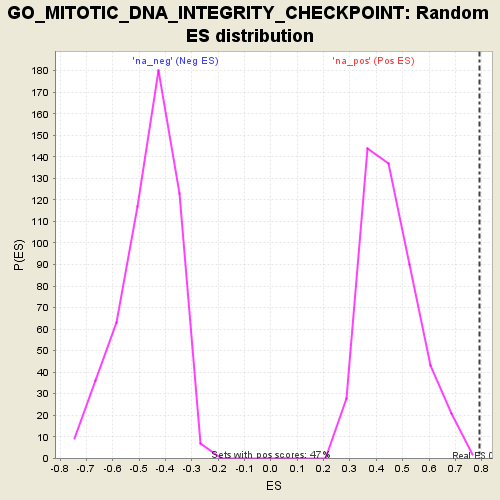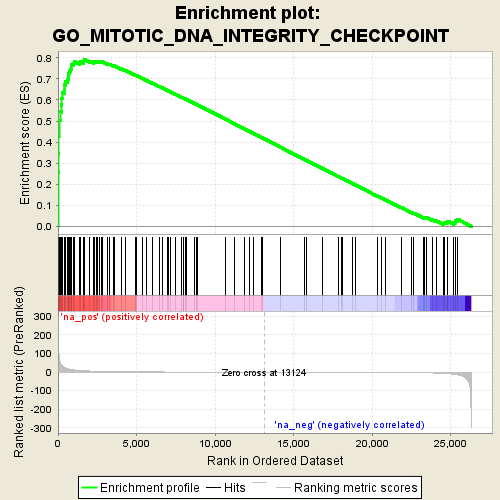
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | NS50832_padj |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_pos |
| GeneSet | GO_MITOTIC_DNA_INTEGRITY_CHECKPOINT |
| Enrichment Score (ES) | 0.7916707 |
| Normalized Enrichment Score (NES) | 1.7431179 |
| Nominal p-value | 0.0021505377 |
| FDR q-value | 0.1417673 |
| FWER p-Value | 0.726 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | CDKN1A | 2 | 299.207 | 0.2556 | Yes | ||
| 2 | RPS27L | 40 | 106.999 | 0.3456 | Yes | ||
| 3 | BAX | 53 | 95.755 | 0.4270 | Yes | ||
| 4 | BTG2 | 60 | 90.877 | 0.5045 | Yes | ||
| 5 | MDM2 | 156 | 51.042 | 0.5445 | Yes | ||
| 6 | XPC | 204 | 40.284 | 0.5771 | Yes | ||
| 7 | TRIAP1 | 239 | 35.685 | 0.6063 | Yes | ||
| 8 | PCNA | 255 | 34.423 | 0.6351 | Yes | ||
| 9 | CCNB1 | 387 | 24.655 | 0.6512 | Yes | ||
| 10 | GTSE1 | 394 | 24.176 | 0.6716 | Yes | ||
| 11 | PLK2 | 461 | 21.783 | 0.6877 | Yes | ||
| 12 | SFN | 603 | 18.262 | 0.6980 | Yes | ||
| 13 | RBM38 | 652 | 17.278 | 0.7109 | Yes | ||
| 14 | CDK1 | 676 | 16.831 | 0.7244 | Yes | ||
| 15 | PLK3 | 703 | 16.241 | 0.7373 | Yes | ||
| 16 | CASP2 | 802 | 14.851 | 0.7463 | Yes | ||
| 17 | E2F7 | 830 | 14.296 | 0.7574 | Yes | ||
| 18 | CLSPN | 837 | 14.160 | 0.7693 | Yes | ||
| 19 | CRADD | 1008 | 11.968 | 0.7731 | Yes | ||
| 20 | FOXO4 | 1024 | 11.812 | 0.7826 | Yes | ||
| 21 | CCNA2 | 1369 | 9.062 | 0.7772 | Yes | ||
| 22 | RFWD3 | 1438 | 8.664 | 0.7820 | Yes | ||
| 23 | FBXO31 | 1619 | 7.654 | 0.7817 | Yes | ||
| 24 | PRMT1 | 1642 | 7.527 | 0.7873 | Yes | ||
| 25 | SOX4 | 1692 | 7.315 | 0.7917 | Yes | ||
| 26 | E2F4 | 2009 | 6.019 | 0.7848 | No | ||
| 27 | HMGA2 | 2268 | 5.208 | 0.7794 | No | ||
| 28 | FOXN3 | 2329 | 5.023 | 0.7814 | No | ||
| 29 | GIGYF2 | 2422 | 4.780 | 0.7820 | No | ||
| 30 | TP73 | 2507 | 4.622 | 0.7827 | No | ||
| 31 | HUS1 | 2636 | 4.337 | 0.7815 | No | ||
| 32 | CDK2 | 2749 | 4.087 | 0.7808 | No | ||
| 33 | CENPJ | 2831 | 3.907 | 0.7810 | No | ||
| 34 | ZNF385A | 3164 | 3.340 | 0.7712 | No | ||
| 35 | UBB | 3255 | 3.201 | 0.7705 | No | ||
| 36 | CCND1 | 3538 | 2.814 | 0.7622 | No | ||
| 37 | UBA52 | 3612 | 2.746 | 0.7617 | No | ||
| 38 | CNOT8 | 4010 | 2.306 | 0.7486 | No | ||
| 39 | MDC1 | 4291 | 2.024 | 0.7396 | No | ||
| 40 | ARID3A | 4941 | 1.515 | 0.7162 | No | ||
| 41 | E2F1 | 4963 | 1.500 | 0.7166 | No | ||
