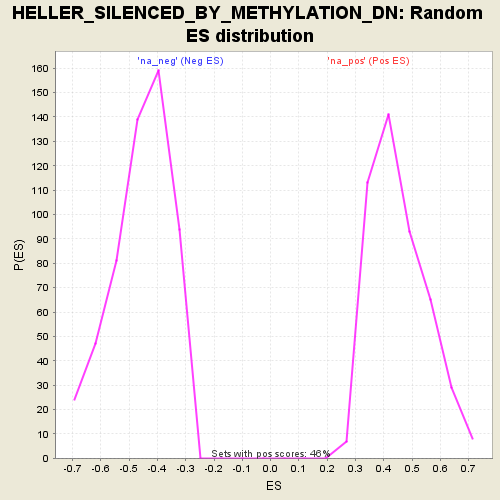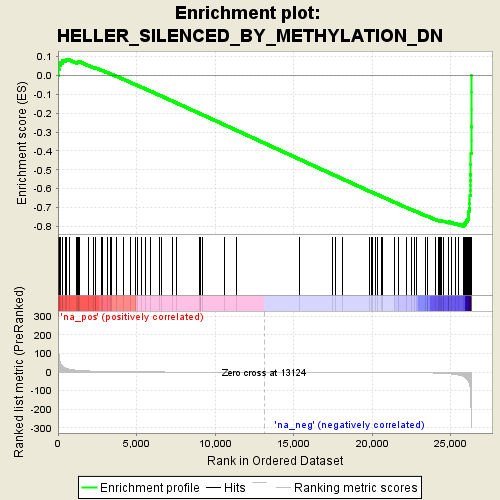
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | NS50832_padj |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_neg |
| GeneSet | HELLER_SILENCED_BY_METHYLATION_DN |
| Enrichment Score (ES) | -0.79885685 |
| Normalized Enrichment Score (NES) | -1.7492001 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.06488052 |
| FWER p-Value | 0.488 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | HSPA1A | 44 | 101.439 | 0.0292 | No | ||
| 2 | HSPA1B | 67 | 84.429 | 0.0541 | No | ||
| 3 | DNAJB1 | 161 | 49.447 | 0.0657 | No | ||
| 4 | CRELD2 | 258 | 34.386 | 0.0725 | No | ||
| 5 | GDF15 | 292 | 30.340 | 0.0805 | No | ||
| 6 | IDH1 | 464 | 21.758 | 0.0806 | No | ||
| 7 | SYT11 | 548 | 19.376 | 0.0833 | No | ||
| 8 | MZF1 | 707 | 16.196 | 0.0822 | No | ||
| 9 | SEC24D | 1182 | 10.500 | 0.0673 | No | ||
| 10 | RRM2 | 1235 | 10.035 | 0.0684 | No | ||
| 11 | CA2 | 1305 | 9.624 | 0.0687 | No | ||
| 12 | UFM1 | 1317 | 9.492 | 0.0712 | No | ||
| 13 | HSPH1 | 1350 | 9.216 | 0.0728 | No | ||
| 14 | TPP1 | 1930 | 6.284 | 0.0526 | No | ||
| 15 | ENPP2 | 2284 | 5.147 | 0.0407 | No | ||
| 16 | NEAT1 | 2398 | 4.852 | 0.0379 | No | ||
| 17 | MAF | 2406 | 4.831 | 0.0391 | No | ||
| 18 | DNAJB9 | 2733 | 4.122 | 0.0279 | No | ||
| 19 | PLTP | 2799 | 3.974 | 0.0267 | No | ||
| 20 | HMOX1 | 3141 | 3.368 | 0.0147 | No | ||
| 21 | ATF4 | 3319 | 3.105 | 0.0089 | No | ||
| 22 | P2RX5 | 3405 | 2.982 | 0.0065 | No | ||
| 23 | TOX3 | 3699 | 2.646 | -0.0038 | No | ||
| 24 | THBS1 | 4168 | 2.135 | -0.0210 | No | ||
| 25 | PGM3 | 4594 | 1.778 | -0.0367 | No | ||
| 26 | SLC5A3 | 4920 | 1.534 | -0.0486 | No | ||
| 27 | BAG3 | 5070 | 1.426 | -0.0539 | No | ||
| 28 | RPS19 | 5342 | 1.269 | -0.0638 | No | ||
| 29 | SERPINH1 | 5549 | 1.146 | -0.0713 | No | ||
| 30 | CBS | 5888 | 0.974 | -0.0839 | No | ||
| 31 | USH2A | 6430 | 0.784 | -0.1043 | No | ||
| 32 | TXNDC5 | 6581 | 0.732 | -0.1098 | No | ||
| 33 | PKD1 | 7301 | 0.528 | -0.1371 | No | ||
| 34 | LAPTM5 | 7539 | 0.478 | -0.1459 | No | ||
| 35 | LRP2BP | 9006 | 0.260 | -0.2018 | No | ||
| 36 | CATSPER2 | 9043 | 0.255 | -0.2031 | No | ||
| 37 | ADRB2 | 9079 | 0.251 | -0.2043 | No | ||
| 38 | EPB41L2 | 9187 | 0.241 | -0.2083 | No | ||
| 39 | LSAMP | 10575 | 0.120 | -0.2612 | No | ||
| 40 | CRBN | 11367 | 0.072 | -0.2913 | No | ||
| 41 | PTPRC | 15352 | -0.094 | -0.4432 | No | ||
| 42 | TSHR | 17457 | -0.270 | -0.5234 | No | ||
| 43 | OSGIN1 | 17673 | -0.294 | -0.5315 | No | ||
| 44 | FAM149A | 18127 | -0.352 | -0.5487 | No | ||
| 45 | DNAJB4 | 19862 | -0.738 | -0.6146 | No | ||
