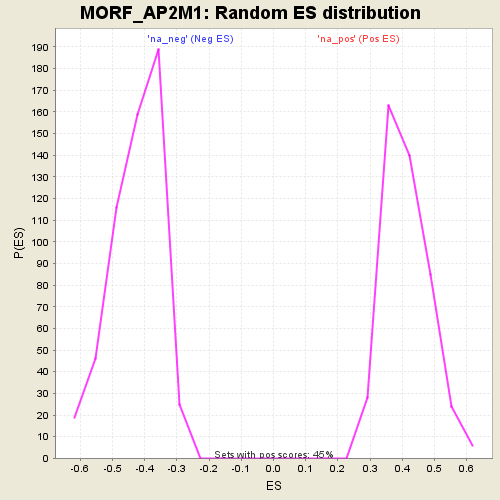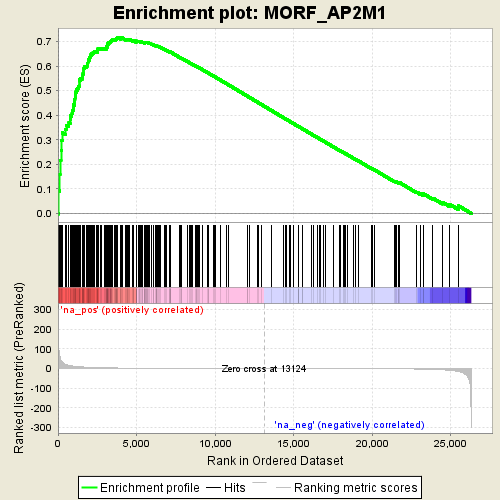
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | NS50832_padj |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_pos |
| GeneSet | MORF_AP2M1 |
| Enrichment Score (ES) | 0.71616745 |
| Normalized Enrichment Score (NES) | 1.7442424 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.16723655 |
| FWER p-Value | 0.71 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | HSPA8 | 56 | 93.481 | 0.0928 | Yes | ||
| 2 | TUBB2A | 105 | 65.807 | 0.1577 | Yes | ||
| 3 | ARHGDIA | 126 | 58.390 | 0.2162 | Yes | ||
| 4 | VCP | 188 | 42.258 | 0.2568 | Yes | ||
| 5 | EIF4E2 | 201 | 40.595 | 0.2975 | Yes | ||
| 6 | CALR | 266 | 32.957 | 0.3285 | Yes | ||
| 7 | PGD | 477 | 21.362 | 0.3422 | Yes | ||
| 8 | TMED9 | 539 | 19.512 | 0.3597 | Yes | ||
| 9 | CAPN1 | 690 | 16.522 | 0.3707 | Yes | ||
| 10 | SSR2 | 766 | 15.156 | 0.3832 | Yes | ||
| 11 | POLR2L | 794 | 14.910 | 0.3973 | Yes | ||
| 12 | LYPLA2 | 884 | 13.611 | 0.4077 | Yes | ||
| 13 | MRPS12 | 910 | 13.251 | 0.4202 | Yes | ||
| 14 | TUBA1B | 982 | 12.363 | 0.4300 | Yes | ||
| 15 | PPIA | 997 | 12.201 | 0.4419 | Yes | ||
| 16 | NDUFA2 | 1051 | 11.590 | 0.4516 | Yes | ||
| 17 | RPL36AL | 1064 | 11.483 | 0.4628 | Yes | ||
| 18 | PPP2R1A | 1078 | 11.290 | 0.4738 | Yes | ||
| 19 | RAB1A | 1098 | 11.114 | 0.4843 | Yes | ||
| 20 | GNAS | 1126 | 10.933 | 0.4944 | Yes | ||
| 21 | CALM1 | 1171 | 10.568 | 0.5034 | Yes | ||
| 22 | SUMO2 | 1250 | 9.927 | 0.5105 | Yes | ||
| 23 | TRAPPC3 | 1309 | 9.579 | 0.5180 | Yes | ||
| 24 | FKBP1A | 1342 | 9.289 | 0.5262 | Yes | ||
| 25 | HSP90AA1 | 1345 | 9.282 | 0.5356 | Yes | ||
| 26 | UBE2L3 | 1351 | 9.211 | 0.5447 | Yes | ||
| 27 | DYNC1I2 | 1442 | 8.644 | 0.5501 | Yes | ||
| 28 | CBX3 | 1527 | 8.187 | 0.5552 | Yes | ||
| 29 | MORF4L2 | 1544 | 8.093 | 0.5628 | Yes | ||
| 30 | PUF60 | 1584 | 7.822 | 0.5692 | Yes | ||
| 31 | SEC61B | 1634 | 7.567 | 0.5750 | Yes | ||
| 32 | DAZAP2 | 1638 | 7.534 | 0.5826 | Yes | ||
