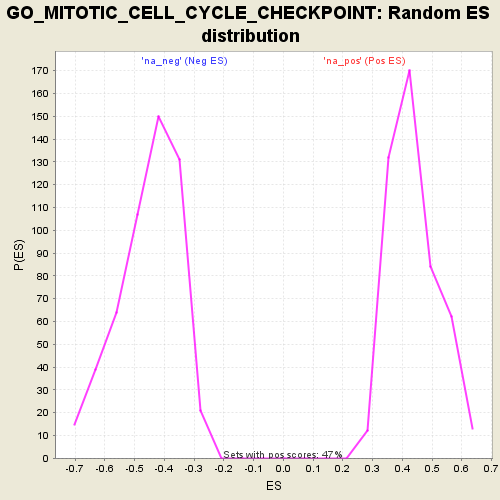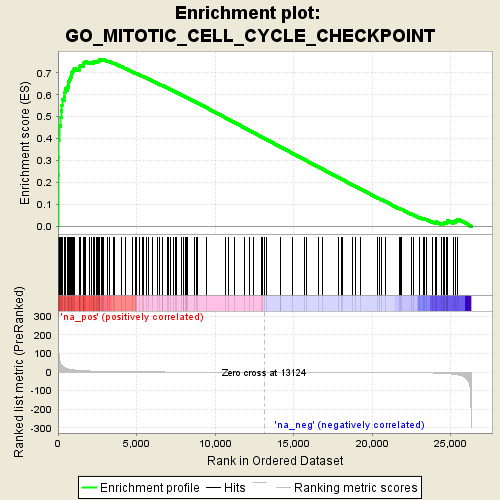
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | NS50832_padj |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_pos |
| GeneSet | GO_MITOTIC_CELL_CYCLE_CHECKPOINT |
| Enrichment Score (ES) | 0.7602351 |
| Normalized Enrichment Score (NES) | 1.7396506 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.13268022 |
| FWER p-Value | 0.754 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | CDKN1A | 2 | 299.207 | 0.2335 | Yes | ||
| 2 | RPS27L | 40 | 106.999 | 0.3156 | Yes | ||
| 3 | BAX | 53 | 95.755 | 0.3899 | Yes | ||
| 4 | BTG2 | 60 | 90.877 | 0.4606 | Yes | ||
| 5 | MDM2 | 156 | 51.042 | 0.4969 | Yes | ||
| 6 | XPC | 204 | 40.284 | 0.5265 | Yes | ||
| 7 | TRIAP1 | 239 | 35.685 | 0.5531 | Yes | ||
| 8 | PCNA | 255 | 34.423 | 0.5794 | Yes | ||
| 9 | CCNB1 | 387 | 24.655 | 0.5936 | Yes | ||
| 10 | GTSE1 | 394 | 24.176 | 0.6123 | Yes | ||
| 11 | PLK2 | 461 | 21.783 | 0.6267 | Yes | ||
| 12 | SFN | 603 | 18.262 | 0.6356 | Yes | ||
| 13 | RBM38 | 652 | 17.278 | 0.6473 | Yes | ||
| 14 | CDK1 | 676 | 16.831 | 0.6595 | Yes | ||
| 15 | PLK3 | 703 | 16.241 | 0.6712 | Yes | ||
| 16 | CASP2 | 802 | 14.851 | 0.6791 | Yes | ||
| 17 | E2F7 | 830 | 14.296 | 0.6892 | Yes | ||
| 18 | CLSPN | 837 | 14.160 | 0.7000 | Yes | ||
| 19 | TGFB1 | 913 | 13.218 | 0.7075 | Yes | ||
| 20 | CRADD | 1008 | 11.968 | 0.7132 | Yes | ||
| 21 | FOXO4 | 1024 | 11.812 | 0.7219 | Yes | ||
| 22 | MAD2L1 | 1335 | 9.368 | 0.7174 | Yes | ||
| 23 | KLHL22 | 1352 | 9.207 | 0.7239 | Yes | ||
| 24 | CCNA2 | 1369 | 9.062 | 0.7304 | Yes | ||
| 25 | RFWD3 | 1438 | 8.664 | 0.7346 | Yes | ||
| 26 | ZNF207 | 1597 | 7.765 | 0.7346 | Yes | ||
| 27 | FBXO31 | 1619 | 7.654 | 0.7398 | Yes | ||
| 28 | PRMT1 | 1642 | 7.527 | 0.7448 | Yes | ||
| 29 | SOX4 | 1692 | 7.315 | 0.7486 | Yes | ||
| 30 | CENPF | 1716 | 7.186 | 0.7534 | Yes | ||
| 31 | E2F4 | 2009 | 6.019 | 0.7469 | Yes | ||
| 32 | HRAS | 2126 | 5.642 | 0.7469 | Yes | ||
| 33 | PSMG2 | 2264 | 5.218 | 0.7457 | Yes | ||
| 34 | HMGA2 | 2268 | 5.208 | 0.7497 | Yes | ||
| 35 | FOXN3 | 2329 | 5.023 | 0.7513 | Yes | ||
| 36 | GIGYF2 | 2422 | 4.780 | 0.7515 | Yes | ||
