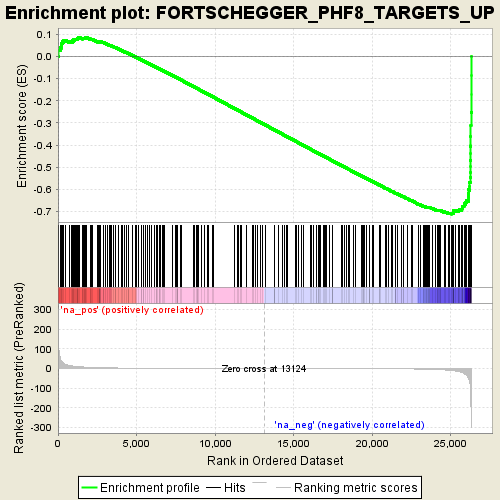
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | NS50832_padj |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_neg |
| GeneSet | FORTSCHEGGER_PHF8_TARGETS_UP |
| Enrichment Score (ES) | -0.71210164 |
| Normalized Enrichment Score (NES) | -1.6932075 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.17979105 |
| FWER p-Value | 0.984 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | FZD5 | 38 | 109.643 | 0.0297 | No | ||
| 2 | CITED2 | 143 | 54.021 | 0.0410 | No | ||
| 3 | FZD7 | 200 | 40.889 | 0.0505 | No | ||
| 4 | LRP8 | 208 | 39.535 | 0.0614 | No | ||
| 5 | VTI1B | 293 | 30.294 | 0.0668 | No | ||
| 6 | CMTM7 | 365 | 25.490 | 0.0713 | No | ||
| 7 | ARL6IP1 | 472 | 21.533 | 0.0733 | No | ||
| 8 | C6orf62 | 723 | 15.826 | 0.0682 | No | ||
| 9 | FUBP1 | 847 | 14.067 | 0.0675 | No | ||
| 10 | ADAMTS1 | 922 | 13.023 | 0.0684 | No | ||
| 11 | DHRS3 | 933 | 12.896 | 0.0717 | No | ||
| 12 | NQO2 | 988 | 12.307 | 0.0731 | No | ||
| 13 | KPNB1 | 995 | 12.236 | 0.0763 | No | ||
| 14 | MGAT5 | 1043 | 11.670 | 0.0778 | No | ||
| 15 | SCD | 1102 | 11.098 | 0.0787 | No | ||
| 16 | ZNF76 | 1176 | 10.523 | 0.0789 | No | ||
| 17 | IQCG | 1214 | 10.268 | 0.0804 | No | ||
| 18 | KLHL7 | 1246 | 9.949 | 0.0821 | No | ||
| 19 | IVNS1ABP | 1265 | 9.846 | 0.0842 | No | ||
| 20 | RAB8A | 1273 | 9.818 | 0.0867 | No | ||
| 21 | FKBP1A | 1342 | 9.289 | 0.0867 | No | ||
| 22 | BCL2L11 | 1528 | 8.185 | 0.0819 | No | ||
| 23 | MOCS2 | 1563 | 7.960 | 0.0829 | No | ||
| 24 | TFPI | 1640 | 7.533 | 0.0821 | No | ||
| 25 | TUSC2 | 1654 | 7.495 | 0.0837 | No | ||
| 26 | VAMP1 | 1711 | 7.211 | 0.0836 | No | ||
| 27 | NKAIN1 | 1725 | 7.159 | 0.0852 | No | ||
| 28 | TGFBR2 | 1740 | 7.119 | 0.0867 | No | ||
| 29 | PABPN1 | 1810 | 6.770 | 0.0859 | No | ||
| 30 | GRAMD1B | 2042 | 5.911 | 0.0788 | No | ||
| 31 | CTSC | 2055 | 5.878 | 0.0800 | No | ||
| 32 | IGF1R | 2148 | 5.588 | 0.0780 | No | ||
| 33 | PTP4A1 | 2168 | 5.519 | 0.0789 | No | ||
| 34 | CERK | 2493 | 4.644 | 0.0678 | No | ||
| 35 | RPIA | 2593 | 4.443 | 0.0652 | No | ||
| 36 | FAM171A1 | 2617 | 4.377 | 0.0656 | No | ||
| 37 | FAM105A | 2637 | 4.337 | 0.0661 | No | ||
| 38 | CDH24 | 2648 | 4.308 | 0.0669 | No | ||
| 39 | GCLC | 2659 | 4.289 | 0.0678 | No | ||
| 40 | INSR | 2697 | 4.181 | 0.0675 | No | ||
| 41 | STEAP3 | 2717 | 4.149 | 0.0680 | No | ||
| 42 | SYNCRIP | 2873 | 3.833 | 0.0631 | No | ||
| 43 | CDC26 | 2882 | 3.820 | 0.0639 | No | ||
| 44 | ITPR1 | 2995 | 3.635 | 0.0606 | No | ||
