Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | NS50832_padj |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_neg |
| GeneSet | HELLER_HDAC_TARGETS_SILENCED_BY_METHYLATION_DN |
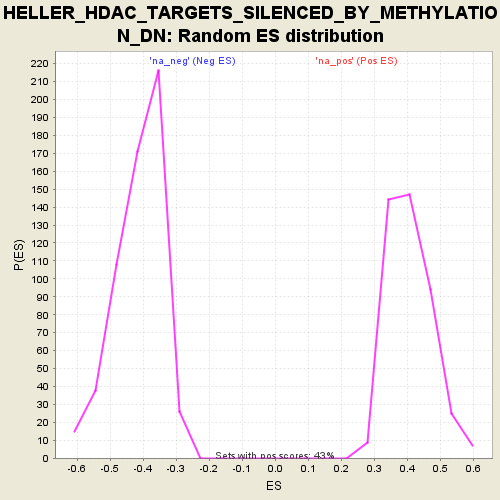
| Enrichment Score (ES) | -0.7274519 |
| Normalized Enrichment Score (NES) | -1.7666788 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.038905274 |
| FWER p-Value | 0.291 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | HSPA1A | 44 | 101.439 | 0.0200 | No | ||
| 2 | HSPA1B | 67 | 84.429 | 0.0372 | No | ||
| 3 | DNAJB1 | 161 | 49.447 | 0.0442 | No | ||
| 4 | GJA1 | 172 | 45.980 | 0.0536 | No | ||
| 5 | FGFR3 | 175 | 44.919 | 0.0631 | No | ||
| 6 | SDC1 | 189 | 42.210 | 0.0716 | No | ||
| 7 | PHACTR2 | 202 | 40.583 | 0.0798 | No | ||
| 8 | LRP8 | 208 | 39.535 | 0.0881 | No | ||
| 9 | GDF15 | 292 | 30.340 | 0.0914 | No | ||
| 10 | TNFRSF10D | 306 | 29.057 | 0.0971 | No | ||
| 11 | BTN3A2 | 653 | 17.260 | 0.0875 | No | ||
| 12 | MZF1 | 707 | 16.196 | 0.0889 | No | ||
| 13 | CADM1 | 816 | 14.612 | 0.0879 | No | ||
| 14 | HSPE1 | 829 | 14.316 | 0.0905 | No | ||
| 15 | PLCL2 | 834 | 14.251 | 0.0934 | No | ||
| 16 | PFAS | 937 | 12.868 | 0.0922 | No | ||
| 17 | FKBP11 | 938 | 12.867 | 0.0950 | No | ||
| 18 | KLHL21 | 955 | 12.622 | 0.0971 | No | ||
| 19 | PDK1 | 1096 | 11.129 | 0.0941 | No | ||
| 20 | SCD | 1102 | 11.098 | 0.0963 | No | ||
| 21 | SEC24D | 1182 | 10.500 | 0.0955 | No | ||
| 22 | TXNDC15 | 1279 | 9.771 | 0.0939 | No | ||
| 23 | HSPH1 | 1350 | 9.216 | 0.0932 | No | ||
| 24 | FAS | 1372 | 9.028 | 0.0943 | No | ||
| 25 | MAGEH1 | 1585 | 7.816 | 0.0878 | No | ||
| 26 | KIAA0368 | 1586 | 7.811 | 0.0895 | No | ||
| 27 | SUMO3 | 1929 | 6.287 | 0.0777 | No | ||
| 28 | TPP1 | 1930 | 6.284 | 0.0790 | No | ||
| 29 | GGT1 | 1956 | 6.204 | 0.0794 | No | ||
| 30 | PDIA6 | 1980 | 6.116 | 0.0798 | No | ||
| 31 | PIM2 | 1989 | 6.092 | 0.0808 | No | ||
| 32 | CDK14 | 2003 | 6.054 | 0.0816 | No | ||
| 33 | RAB15 | 2033 | 5.941 | 0.0818 | No | ||
| 34 | HNRNPUL1 | 2169 | 5.509 | 0.0778 | No | ||
| 35 | PPRC1 | 2209 | 5.371 | 0.0774 | No | ||
| 36 | ALG8 | 2241 | 5.280 | 0.0774 | No | ||
| 37 | MARCH6 | 2381 | 4.897 | 0.0731 | No | ||
| 38 | MAF | 2406 | 4.831 | 0.0732 | No | ||
| 39 | MYB | 2458 | 4.714 | 0.0722 | No | ||
| 40 | KCNN3 | 2468 | 4.690 | 0.0729 | No | ||
| 41 | TYMS | 2510 | 4.612 | 0.0723 | No | ||
| 42 | DNAJB9 | 2733 | 4.122 | 0.0647 | No | ||
| 43 | EHD3 | 2786 | 4.011 | 0.0635 | No | ||
| 44 | SPCS2 | 2975 | 3.654 | 0.0571 | No | ||
| 45 | CELF2 | 3125 | 3.391 | 0.0521 | No | ||
| 46 | HMOX1 | 3141 | 3.368 | 0.0523 | No | ||