| 46 | CCPG1 | 19990 | -0.782 | -0.6192 | No | ||
| 47 | CEACAM1 | 20024 | -0.790 | -0.6202 | No | ||
| 48 | RFPL2 | 20203 | -0.849 | -0.6267 | No | ||
| 49 | MAFF | 20369 | -0.911 | -0.6327 | No | ||
| 50 | F2RL1 | 20607 | -1.003 | -0.6415 | No | ||
| 51 | MAP1B | 20641 | -1.018 | -0.6424 | No | ||
| 52 | PPFIBP1 | 21445 | -1.449 | -0.6726 | No | ||
| 53 | HSPA13 | 21655 | -1.583 | -0.6801 | No | ||
| 54 | HGF | 22197 | -2.029 | -0.7001 | No | ||
| 55 | PPARGC1A | 22501 | -2.331 | -0.7110 | No | ||
| 56 | STK39 | 22707 | -2.573 | -0.7180 | No | ||
| 57 | RCAN2 | 22804 | -2.692 | -0.7208 | No | ||
| 58 | AAK1 | 23400 | -3.617 | -0.7424 | No | ||
| 59 | MMP16 | 23530 | -3.846 | -0.7462 | No | ||
| 60 | SLC6A15 | 23553 | -3.909 | -0.7458 | No | ||
| 61 | GOT1 | 24030 | -5.126 | -0.7624 | No | ||
| 62 | PHGDH | 24203 | -5.598 | -0.7673 | No | ||
| 63 | PPP1R15A | 24307 | -6.013 | -0.7694 | No | ||
| 64 | CLK1 | 24334 | -6.145 | -0.7685 | No | ||
| 65 | PLEKHA8P1 | 24397 | -6.381 | -0.7689 | No | ||
| 66 | SLC47A1 | 24538 | -6.937 | -0.7721 | No | ||
| 67 | EIF4EBP1 | 24571 | -7.108 | -0.7712 | No | ||
| 68 | SLCO4C1 | 24842 | -8.472 | -0.7789 | No | ||
| 69 | E2F2 | 24851 | -8.537 | -0.7766 | No | ||
| 70 | MARS | 24894 | -8.832 | -0.7755 | No | ||
| 71 | SLC3A2 | 25078 | -10.370 | -0.7793 | No | ||
| 72 | DDAH1 | 25308 | -12.795 | -0.7841 | No | ||
| 73 | CTAGE5 | 25523 | -15.807 | -0.7875 | No | ||
| 74 | CDS1 | 25822 | -23.725 | -0.7916 | Yes | ||
| 75 | RPS6KA2 | 25884 | -26.388 | -0.7859 | Yes | ||
| 76 | CLGN | 25962 | -31.385 | -0.7793 | Yes | ||
| 77 | FAM129A | 25984 | -32.982 | -0.7700 | Yes | ||
| 78 | TIMM44 | 26063 | -40.273 | -0.7607 | Yes | ||
| 79 | WARS | 26117 | -46.483 | -0.7486 | Yes | ||
| 80 | SLC7A11 | 26126 | -47.089 | -0.7345 | Yes | ||
| 81 | SLC1A4 | 26133 | -48.490 | -0.7200 | Yes | ||
| 82 | SHMT2 | 26204 | -64.915 | -0.7029 | Yes | ||
| 83 | MYC | 26225 | -75.265 | -0.6807 | Yes | ||
| 84 | PSAT1 | 26230 | -76.008 | -0.6577 | Yes | ||
| 85 | RGS16 | 26231 | -76.235 | -0.6345 | Yes | ||
| 86 | ASNS | 26241 | -83.978 | -0.6092 | Yes | ||
| 87 | CEBPB | 26244 | -86.643 | -0.5829 | Yes | ||
| 88 | DNAJC15 | 26246 | -88.122 | -0.5560 | Yes | ||
| 89 | CTH | 26265 | -99.001 | -0.5266 | Yes | ||
| 90 | DDIT3 | 26291 | -176.939 | -0.4736 | Yes | ||
| 91 | ATF3 | 26295 | -192.665 | -0.4150 | Yes | ||
| 92 | IFRD1 | 26303 | -230.785 | -0.3449 | Yes | ||
| 93 | TRIB3 | 26304 | -235.323 | -0.2732 | Yes | ||
| 94 | DDIT4 | 26311 | -299.207 | -0.1822 | Yes | ||
| 95 | INHBE | 26312 | -299.207 | -0.0910 | Yes | ||
| 96 | CHAC1 | 26314 | -299.207 | 0.0001 | Yes |