| 45 | HMOX1 | 3141 | 3.368 | 0.0560 | No | ||
| 46 | MIR17HG | 3274 | 3.178 | 0.0519 | No | ||
| 47 | ATF4 | 3319 | 3.105 | 0.0511 | No | ||
| 48 | ACVR2B | 3393 | 3.001 | 0.0491 | No | ||
| 49 | FAM173B | 3522 | 2.829 | 0.0450 | No | ||
| 50 | GSK3B | 3658 | 2.696 | 0.0406 | No | ||
| 51 | C9orf116 | 3667 | 2.689 | 0.0410 | No | ||
| 52 | ZP3 | 3845 | 2.475 | 0.0350 | No | ||
| 53 | FZD10 | 4037 | 2.285 | 0.0283 | No | ||
| 54 | CENPV | 4128 | 2.195 | 0.0255 | No | ||
| 55 | MCF2L | 4257 | 2.058 | 0.0211 | No | ||
| 56 | CALU | 4369 | 1.958 | 0.0174 | No | ||
| 57 | MAT2B | 4462 | 1.872 | 0.0144 | No | ||
| 58 | PABPC4 | 4481 | 1.862 | 0.0143 | No | ||
| 59 | RCOR3 | 4723 | 1.675 | 0.0055 | No | ||
| 60 | YKT6 | 4922 | 1.531 | -0.0017 | No | ||
| 61 | TSPAN13 | 4975 | 1.491 | -0.0032 | No | ||
| 62 | HLX | 5146 | 1.381 | -0.0094 | No | ||
| 63 | IQCK | 5285 | 1.307 | -0.0143 | No | ||
| 64 | ARHGAP28 | 5312 | 1.285 | -0.0149 | No | ||
| 65 | PDGFC | 5424 | 1.228 | -0.0188 | No | ||
| 66 | FBXW7 | 5460 | 1.204 | -0.0198 | No | ||
| 67 | DPCD | 5572 | 1.136 | -0.0238 | No | ||
| 68 | CHML | 5695 | 1.069 | -0.0281 | No | ||
| 69 | ACPP | 5847 | 0.991 | -0.0336 | No | ||
| 70 | SET | 5935 | 0.954 | -0.0367 | No | ||
| 71 | RASD1 | 6115 | 0.890 | -0.0433 | No | ||
| 72 | XYLT2 | 6249 | 0.839 | -0.0482 | No | ||
| 73 | FAM120C | 6268 | 0.834 | -0.0486 | No | ||
| 74 | CBX8 | 6304 | 0.823 | -0.0497 | No | ||
| 75 | RNLS | 6427 | 0.786 | -0.0542 | No | ||
| 76 | JAG2 | 6456 | 0.776 | -0.0551 | No | ||
| 77 | TBC1D4 | 6526 | 0.754 | -0.0575 | No | ||
| 78 | PTPN2 | 6643 | 0.712 | -0.0617 | No | ||
| 79 | DCBLD1 | 6740 | 0.673 | -0.0652 | No | ||
| 80 | SLC19A2 | 6796 | 0.660 | -0.0671 | No | ||
| 81 | GPR180 | 7261 | 0.536 | -0.0848 | No | ||
| 82 | CNOT6 | 7450 | 0.498 | -0.0919 | No | ||
| 83 | NLN | 7555 | 0.474 | -0.0957 | No | ||
| 84 | CLN3 | 7582 | 0.466 | -0.0966 | No | ||
| 85 | ID1 | 7629 | 0.457 | -0.0982 | No | ||
| 86 | TSPAN3 | 7636 | 0.456 | -0.0983 | No | ||
| 87 | DUSP2 | 7799 | 0.427 | -0.1044 | No | ||
| 88 | ATF4P3 | 7872 | 0.417 | -0.1070 | No | ||
| 89 | UCP3 | 8630 | 0.305 | -0.1360 | No | ||
| 90 | MND1 | 8687 | 0.299 | -0.1381 | No | ||
| 91 | KLHL15 | 8831 | 0.280 | -0.1435 | No | ||
| 92 | KCNK3 | 8877 | 0.274 | -0.1451 | No | ||
| 93 | CALML4 | 8889 | 0.273 | -0.1454 | No | ||
| 94 | C11orf57 | 8922 | 0.269 | -0.1466 | No | ||
| 95 | FNBP1L | 9119 | 0.247 | -0.1540 | No | ||
| 96 | NEK10 | 9338 | 0.222 | -0.1623 | No | ||
| 97 | NQO1 | 9499 | 0.207 | -0.1684 | No | ||