| 33 | PRMT1 | 1642 | 7.527 | 0.5901 | Yes | ||
| 34 | COX6B1 | 1668 | 7.436 | 0.5967 | Yes | ||
| 35 | PSMA4 | 1791 | 6.873 | 0.5990 | Yes | ||
| 36 | WBP2 | 1846 | 6.654 | 0.6037 | Yes | ||
| 37 | ENSA | 1868 | 6.531 | 0.6095 | Yes | ||
| 38 | HSP90AB1 | 1905 | 6.375 | 0.6146 | Yes | ||
| 39 | SNRPE | 1908 | 6.371 | 0.6210 | Yes | ||
| 40 | SUMO3 | 1929 | 6.287 | 0.6266 | Yes | ||
| 41 | HNRNPAB | 2011 | 6.002 | 0.6296 | Yes | ||
| 42 | ARF3 | 2015 | 5.993 | 0.6355 | Yes | ||
| 43 | NDUFA1 | 2039 | 5.918 | 0.6407 | Yes | ||
| 44 | POLR2H | 2080 | 5.778 | 0.6450 | Yes | ||
| 45 | COMMD4 | 2120 | 5.661 | 0.6493 | Yes | ||
| 46 | HNRNPUL1 | 2169 | 5.509 | 0.6530 | Yes | ||
| 47 | GABARAP | 2255 | 5.236 | 0.6551 | Yes | ||
| 48 | MAP3K11 | 2303 | 5.094 | 0.6584 | Yes | ||
| 49 | DNAJC8 | 2433 | 4.759 | 0.6583 | Yes | ||
| 50 | BANF1 | 2505 | 4.630 | 0.6603 | Yes | ||
| 51 | MRPL9 | 2515 | 4.602 | 0.6646 | Yes | ||
| 52 | TPI1 | 2516 | 4.596 | 0.6693 | Yes | ||
| 53 | TCF25 | 2585 | 4.455 | 0.6712 | Yes | ||
| 54 | ARPC5 | 2685 | 4.224 | 0.6717 | Yes | ||
| 55 | VAMP3 | 2793 | 3.991 | 0.6717 | Yes | ||
| 56 | RPL6 | 2962 | 3.681 | 0.6690 | Yes | ||
| 57 | TMED2 | 3029 | 3.568 | 0.6701 | Yes | ||
| 58 | P4HB | 3059 | 3.515 | 0.6725 | Yes | ||
| 59 | ATP6V1F | 3067 | 3.507 | 0.6758 | Yes | ||
| 60 | UBA1 | 3077 | 3.481 | 0.6790 | Yes | ||
| 61 | RXRA | 3128 | 3.386 | 0.6805 | Yes | ||
| 62 | GPI | 3139 | 3.369 | 0.6836 | Yes | ||
| 63 | C9orf16 | 3149 | 3.353 | 0.6866 | Yes | ||
| 64 | PCMT1 | 3154 | 3.350 | 0.6899 | Yes | ||
| 65 | AP2M1 | 3189 | 3.303 | 0.6919 | Yes | ||
| 66 | SLC25A3 | 3213 | 3.255 | 0.6943 | Yes | ||
| 67 | AP2S1 | 3247 | 3.210 | 0.6963 | Yes | ||
| 68 | EEF1D | 3308 | 3.128 | 0.6972 | Yes | ||
| 69 | LDHA | 3339 | 3.076 | 0.6992 | Yes | ||
| 70 | ATP5J2 | 3356 | 3.050 | 0.7017 | Yes | ||
| 71 | SKP1 | 3377 | 3.018 | 0.7040 | Yes | ||
| 72 | CFL1 | 3416 | 2.966 | 0.7055 | Yes | ||
| 73 | RAN | 3473 | 2.894 | 0.7063 | Yes | ||
