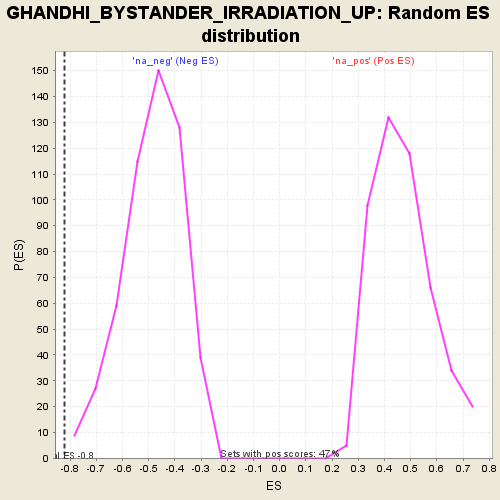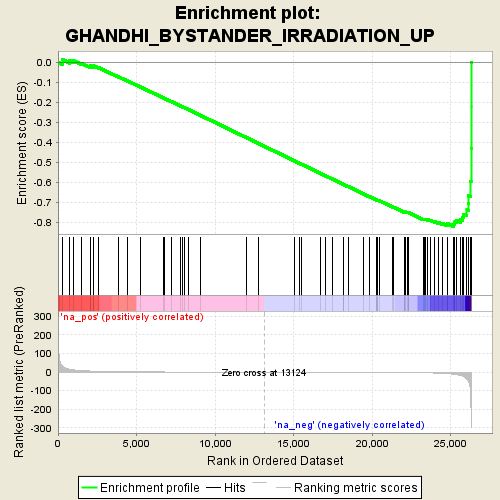
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | NS50832_padj |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_neg |
| GeneSet | GHANDHI_BYSTANDER_IRRADIATION_UP |
| Enrichment Score (ES) | -0.8198635 |
| Normalized Enrichment Score (NES) | -1.694819 |
| Nominal p-value | 0.0018975332 |
| FDR q-value | 0.18162522 |
| FWER p-Value | 0.981 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | NFKBIZ | 270 | 32.545 | 0.0145 | No | ||
| 2 | STC1 | 740 | 15.599 | 0.0085 | No | ||
| 3 | LIF | 967 | 12.489 | 0.0094 | No | ||
| 4 | ATP6V1C1 | 1506 | 8.259 | -0.0048 | No | ||
| 5 | B3GNT5 | 2035 | 5.934 | -0.0204 | No | ||
| 6 | FGF2 | 2037 | 5.925 | -0.0159 | No | ||
| 7 | ZCCHC6 | 2263 | 5.228 | -0.0205 | No | ||
| 8 | INHBA | 2282 | 5.152 | -0.0173 | No | ||
| 9 | PLEK2 | 2556 | 4.522 | -0.0242 | No | ||
| 10 | RGS9 | 3838 | 2.487 | -0.0711 | No | ||
| 11 | SLC16A3 | 4393 | 1.933 | -0.0908 | No | ||
| 12 | MMP10 | 5239 | 1.331 | -0.1220 | No | ||
| 13 | PTGES | 6696 | 0.689 | -0.1769 | No | ||
| 14 | AHR | 6763 | 0.668 | -0.1789 | No | ||
| 15 | C2CD4B | 7246 | 0.539 | -0.1969 | No | ||
| 16 | DUSP2 | 7799 | 0.427 | -0.2176 | No | ||
| 17 | MCTP1 | 7915 | 0.410 | -0.2216 | No | ||
| 18 | SLC39A8 | 8030 | 0.392 | -0.2257 | No | ||
| 19 | LRRC8B | 8283 | 0.357 | -0.2350 | No | ||
| 20 | RELB | 9064 | 0.253 | -0.2645 | No | ||
| 21 | PHEX | 11973 | 0.045 | -0.3753 | No | ||
| 22 | SAT1 | 12782 | 0.011 | -0.4061 | No | ||
| 23 | CCK | 15027 | -0.075 | -0.4915 | No | ||
| 24 | IFNE | 15346 | -0.094 | -0.5035 | No | ||
| 25 | LAMB3 | 15522 | -0.104 | -0.5101 | No | ||
| 26 | MMP3 | 16692 | -0.189 | -0.5545 | No | ||
| 27 | C8orf4 | 17032 | -0.220 | -0.5673 | No | ||
| 28 | KYNU | 17504 | -0.275 | -0.5850 | No | ||
| 29 | PLA2G4A | 18148 | -0.355 | -0.6092 | No | ||
| 30 | MCTP2 | 18206 | -0.364 | -0.6111 | No | ||
| 31 | BIRC3 | 18505 | -0.418 | -0.6222 | No | ||