| 98 | SLC16A1 | 9606 | 0.195 | -0.1724 | No | ||
| 99 | NEK4 | 9830 | 0.176 | -0.1809 | No | ||
| 100 | BMP6 | 9894 | 0.171 | -0.1833 | No | ||
| 101 | KLRG1 | 11244 | 0.080 | -0.2350 | No | ||
| 102 | PTGES2 | 11447 | 0.068 | -0.2427 | No | ||
| 103 | SERF1B | 11505 | 0.065 | -0.2449 | No | ||
| 104 | CRB1 | 11609 | 0.060 | -0.2488 | No | ||
| 105 | RELL1 | 11680 | 0.056 | -0.2515 | No | ||
| 106 | DDX3X | 11986 | 0.044 | -0.2632 | No | ||
| 107 | SPA17 | 12376 | 0.027 | -0.2781 | No | ||
| 108 | RASSF9 | 12435 | 0.025 | -0.2803 | No | ||
| 109 | SMC5 | 12541 | 0.021 | -0.2843 | No | ||
| 110 | FBXW10 | 12696 | 0.015 | -0.2902 | No | ||
| 111 | LYG1 | 12883 | 0.007 | -0.2974 | No | ||
| 112 | NFYB | 12995 | 0.005 | -0.3016 | No | ||
| 113 | KRTDAP | 13237 | -0.004 | -0.3109 | No | ||
| 114 | TCP10L | 13769 | -0.023 | -0.3312 | No | ||
| 115 | HBP1 | 13784 | -0.023 | -0.3318 | No | ||
| 116 | PDK4 | 13791 | -0.024 | -0.3320 | No | ||
| 117 | XIAP | 14046 | -0.034 | -0.3417 | No | ||
| 118 | CCNYL1 | 14301 | -0.044 | -0.3514 | No | ||
| 119 | CCNY | 14406 | -0.048 | -0.3554 | No | ||
| 120 | PDE1A | 14543 | -0.052 | -0.3606 | No | ||
| 121 | ZG16B | 14580 | -0.053 | -0.3620 | No | ||
| 122 | WWP1 | 15138 | -0.081 | -0.3833 | No | ||
| 123 | HMGB2 | 15209 | -0.085 | -0.3860 | No | ||
| 124 | LYRM7 | 15279 | -0.088 | -0.3886 | No | ||
| 125 | CDC37L1 | 15503 | -0.102 | -0.3971 | No | ||
| 126 | TNPO1 | 15597 | -0.108 | -0.4007 | No | ||
| 127 | MRRF | 16093 | -0.140 | -0.4196 | No | ||
| 128 | OSCP1 | 16145 | -0.144 | -0.4215 | No | ||
| 129 | MCM3AP-AS1 | 16270 | -0.152 | -0.4262 | No | ||
| 130 | NR0B1 | 16431 | -0.164 | -0.4323 | No | ||
| 131 | MOK | 16452 | -0.166 | -0.4330 | No | ||
| 132 | FAM60A | 16563 | -0.175 | -0.4372 | No | ||
| 133 | A2M | 16583 | -0.178 | -0.4379 | No | ||
| 134 | RNF141 | 16632 | -0.183 | -0.4397 | No | ||
| 135 | BNIP2 | 16702 | -0.189 | -0.4423 | No | ||
| 136 | LZTFL1 | 16900 | -0.208 | -0.4498 | No | ||
| 137 | ENAH | 16954 | -0.213 | -0.4517 | No | ||
| 138 | RIMS4 | 17054 | -0.223 | -0.4555 | No | ||
| 139 | CBX5 | 17102 | -0.230 | -0.4572 | No | ||
| 140 | TCTEX1D2 | 17263 | -0.245 | -0.4633 | No | ||
| 141 | TMTC4 | 17489 | -0.274 | -0.4718 | No | ||
| 142 | YES1 | 17492 | -0.274 | -0.4718 | No | ||
| 143 | LIN9 | 17505 | -0.275 | -0.4722 | No | ||
| 144 | SLC2A3 | 18053 | -0.342 | -0.4931 | No | ||
| 145 | SUZ12 | 18093 | -0.347 | -0.4945 | No | ||
| 146 | DCAF17 | 18266 | -0.375 | -0.5010 | No | ||