| 74 | TMEM147 | 3582 | 2.764 | 0.7050 | Yes | ||
| 75 | NME2 | 3661 | 2.694 | 0.7047 | Yes | ||
| 76 | RAB5B | 3672 | 2.683 | 0.7071 | Yes | ||
| 77 | OTUB1 | 3704 | 2.634 | 0.7086 | Yes | ||
| 78 | PSMB4 | 3707 | 2.631 | 0.7112 | Yes | ||
| 79 | CALM2 | 3713 | 2.622 | 0.7136 | Yes | ||
| 80 | SUMO1 | 3763 | 2.565 | 0.7143 | Yes | ||
| 81 | CLIC1 | 3784 | 2.545 | 0.7162 | Yes | ||
| 82 | PSMB7 | 3958 | 2.358 | 0.7119 | No | ||
| 83 | CAPZA1 | 3980 | 2.335 | 0.7135 | No | ||
| 84 | SLC4A2 | 4007 | 2.309 | 0.7148 | No | ||
| 85 | ATP5I | 4079 | 2.238 | 0.7144 | No | ||
| 86 | COX7C | 4312 | 2.006 | 0.7076 | No | ||
| 87 | SNRPB | 4336 | 1.985 | 0.7087 | No | ||
| 88 | HMGN1 | 4411 | 1.921 | 0.7078 | No | ||
| 89 | SRSF9 | 4444 | 1.892 | 0.7085 | No | ||
| 90 | YWHAB | 4514 | 1.842 | 0.7077 | No | ||
| 91 | NACA | 4559 | 1.804 | 0.7079 | No | ||
| 92 | HSBP1 | 4750 | 1.652 | 0.7023 | No | ||
| 93 | PPP1CA | 4801 | 1.621 | 0.7020 | No | ||
| 94 | SSNA1 | 4833 | 1.598 | 0.7024 | No | ||
| 95 | SF3B2 | 4967 | 1.495 | 0.6989 | No | ||
| 96 | BCAP31 | 4974 | 1.492 | 0.7001 | No | ||
| 97 | TADA3 | 4996 | 1.477 | 0.7008 | No | ||
| 98 | YWHAQ | 5008 | 1.472 | 0.7019 | No | ||
| 99 | TBCB | 5095 | 1.411 | 0.7000 | No | ||
| 100 | PGAM1 | 5155 | 1.378 | 0.6992 | No | ||
| 101 | COX6A1 | 5180 | 1.364 | 0.6996 | No | ||
| 102 | C14orf2 | 5242 | 1.329 | 0.6987 | No | ||
| 103 | PSMB6 | 5289 | 1.303 | 0.6982 | No | ||
| 104 | COX5B | 5352 | 1.262 | 0.6971 | No | ||
| 105 | ARF5 | 5406 | 1.238 | 0.6964 | No | ||
| 106 | GNAI2 | 5502 | 1.177 | 0.6939 | No | ||
| 107 | RALY | 5504 | 1.175 | 0.6951 | No | ||
| 108 | COPE | 5520 | 1.165 | 0.6957 | No | ||
| 109 | PSMD8 | 5541 | 1.152 | 0.6961 | No | ||
| 110 | PSMC1 | 5559 | 1.143 | 0.6966 | No | ||
| 111 | IMPDH1 | 5601 | 1.119 | 0.6962 | No | ||
| 112 | FAU | 5616 | 1.115 | 0.6967 | No | ||
| 113 | PSMB1 | 5677 | 1.079 | 0.6955 | No | ||
| 114 | TMBIM6 | 5680 | 1.077 | 0.6966 | No | ||
| 115 | TUG1 | 5764 | 1.037 | 0.6944 | No | ||
| 116 | COX4I1 | 5820 | 1.005 | 0.6933 | No | ||