| 47 | CDKN2A | 3158 | 3.344 | 0.0524 | No | ||
| 48 | ATF4 | 3319 | 3.105 | 0.0469 | No | ||
| 49 | P2RX5 | 3405 | 2.982 | 0.0443 | No | ||
| 50 | WISP1 | 3499 | 2.869 | 0.0413 | No | ||
| 51 | CCND1 | 3538 | 2.814 | 0.0404 | No | ||
| 52 | ILF3 | 4120 | 2.204 | 0.0186 | No | ||
| 53 | BCL3 | 4227 | 2.078 | 0.0150 | No | ||
| 54 | KCNK1 | 4595 | 1.778 | 0.0013 | No | ||
| 55 | MYBL2 | 4689 | 1.701 | -0.0019 | No | ||
| 56 | GSTO1 | 4694 | 1.696 | -0.0017 | No | ||
| 57 | PMF1 | 4716 | 1.681 | -0.0021 | No | ||
| 58 | ELF1 | 4893 | 1.551 | -0.0086 | No | ||
| 59 | BAG3 | 5070 | 1.426 | -0.0150 | No | ||
| 60 | CDKN2C | 5164 | 1.373 | -0.0183 | No | ||
| 61 | CCDC69 | 5271 | 1.316 | -0.0221 | No | ||
| 62 | SMYD5 | 5332 | 1.273 | -0.0241 | No | ||
| 63 | LRRC41 | 5445 | 1.212 | -0.0281 | No | ||
| 64 | SERPINH1 | 5549 | 1.146 | -0.0318 | No | ||
| 65 | ZMYND8 | 5671 | 1.083 | -0.0362 | No | ||
| 66 | NFIC | 5704 | 1.067 | -0.0372 | No | ||
| 67 | MAP4K1 | 5738 | 1.046 | -0.0383 | No | ||
| 68 | MYCN | 5775 | 1.031 | -0.0394 | No | ||
| 69 | CBS | 5888 | 0.974 | -0.0435 | No | ||
| 70 | PMM2 | 5996 | 0.931 | -0.0474 | No | ||
| 71 | RBM3 | 6525 | 0.754 | -0.0675 | No | ||
| 72 | TXNDC5 | 6581 | 0.732 | -0.0695 | No | ||
| 73 | MYOM2 | 6638 | 0.713 | -0.0715 | No | ||
| 74 | HSPD1 | 6645 | 0.711 | -0.0716 | No | ||
| 75 | FLI1 | 7002 | 0.603 | -0.0851 | No | ||
| 76 | DST | 7044 | 0.590 | -0.0865 | No | ||
| 77 | PKD1 | 7301 | 0.528 | -0.0962 | No | ||
| 78 | AUTS2 | 7319 | 0.524 | -0.0968 | No | ||
| 79 | TNFRSF14 | 7346 | 0.519 | -0.0977 | No | ||
| 80 | SELPLG | 7378 | 0.512 | -0.0987 | No | ||
| 81 | CD58 | 7427 | 0.501 | -0.1005 | No | ||
| 82 | LAPTM5 | 7539 | 0.478 | -0.1046 | No | ||
| 83 | ST6GAL1 | 7574 | 0.470 | -0.1058 | No | ||
| 84 | PXDN | 7673 | 0.452 | -0.1095 | No | ||
| 85 | CBR3 | 7805 | 0.426 | -0.1144 | No | ||
| 86 | SRM | 7877 | 0.416 | -0.1171 | No | ||
| 87 | MRPS15 | 8224 | 0.364 | -0.1303 | No | ||
| 88 | PITX1 | 8450 | 0.329 | -0.1388 | No | ||
| 89 | OAS2 | 8916 | 0.269 | -0.1566 | No | ||
| 90 | LRP2BP | 9006 | 0.260 | -0.1600 | No | ||
| 91 | PSMB10 | 9152 | 0.244 | -0.1655 | No | ||
| 92 | IQGAP2 | 9427 | 0.214 | -0.1759 | No | ||
| 93 | HSPB1 | 9513 | 0.206 | -0.1792 | No | ||
| 94 | SRSF7 | 9632 | 0.194 | -0.1836 | No | ||
| 95 | RASGRP3 | 9942 | 0.167 | -0.1955 | No | ||
| 96 | DNAJB12 | 10155 | 0.148 | -0.2036 | No | ||
| 97 | ZMIZ1 | 10165 | 0.148 | -0.2039 | No | ||
| 98 | LSAMP | 10575 | 0.120 | -0.2195 | No | ||
| 99 | ITGB5 | 10693 | 0.111 | -0.2240 | No | ||
| 100 | FARP1 | 10728 | 0.108 | -0.2253 | No | ||