| 147 | HEY1 | 18340 | -0.387 | -0.5037 | No | ||
| 148 | GABPB1 | 18483 | -0.413 | -0.5090 | No | ||
| 149 | PTPRJ | 18542 | -0.424 | -0.5111 | No | ||
| 150 | PDHX | 18835 | -0.480 | -0.5222 | No | ||
| 151 | ECM2 | 18915 | -0.498 | -0.5251 | No | ||
| 152 | AKR1C1 | 19331 | -0.586 | -0.5408 | No | ||
| 153 | MAN1A1 | 19387 | -0.603 | -0.5427 | No | ||
| 154 | CCDC6 | 19444 | -0.616 | -0.5447 | No | ||
| 155 | TFPI2 | 19448 | -0.618 | -0.5447 | No | ||
| 156 | MBTD1 | 19495 | -0.629 | -0.5462 | No | ||
| 157 | SEC62 | 19620 | -0.664 | -0.5508 | No | ||
| 158 | RASA4 | 19832 | -0.731 | -0.5587 | No | ||
| 159 | RBM33 | 19846 | -0.733 | -0.5590 | No | ||
| 160 | DNAJB4 | 19862 | -0.738 | -0.5593 | No | ||
| 161 | ECE1 | 20039 | -0.794 | -0.5659 | No | ||
| 162 | TMEM79 | 20100 | -0.811 | -0.5679 | No | ||
| 163 | SLC16A6 | 20442 | -0.938 | -0.5808 | No | ||
| 164 | TNFRSF11B | 20444 | -0.938 | -0.5805 | No | ||
| 165 | STX6 | 20551 | -0.981 | -0.5843 | No | ||
| 166 | SORBS2 | 20831 | -1.108 | -0.5947 | No | ||
| 167 | BET1 | 20912 | -1.144 | -0.5974 | No | ||
| 168 | DPY19L4 | 21056 | -1.226 | -0.6026 | No | ||
| 169 | FAM86C1 | 21238 | -1.332 | -0.6091 | No | ||
| 170 | TPCN1 | 21306 | -1.366 | -0.6113 | No | ||
| 171 | PIAS2 | 21474 | -1.471 | -0.6173 | No | ||
| 172 | SYT15 | 21506 | -1.488 | -0.6181 | No | ||
| 173 | ZNF721 | 21519 | -1.494 | -0.6181 | No | ||
| 174 | RAB10 | 21628 | -1.564 | -0.6218 | No | ||
| 175 | TOP2B | 21868 | -1.742 | -0.6305 | No | ||
| 176 | SSU72 | 21877 | -1.748 | -0.6303 | No | ||
| 177 | BID | 22004 | -1.869 | -0.6346 | No | ||
| 178 | SERF1A | 22007 | -1.872 | -0.6341 | No | ||
| 179 | SETD4 | 22251 | -2.078 | -0.6429 | No | ||
| 180 | LINC00116 | 22495 | -2.324 | -0.6515 | No | ||
| 181 | MRPL42 | 22541 | -2.367 | -0.6526 | No | ||
| 182 | MLXIPL | 22964 | -2.895 | -0.6679 | No | ||
| 183 | ANKRD46 | 23064 | -3.030 | -0.6709 | No | ||
| 184 | JMY | 23274 | -3.370 | -0.6779 | No | ||
| 185 | SERP1 | 23293 | -3.403 | -0.6777 | No | ||
| 186 | ZNF280C | 23303 | -3.423 | -0.6770 | No | ||
| 187 | PDE4D | 23393 | -3.607 | -0.6794 | No | ||
| 188 | VAV3 | 23486 | -3.765 | -0.6819 | No | ||
| 189 | EIF2AK4 | 23487 | -3.765 | -0.6808 | No | ||
| 190 | NMNAT1 | 23557 | -3.917 | -0.6824 | No | ||
| 191 | TMEM116 | 23609 | -4.020 | -0.6832 | No | ||
| 192 | GABPB2 | 23614 | -4.040 | -0.6822 | No | ||
| 193 | RERG | 23653 | -4.122 | -0.6825 | No | ||
| 194 | CCDC77 | 23665 | -4.142 | -0.6817 | No | ||
| 195 | FAM86B1 | 23843 | -4.575 | -0.6872 | No | ||