| 37 | TP73 | 2507 | 4.622 | 0.7519 | Yes | ||
| 38 | TTK | 2547 | 4.538 | 0.7540 | Yes | ||
| 39 | TERF1 | 2628 | 4.350 | 0.7543 | Yes | ||
| 40 | HUS1 | 2636 | 4.337 | 0.7575 | Yes | ||
| 41 | MAD2L1BP | 2662 | 4.285 | 0.7599 | Yes | ||
| 42 | CDK2 | 2749 | 4.087 | 0.7598 | Yes | ||
| 43 | CENPJ | 2831 | 3.907 | 0.7597 | Yes | ||
| 44 | SMC1A | 2896 | 3.795 | 0.7602 | Yes | ||
| 45 | ZNF385A | 3164 | 3.340 | 0.7526 | No | ||
| 46 | UBB | 3255 | 3.201 | 0.7517 | No | ||
| 47 | CCND1 | 3538 | 2.814 | 0.7431 | No | ||
| 48 | UBA52 | 3612 | 2.746 | 0.7425 | No | ||
| 49 | CNOT8 | 4010 | 2.306 | 0.7291 | No | ||
| 50 | MDC1 | 4291 | 2.024 | 0.7200 | No | ||
| 51 | ZW10 | 4769 | 1.642 | 0.7031 | No | ||
| 52 | ARID3A | 4941 | 1.515 | 0.6977 | No | ||
| 53 | E2F1 | 4963 | 1.500 | 0.6981 | No | ||
| 54 | APC | 5170 | 1.370 | 0.6913 | No | ||
| 55 | PML | 5388 | 1.243 | 0.6840 | No | ||
| 56 | RPS6 | 5456 | 1.206 | 0.6824 | No | ||
| 57 | CARM1 | 5636 | 1.100 | 0.6764 | No | ||
| 58 | WNT9A | 5781 | 1.030 | 0.6717 | No | ||
| 59 | TOP2A | 6015 | 0.925 | 0.6635 | No | ||
| 60 | CDKN2B | 6331 | 0.813 | 0.6522 | No | ||
| 61 | AURKA | 6451 | 0.778 | 0.6482 | No | ||
| 62 | NAE1 | 6621 | 0.718 | 0.6423 | No | ||
| 63 | ZWINT | 6636 | 0.714 | 0.6423 | No | ||
| 64 | TFDP2 | 6940 | 0.619 | 0.6313 | No | ||
| 65 | SYF2 | 7031 | 0.593 | 0.6283 | No | ||
| 66 | CDK5RAP3 | 7133 | 0.567 | 0.6249 | No | ||
| 67 | PRCC | 7372 | 0.513 | 0.6162 | No | ||
| 68 | CNOT6 | 7450 | 0.498 | 0.6136 | No | ||
| 69 | TPR | 7525 | 0.481 | 0.6112 | No | ||
| 70 | EP300 | 7882 | 0.416 | 0.5979 | No | ||
| 71 | RPS27A | 7967 | 0.401 | 0.5950 | No | ||
| 72 | NPM1 | 8001 | 0.394 | 0.5941 | No | ||
| 73 | ZNF830 | 8127 | 0.376 | 0.5896 | No | ||
| 74 | INTS3 | 8168 | 0.371 | 0.5884 | No | ||
| 75 | HUS1B | 8191 | 0.368 | 0.5878 | No | ||
| 76 | BUB1B | 8264 | 0.360 | 0.5853 | No | ||
| 77 | CHEK2 | 8686 | 0.299 | 0.5695 | No | ||
| 78 | UBC | 8798 | 0.286 | 0.5655 | No | ||
| 79 | PRKDC | 8863 | 0.276 | 0.5632 | No | ||
| 80 | BUB1 | 9451 | 0.212 | 0.5410 | No | ||
| 81 | BLM | 10670 | 0.112 | 0.4946 | No | ||
| 82 | DLG1 | 10821 | 0.102 | 0.4889 | No | ||
| 83 | TNKS1BP1 | 11231 | 0.081 | 0.4734 | No | ||