| 101 | ISG20 | 10851 | 0.100 | -0.2299 | No | ||
| 102 | PHF11 | 11659 | 0.057 | -0.2609 | No | ||
| 103 | ST14 | 11707 | 0.055 | -0.2627 | No | ||
| 104 | GADD45A | 11891 | 0.047 | -0.2697 | No | ||
| 105 | TRIB1 | 12362 | 0.028 | -0.2877 | No | ||
| 106 | CLEC7A | 13570 | -0.015 | -0.3340 | No | ||
| 107 | USF2 | 14149 | -0.038 | -0.3562 | No | ||
| 108 | MAFG | 14243 | -0.042 | -0.3597 | No | ||
| 109 | ANP32B | 15328 | -0.092 | -0.4013 | No | ||
| 110 | PTPRC | 15352 | -0.094 | -0.4022 | No | ||
| 111 | GSDMB | 15507 | -0.102 | -0.4081 | No | ||
| 112 | LONP1 | 15779 | -0.119 | -0.4184 | No | ||
| 113 | ST3GAL6 | 15813 | -0.121 | -0.4197 | No | ||
| 114 | ITGB7 | 15899 | -0.127 | -0.4229 | No | ||
| 115 | IRF4 | 15929 | -0.129 | -0.4240 | No | ||
| 116 | SH2D1A | 16114 | -0.142 | -0.4310 | No | ||
| 117 | ZNF804A | 16160 | -0.144 | -0.4327 | No | ||
| 118 | CLIC2 | 16590 | -0.179 | -0.4491 | No | ||
| 119 | EYA2 | 16789 | -0.198 | -0.4567 | No | ||
| 120 | FERMT1 | 16875 | -0.205 | -0.4599 | No | ||
| 121 | HDGF | 17029 | -0.220 | -0.4657 | No | ||
| 122 | TRAF3IP3 | 17293 | -0.249 | -0.4758 | No | ||
| 123 | RPL10 | 17319 | -0.251 | -0.4767 | No | ||
| 124 | GAS2 | 17364 | -0.258 | -0.4783 | No | ||
| 125 | OAS1 | 17373 | -0.259 | -0.4785 | No | ||
| 126 | TSHR | 17457 | -0.270 | -0.4817 | No | ||
| 127 | DCPS | 17675 | -0.295 | -0.4899 | No | ||
| 128 | PDE3B | 17718 | -0.300 | -0.4915 | No | ||
| 129 | SLA | 17738 | -0.302 | -0.4921 | No | ||
| 130 | AVEN | 17920 | -0.325 | -0.4990 | No | ||
| 131 | LRMP | 18011 | -0.337 | -0.5024 | No | ||
| 132 | IL21R | 18015 | -0.337 | -0.5024 | No | ||
| 133 | FAM149A | 18127 | -0.352 | -0.5066 | No | ||
| 134 | ZEB2 | 18149 | -0.355 | -0.5074 | No | ||
| 135 | CYC1 | 18254 | -0.373 | -0.5113 | No | ||
| 136 | EPCAM | 18272 | -0.376 | -0.5118 | No | ||
| 137 | HOMER3 | 18470 | -0.412 | -0.5193 | No | ||
| 138 | NOL12 | 18484 | -0.413 | -0.5197 | No | ||
| 139 | IGF2BP3 | 18610 | -0.436 | -0.5244 | No | ||
| 140 | COPZ2 | 18841 | -0.481 | -0.5331 | No | ||
| 141 | HERC4 | 18976 | -0.509 | -0.5382 | No | ||
| 142 | IL27RA | 19100 | -0.534 | -0.5428 | No | ||
| 143 | CD79B | 19615 | -0.663 | -0.5623 | No | ||
| 144 | DNAJB4 | 19862 | -0.738 | -0.5716 | No | ||
| 145 | UCK2 | 19951 | -0.767 | -0.5748 | No | ||
| 146 | DEPTOR | 19962 | -0.770 | -0.5751 | No | ||
| 147 | CCPG1 | 19990 | -0.782 | -0.5759 | No | ||
| 148 | SLC25A15 | 20017 | -0.788 | -0.5768 | No | ||
| 149 | CEACAM1 | 20024 | -0.790 | -0.5768 | No | ||
| 150 | ST7 | 20236 | -0.862 | -0.5847 | No | ||
| 151 | XBP1 | 20283 | -0.879 | -0.5863 | No | ||