| 117 | MRPL23 | 5971 | 0.941 | 0.6886 | No | ||
| 118 | GDI2 | 6048 | 0.913 | 0.6866 | No | ||
| 119 | GANAB | 6086 | 0.901 | 0.6861 | No | ||
| 120 | PPP1R7 | 6202 | 0.856 | 0.6825 | No | ||
| 121 | NEDD8 | 6263 | 0.836 | 0.6811 | No | ||
| 122 | DNPEP | 6277 | 0.830 | 0.6814 | No | ||
| 123 | YWHAZ | 6285 | 0.828 | 0.6820 | No | ||
| 124 | ATP6V0C | 6359 | 0.805 | 0.6800 | No | ||
| 125 | DRG1 | 6387 | 0.795 | 0.6798 | No | ||
| 126 | CCT7 | 6486 | 0.767 | 0.6768 | No | ||
| 127 | CTDNEP1 | 6536 | 0.751 | 0.6757 | No | ||
| 128 | ACTN4 | 6787 | 0.661 | 0.6668 | No | ||
| 129 | SRP14 | 6816 | 0.654 | 0.6664 | No | ||
| 130 | PIEZO1 | 6907 | 0.630 | 0.6636 | No | ||
| 131 | XRCC6 | 7077 | 0.582 | 0.6577 | No | ||
| 132 | STARD7 | 7115 | 0.571 | 0.6569 | No | ||
| 133 | SDHA | 7134 | 0.567 | 0.6567 | No | ||
| 134 | POLR2I | 7137 | 0.567 | 0.6572 | No | ||
| 135 | PSMD7 | 7143 | 0.565 | 0.6576 | No | ||
| 136 | AP1B1 | 7735 | 0.439 | 0.6354 | No | ||
| 137 | CLTA | 7765 | 0.433 | 0.6348 | No | ||
| 138 | COX7A2L | 7840 | 0.421 | 0.6324 | No | ||
| 139 | RHEB | 7866 | 0.418 | 0.6318 | No | ||
| 140 | HUWE1 | 8212 | 0.366 | 0.6190 | No | ||
| 141 | CANX | 8348 | 0.345 | 0.6142 | No | ||
| 142 | EIF4A1 | 8417 | 0.335 | 0.6119 | No | ||
| 143 | SOD1 | 8489 | 0.323 | 0.6095 | No | ||
| 144 | LASP1 | 8536 | 0.318 | 0.6081 | No | ||
| 145 | GPX4 | 8740 | 0.292 | 0.6006 | No | ||
| 146 | UBC | 8798 | 0.286 | 0.5987 | No | ||
| 147 | COX8A | 8840 | 0.280 | 0.5974 | No | ||
| 148 | ATOX1 | 8872 | 0.275 | 0.5965 | No | ||
| 149 | EIF3C | 8949 | 0.266 | 0.5939 | No | ||
| 150 | ARPC2 | 9033 | 0.256 | 0.5909 | No | ||
| 151 | UBE2I | 9215 | 0.237 | 0.5842 | No | ||
| 152 | COPS5 | 9514 | 0.206 | 0.5730 | No | ||
| 153 | GMFB | 9562 | 0.202 | 0.5714 | No | ||
| 154 | HNRNPC | 9576 | 0.199 | 0.5711 | No | ||
| 155 | PGK1 | 9909 | 0.170 | 0.5586 | No | ||
| 156 | EIF4G2 | 9946 | 0.167 | 0.5574 | No | ||
| 157 | VPS52 | 10028 | 0.160 | 0.5545 | No | ||