| 84 | GADD45A | 11891 | 0.047 | 0.4483 | No | ||
| 85 | CNOT1 | 12196 | 0.035 | 0.4367 | No | ||
| 86 | CNOT2 | 12436 | 0.025 | 0.4276 | No | ||
| 87 | NEK11 | 12948 | 0.006 | 0.4081 | No | ||
| 88 | MDM4 | 13001 | 0.005 | 0.4061 | No | ||
| 89 | RB1 | 13128 | -0.000 | 0.4013 | No | ||
| 90 | ZWILCH | 13284 | -0.005 | 0.3954 | No | ||
| 91 | CSNK2A1 | 14143 | -0.038 | 0.3626 | No | ||
| 92 | PLAGL1 | 14158 | -0.039 | 0.3621 | No | ||
| 93 | PLK1 | 14929 | -0.069 | 0.3328 | No | ||
| 94 | ATF2 | 15697 | -0.113 | 0.3036 | No | ||
| 95 | PPP2R5C | 15792 | -0.120 | 0.3001 | No | ||
| 96 | KNTC1 | 16608 | -0.180 | 0.2691 | No | ||
| 97 | ATM | 16808 | -0.200 | 0.2617 | No | ||
| 98 | SETMAR | 17845 | -0.316 | 0.2223 | No | ||
| 99 | RAD9B | 18061 | -0.343 | 0.2144 | No | ||
| 100 | CNOT3 | 18104 | -0.349 | 0.2131 | No | ||
| 101 | CNOT4 | 18759 | -0.466 | 0.1885 | No | ||
| 102 | DGKZ | 18924 | -0.501 | 0.1826 | No | ||
| 103 | CHFR | 19283 | -0.574 | 0.1694 | No | ||
| 104 | CNOT10 | 20323 | -0.892 | 0.1304 | No | ||
| 105 | MAD1L1 | 20474 | -0.951 | 0.1254 | No | ||
| 106 | CNOT7 | 20626 | -1.009 | 0.1204 | No | ||
| 107 | FANCI | 20836 | -1.109 | 0.1133 | No | ||
| 108 | CSNK2A2 | 21713 | -1.630 | 0.0811 | No | ||
| 109 | BCL2L1 | 21802 | -1.687 | 0.0791 | No | ||
| 110 | TOP2B | 21868 | -1.742 | 0.0780 | No | ||
| 111 | CDC25C | 22485 | -2.318 | 0.0563 | No | ||
| 112 | RPA2 | 22655 | -2.518 | 0.0518 | No | ||
| 113 | BUB3 | 23035 | -2.995 | 0.0397 | No | ||
| 114 | CDKN1B | 23273 | -3.364 | 0.0332 | No | ||
| 115 | TFDP1 | 23324 | -3.462 | 0.0340 | No | ||
| 116 | RBL2 | 23437 | -3.679 | 0.0326 | No | ||
| 117 | NBN | 23844 | -4.578 | 0.0207 | No | ||
| 118 | MAD2L2 | 24038 | -5.145 | 0.0173 | No | ||
| 119 | MUC1 | 24081 | -5.263 | 0.0198 | No | ||
| 120 | RAD17 | 24393 | -6.352 | 0.0129 | No | ||
| 121 | CNOT6L | 24555 | -7.042 | 0.0123 | No | ||
| 122 | RAD9A | 24579 | -7.142 | 0.0170 | No | ||
| 123 | TEX14 | 24718 | -7.773 | 0.0178 | No | ||
| 124 | MSH2 | 24778 | -8.121 | 0.0219 | No | ||
| 125 | TAOK3 | 24789 | -8.188 | 0.0279 | No | ||
| 126 | TIPIN | 25187 | -11.436 | 0.0216 | No | ||
| 127 | TP53 | 25300 | -12.686 | 0.0273 | No | ||
| 128 | PCBP4 | 25454 | -14.763 | 0.0330 | No |