| 152 | MAFF | 20369 | -0.911 | -0.5894 | No | ||
| 153 | TAPBPL | 20371 | -0.913 | -0.5892 | No | ||
| 154 | IFNAR2 | 20785 | -1.083 | -0.6048 | No | ||
| 155 | TLK1 | 20820 | -1.102 | -0.6059 | No | ||
| 156 | SLC38A1 | 20923 | -1.149 | -0.6096 | No | ||
| 157 | ICK | 21072 | -1.238 | -0.6150 | No | ||
| 158 | BCL2 | 21242 | -1.334 | -0.6212 | No | ||
| 159 | FAIM | 21475 | -1.471 | -0.6298 | No | ||
| 160 | CYTIP | 21723 | -1.636 | -0.6389 | No | ||
| 161 | LIMD2 | 21769 | -1.661 | -0.6403 | No | ||
| 162 | BCL2L1 | 21802 | -1.687 | -0.6411 | No | ||
| 163 | IRF8 | 21907 | -1.778 | -0.6447 | No | ||
| 164 | JMJD1C | 21982 | -1.851 | -0.6472 | No | ||
| 165 | ZNF532 | 22102 | -1.943 | -0.6513 | No | ||
| 166 | HGF | 22197 | -2.029 | -0.6545 | No | ||
| 167 | ITGA4 | 22201 | -2.032 | -0.6542 | No | ||
| 168 | UBE2J1 | 22476 | -2.310 | -0.6642 | No | ||
| 169 | THUMPD2 | 22647 | -2.508 | -0.6702 | No | ||
| 170 | CD48 | 22693 | -2.563 | -0.6714 | No | ||
| 171 | FMNL1 | 22910 | -2.832 | -0.6790 | No | ||
| 172 | FNDC3A | 23128 | -3.121 | -0.6867 | No | ||
| 173 | DOCK10 | 23169 | -3.186 | -0.6876 | No | ||
| 174 | TNS3 | 23204 | -3.248 | -0.6882 | No | ||
| 175 | CDKN1B | 23273 | -3.364 | -0.6901 | No | ||
| 176 | NUAK1 | 23392 | -3.606 | -0.6938 | No | ||
| 177 | PDE4D | 23393 | -3.607 | -0.6930 | No | ||
| 178 | CERS2 | 23432 | -3.665 | -0.6937 | No | ||
| 179 | SETBP1 | 23585 | -3.977 | -0.6987 | No | ||
| 180 | SLC7A5 | 23621 | -4.053 | -0.6992 | No | ||
| 181 | TCF4 | 23715 | -4.268 | -0.7018 | No | ||
| 182 | HSPA9 | 23764 | -4.390 | -0.7027 | No | ||
| 183 | FXYD5 | 23766 | -4.399 | -0.7018 | No | ||
| 184 | STAG2 | 23840 | -4.568 | -0.7037 | No | ||
| 185 | PRTN3 | 23937 | -4.847 | -0.7063 | No | ||
| 186 | SUCLG2 | 24056 | -5.192 | -0.7097 | No | ||
| 187 | IL6R | 24075 | -5.248 | -0.7093 | No | ||
| 188 | POLR3E | 24126 | -5.360 | -0.7101 | No | ||
| 189 | BHLHE40 | 24138 | -5.407 | -0.7093 | No | ||
| 190 | MBNL1 | 24169 | -5.494 | -0.7093 | No | ||
| 191 | CFLAR | 24227 | -5.676 | -0.7103 | No | ||
| 192 | OGT | 24233 | -5.695 | -0.7093 | No | ||
| 193 | STAT6 | 24241 | -5.709 | -0.7083 | No | ||
| 194 | PITPNC1 | 24287 | -5.945 | -0.7088 | No | ||
| 195 | PPP1R15A | 24307 | -6.013 | -0.7082 | No | ||
| 196 | CLK1 | 24334 | -6.145 | -0.7079 | No | ||
| 197 | PPP1R16B | 24396 | -6.370 | -0.7089 | No | ||
| 198 | GALNT14 | 24416 | -6.458 | -0.7082 | No | ||
| 199 | S100A11 | 24497 | -6.746 | -0.7099 | No | ||
| 200 | SLC47A1 | 24538 | -6.937 | -0.7099 | No | ||
| 201 | CHST15 | 24645 | -7.422 | -0.7124 | No | ||