| 32 | ZC3H12C | 19462 | -0.621 | -0.6581 | No | ||
| 33 | DNAJB4 | 19862 | -0.738 | -0.6727 | No | ||
| 34 | MT1L | 20272 | -0.876 | -0.6877 | No | ||
| 35 | SRSF12 | 20296 | -0.883 | -0.6879 | No | ||
| 36 | SOD2 | 20330 | -0.895 | -0.6884 | No | ||
| 37 | NAMPT | 20349 | -0.904 | -0.6884 | No | ||
| 38 | SLC16A6 | 20442 | -0.938 | -0.6912 | No | ||
| 39 | GCLM | 21297 | -1.363 | -0.7227 | No | ||
| 40 | EPB41L4B | 21380 | -1.411 | -0.7248 | No | ||
| 41 | OSGIN2 | 22052 | -1.906 | -0.7489 | No | ||
| 42 | PTGS2 | 22073 | -1.921 | -0.7482 | No | ||
| 43 | TNFAIP3 | 22114 | -1.957 | -0.7482 | No | ||
| 44 | TSLP | 22245 | -2.072 | -0.7516 | No | ||
| 45 | CXCL2 | 22288 | -2.121 | -0.7516 | No | ||
| 46 | GJB2 | 22331 | -2.158 | -0.7515 | No | ||
| 47 | MT1H | 23299 | -3.419 | -0.7858 | No | ||
| 48 | POPDC3 | 23338 | -3.483 | -0.7846 | No | ||
| 49 | PTGFR | 23423 | -3.658 | -0.7850 | No | ||
| 50 | SLC6A15 | 23553 | -3.909 | -0.7869 | No | ||
| 51 | C9orf72 | 23691 | -4.212 | -0.7889 | No | ||
| 52 | IL1A | 23980 | -4.957 | -0.7961 | No | ||
| 53 | ICAM1 | 24196 | -5.579 | -0.8001 | No | ||
| 54 | GPR68 | 24457 | -6.568 | -0.8050 | No | ||
| 55 | GCH1 | 24774 | -8.103 | -0.8108 | No | ||
| 56 | CXCL1 | 24785 | -8.163 | -0.8050 | No | ||
| 57 | RNF128 | 25176 | -11.335 | -0.8112 | Yes | ||
| 58 | DUSP5 | 25225 | -11.830 | -0.8041 | Yes | ||
| 59 | IRAK3 | 25275 | -12.366 | -0.7965 | Yes | ||
| 60 | CREB5 | 25351 | -13.315 | -0.7892 | Yes | ||
| 61 | CXCL3 | 25645 | -18.583 | -0.7862 | Yes | ||
| 62 | MT1E | 25733 | -20.906 | -0.7736 | Yes | ||
| 63 | TESC | 25832 | -24.147 | -0.7590 | Yes | ||
| 64 | SLC39A14 | 26039 | -37.712 | -0.7381 | Yes | ||
| 65 | SLC7A11 | 26126 | -47.089 | -0.7055 | Yes | ||
| 66 | GADD45G | 26144 | -50.686 | -0.6676 | Yes | ||
| 67 | MT1G | 26268 | -103.406 | -0.5935 | Yes | ||
| 68 | G0S2 | 26301 | -213.686 | -0.4320 | Yes | ||
| 69 | CXCL5 | 26306 | -277.394 | -0.2209 | Yes | ||
| 70 | MT1X | 26307 | -290.514 | 0.0004 | Yes |