| 42 | PML | 5388 | 1.243 | 0.7015 | No | ||
| 43 | CARM1 | 5636 | 1.100 | 0.6931 | No | ||
| 44 | TOP2A | 6015 | 0.925 | 0.6794 | No | ||
| 45 | AURKA | 6451 | 0.778 | 0.6635 | No | ||
| 46 | NAE1 | 6621 | 0.718 | 0.6577 | No | ||
| 47 | TFDP2 | 6940 | 0.619 | 0.6461 | No | ||
| 48 | SYF2 | 7031 | 0.593 | 0.6432 | No | ||
| 49 | CDK5RAP3 | 7133 | 0.567 | 0.6398 | No | ||
| 50 | CNOT6 | 7450 | 0.498 | 0.6282 | No | ||
| 51 | EP300 | 7882 | 0.416 | 0.6121 | No | ||
| 52 | RPS27A | 7967 | 0.401 | 0.6092 | No | ||
| 53 | NPM1 | 8001 | 0.394 | 0.6083 | No | ||
| 54 | ZNF830 | 8127 | 0.376 | 0.6039 | No | ||
| 55 | HUS1B | 8191 | 0.368 | 0.6018 | No | ||
| 56 | CHEK2 | 8686 | 0.299 | 0.5832 | No | ||
| 57 | UBC | 8798 | 0.286 | 0.5792 | No | ||
| 58 | PRKDC | 8863 | 0.276 | 0.5770 | No | ||
| 59 | BLM | 10670 | 0.112 | 0.5082 | No | ||
| 60 | TNKS1BP1 | 11231 | 0.081 | 0.4869 | No | ||
| 61 | GADD45A | 11891 | 0.047 | 0.4619 | No | ||
| 62 | CNOT1 | 12196 | 0.035 | 0.4503 | No | ||
| 63 | CNOT2 | 12436 | 0.025 | 0.4412 | No | ||
| 64 | NEK11 | 12948 | 0.006 | 0.4217 | No | ||
| 65 | MDM4 | 13001 | 0.005 | 0.4198 | No | ||
| 66 | PLAGL1 | 14158 | -0.039 | 0.3757 | No | ||
| 67 | ATF2 | 15697 | -0.113 | 0.3172 | No | ||
| 68 | PPP2R5C | 15792 | -0.120 | 0.3137 | No | ||
| 69 | ATM | 16808 | -0.200 | 0.2751 | No | ||
| 70 | SETMAR | 17845 | -0.316 | 0.2359 | No | ||
| 71 | RAD9B | 18061 | -0.343 | 0.2280 | No | ||
| 72 | CNOT3 | 18104 | -0.349 | 0.2267 | No | ||
| 73 | CNOT4 | 18759 | -0.466 | 0.2022 | No | ||
| 74 | DGKZ | 18924 | -0.501 | 0.1963 | No | ||
| 75 | CNOT10 | 20323 | -0.892 | 0.1438 | No | ||
| 76 | CNOT7 | 20626 | -1.009 | 0.1331 | No | ||
| 77 | FANCI | 20836 | -1.109 | 0.1261 | No | ||
| 78 | TOP2B | 21868 | -1.742 | 0.0883 | No | ||
| 79 | CDC25C | 22485 | -2.318 | 0.0668 | No | ||
| 80 | RPA2 | 22655 | -2.518 | 0.0625 | No | ||
| 81 | CDKN1B | 23273 | -3.364 | 0.0418 | No | ||
| 82 | TFDP1 | 23324 | -3.462 | 0.0429 | No | ||
| 83 | RBL2 | 23437 | -3.679 | 0.0418 | No | ||
| 84 | NBN | 23844 | -4.578 | 0.0302 | No | ||
| 85 | MUC1 | 24081 | -5.263 | 0.0257 | No | ||
| 86 | CNOT6L | 24555 | -7.042 | 0.0137 | No | ||
| 87 | RAD9A | 24579 | -7.142 | 0.0189 | No | ||
| 88 | MSH2 | 24778 | -8.121 | 0.0183 | No | ||
| 89 | TAOK3 | 24789 | -8.188 | 0.0249 | No | ||
| 90 | TIPIN | 25187 | -11.436 | 0.0196 | No | ||
| 91 | TP53 | 25300 | -12.686 | 0.0261 | No | ||
| 92 | PCBP4 | 25454 | -14.763 | 0.0329 | No |