| 202 | ALDH3B1 | 24749 | -7.958 | -0.7146 | No | ||
| 203 | RHOBTB1 | 24796 | -8.216 | -0.7146 | No | ||
| 204 | E2F2 | 24851 | -8.537 | -0.7149 | No | ||
| 205 | EIF5 | 24866 | -8.648 | -0.7136 | No | ||
| 206 | MAPK13 | 24999 | -9.710 | -0.7166 | No | ||
| 207 | SLC3A2 | 25078 | -10.370 | -0.7174 | No | ||
| 208 | IRAK1 | 25115 | -10.721 | -0.7164 | No | ||
| 209 | ZBTB38 | 25316 | -12.862 | -0.7214 | No | ||
| 210 | UBE2E1 | 25456 | -14.805 | -0.7235 | No | ||
| 211 | IFI16 | 25559 | -16.487 | -0.7239 | Yes | ||
| 212 | SRSF8 | 25564 | -16.617 | -0.7205 | Yes | ||
| 213 | SOHLH2 | 25581 | -17.183 | -0.7175 | Yes | ||
| 214 | DUSP6 | 25608 | -17.968 | -0.7146 | Yes | ||
| 215 | NUP210 | 25609 | -17.999 | -0.7108 | Yes | ||
| 216 | WNT5B | 25670 | -19.151 | -0.7090 | Yes | ||
| 217 | ANK3 | 25676 | -19.334 | -0.7051 | Yes | ||
| 218 | NSMAF | 25735 | -21.031 | -0.7028 | Yes | ||
| 219 | MFNG | 25798 | -22.760 | -0.7003 | Yes | ||
| 220 | ZNF280D | 25876 | -26.152 | -0.6977 | Yes | ||
| 221 | WIPF1 | 25925 | -29.160 | -0.6933 | Yes | ||
| 222 | NR3C1 | 25926 | -29.242 | -0.6871 | Yes | ||
| 223 | C6orf48 | 25975 | -31.926 | -0.6821 | Yes | ||
| 224 | FAM129A | 25984 | -32.982 | -0.6753 | Yes | ||
| 225 | NFE2L1 | 26006 | -34.437 | -0.6688 | Yes | ||
| 226 | CCND2 | 26028 | -36.907 | -0.6617 | Yes | ||
| 227 | TIMM44 | 26063 | -40.273 | -0.6544 | Yes | ||
| 228 | SLC7A11 | 26126 | -47.089 | -0.6467 | Yes | ||
| 229 | QPCT | 26127 | -47.117 | -0.6367 | Yes | ||
| 230 | SLC1A4 | 26133 | -48.490 | -0.6265 | Yes | ||
| 231 | HERPUD1 | 26159 | -53.015 | -0.6161 | Yes | ||
| 232 | SHMT2 | 26204 | -64.915 | -0.6040 | Yes | ||
| 233 | CEBPG | 26210 | -66.827 | -0.5899 | Yes | ||
| 234 | CAPN2 | 26220 | -71.517 | -0.5750 | Yes | ||
| 235 | ZHX2 | 26222 | -72.879 | -0.5594 | Yes | ||
| 236 | MYC | 26225 | -75.265 | -0.5434 | Yes | ||
| 237 | PSAT1 | 26230 | -76.008 | -0.5273 | Yes | ||
| 238 | CEBPB | 26244 | -86.643 | -0.5093 | Yes | ||
| 239 | SNTB1 | 26245 | -87.394 | -0.4907 | Yes | ||
| 240 | DNAJC15 | 26246 | -88.122 | -0.4718 | Yes | ||
| 241 | VEGFA | 26251 | -89.987 | -0.4528 | Yes | ||
| 242 | SULF1 | 26261 | -97.588 | -0.4323 | Yes | ||
| 243 | DDIT3 | 26291 | -176.939 | -0.3956 | Yes | ||
| 244 | ATF3 | 26295 | -192.665 | -0.3546 | Yes | ||
| 245 | IFRD1 | 26303 | -230.785 | -0.3055 | Yes | ||
| 246 | TRIB3 | 26304 | -235.323 | -0.2553 | Yes | ||
| 247 | DDIT4 | 26311 | -299.207 | -0.1916 | Yes | ||
| 248 | INHBE | 26312 | -299.207 | -0.1277 | Yes | ||
| 249 | CHAC1 | 26314 | -299.207 | -0.0638 | Yes | ||
| 250 | ATF5 | 26315 | -299.207 | 0.0001 | Yes |