| 196 | ZNF267 | 24029 | -5.126 | -0.6928 | No | ||
| 197 | RBKS | 24139 | -5.407 | -0.6955 | No | ||
| 198 | EFNA1 | 24176 | -5.516 | -0.6953 | No | ||
| 199 | PHGDH | 24203 | -5.598 | -0.6947 | No | ||
| 200 | PCSK1 | 24286 | -5.940 | -0.6962 | No | ||
| 201 | KLF9 | 24326 | -6.110 | -0.6959 | No | ||
| 202 | DCAF4 | 24371 | -6.276 | -0.6958 | No | ||
| 203 | NSRP1 | 24581 | -7.143 | -0.7018 | No | ||
| 204 | RIMKLB | 24690 | -7.610 | -0.7038 | No | ||
| 205 | EIF5 | 24866 | -8.648 | -0.7081 | No | ||
| 206 | NUPR1 | 24923 | -9.037 | -0.7077 | No | ||
| 207 | HTRA3 | 25040 | -10.089 | -0.7092 | Yes | ||
| 208 | GKAP1 | 25111 | -10.682 | -0.7089 | Yes | ||
| 209 | TIPIN | 25187 | -11.436 | -0.7085 | Yes | ||
| 210 | MAP2 | 25191 | -11.477 | -0.7054 | Yes | ||
| 211 | HINT3 | 25192 | -11.494 | -0.7021 | Yes | ||
| 212 | E2F5 | 25198 | -11.551 | -0.6990 | Yes | ||
| 213 | BMP2 | 25202 | -11.606 | -0.6959 | Yes | ||
| 214 | DDAH1 | 25308 | -12.795 | -0.6963 | Yes | ||
| 215 | MAP7 | 25471 | -14.937 | -0.6982 | Yes | ||
| 216 | NPTX1 | 25528 | -15.889 | -0.6959 | Yes | ||
| 217 | STOX2 | 25537 | -16.055 | -0.6916 | Yes | ||
| 218 | LMO4 | 25682 | -19.607 | -0.6916 | Yes | ||
| 219 | LINC00467 | 25724 | -20.696 | -0.6873 | Yes | ||
| 220 | CTGF | 25736 | -21.035 | -0.6817 | Yes | ||
| 221 | PYCR1 | 25769 | -21.972 | -0.6767 | Yes | ||
| 222 | NCOA7 | 25853 | -25.364 | -0.6727 | Yes | ||
| 223 | SGPL1 | 25871 | -25.891 | -0.6660 | Yes | ||
| 224 | GRB10 | 25941 | -30.158 | -0.6601 | Yes | ||
| 225 | NR4A1 | 25994 | -33.453 | -0.6526 | Yes | ||
| 226 | IL20RB | 26119 | -46.747 | -0.6441 | Yes | ||
| 227 | SLC7A11 | 26126 | -47.089 | -0.6310 | Yes | ||
| 228 | KRCC1 | 26142 | -50.080 | -0.6173 | Yes | ||
| 229 | HERPUD1 | 26159 | -53.015 | -0.6029 | Yes | ||
| 230 | VLDLR | 26190 | -59.843 | -0.5871 | Yes | ||
| 231 | DUSP16 | 26195 | -61.523 | -0.5697 | Yes | ||
| 232 | ASNS | 26241 | -83.978 | -0.5476 | Yes | ||
| 233 | VEGFA | 26251 | -89.987 | -0.5224 | Yes | ||
| 234 | MTHFD2 | 26257 | -95.222 | -0.4956 | Yes | ||
| 235 | STC2 | 26260 | -97.527 | -0.4680 | Yes | ||
| 236 | SESN2 | 26273 | -109.055 | -0.4375 | Yes | ||
| 237 | GPT2 | 26276 | -117.291 | -0.4043 | Yes | ||
| 238 | ALDH1L2 | 26287 | -152.864 | -0.3613 | Yes | ||
| 239 | TXNIP | 26293 | -180.286 | -0.3104 | Yes | ||
| 240 | PCK2 | 26298 | -198.705 | -0.2541 | Yes | ||
| 241 | DDIT4 | 26311 | -299.207 | -0.1697 | Yes | ||
| 242 | INHBE | 26312 | -299.207 | -0.0848 | Yes | ||
| 243 | CHAC1 | 26314 | -299.207 | 0.0001 | Yes |