| 158 | DCTN2 | 10337 | 0.135 | 0.5428 | No | ||
| 159 | IDH3B | 10721 | 0.109 | 0.5282 | No | ||
| 160 | CSNK2B | 10883 | 0.098 | 0.5222 | No | ||
| 161 | AP3D1 | 12055 | 0.042 | 0.4774 | No | ||
| 162 | RER1 | 12078 | 0.041 | 0.4765 | No | ||
| 163 | COPS6 | 12203 | 0.035 | 0.4718 | No | ||
| 164 | UQCRFS1 | 12726 | 0.014 | 0.4519 | No | ||
| 165 | SMARCD2 | 12757 | 0.012 | 0.4507 | No | ||
| 166 | MAP2K2 | 12922 | 0.007 | 0.4444 | No | ||
| 167 | C21orf33 | 13611 | -0.016 | 0.4181 | No | ||
| 168 | GPAA1 | 14371 | -0.047 | 0.3891 | No | ||
| 169 | SNX3 | 14460 | -0.050 | 0.3858 | No | ||
| 170 | SEPT2 | 14558 | -0.053 | 0.3821 | No | ||
| 171 | IST1 | 14724 | -0.058 | 0.3758 | No | ||
| 172 | URM1 | 14825 | -0.063 | 0.3721 | No | ||
| 173 | GNB2 | 15018 | -0.074 | 0.3648 | No | ||
| 174 | UQCRB | 15308 | -0.090 | 0.3538 | No | ||
| 175 | MYL12B | 15325 | -0.092 | 0.3533 | No | ||
| 176 | ANP32B | 15328 | -0.092 | 0.3533 | No | ||
| 177 | DDOST | 15536 | -0.104 | 0.3455 | No | ||
| 178 | RAC1 | 15577 | -0.107 | 0.3441 | No | ||
| 179 | ATP6AP1 | 16142 | -0.143 | 0.3226 | No | ||
| 180 | C1D | 16258 | -0.151 | 0.3183 | No | ||
| 181 | KARS | 16544 | -0.173 | 0.3076 | No | ||
| 182 | DDX49 | 16657 | -0.186 | 0.3035 | No | ||
| 183 | TIAL1 | 16685 | -0.188 | 0.3027 | No | ||
| 184 | STX4 | 16906 | -0.208 | 0.2944 | No | ||
| 185 | HDGF | 17029 | -0.220 | 0.2900 | No | ||
| 186 | PIGC | 17529 | -0.278 | 0.2712 | No | ||
| 187 | KDELR1 | 17905 | -0.323 | 0.2571 | No | ||
| 188 | PDAP1 | 17991 | -0.335 | 0.2542 | No | ||
| 189 | MYL12A | 18203 | -0.363 | 0.2465 | No | ||
| 190 | CYC1 | 18254 | -0.373 | 0.2450 | No | ||
| 191 | BAG5 | 18334 | -0.386 | 0.2423 | No | ||
| 192 | PLOD3 | 18439 | -0.407 | 0.2388 | No | ||
| 193 | CAPZB | 18801 | -0.472 | 0.2254 | No | ||
| 194 | MBD4 | 18947 | -0.505 | 0.2204 | No | ||
| 195 | HADHB | 19125 | -0.541 | 0.2141 | No | ||
| 196 | SEPHS2 | 19936 | -0.761 | 0.1839 | No | ||
| 197 | STOML2 | 20008 | -0.786 | 0.1820 | No | ||
| 198 | RPN2 | 20141 | -0.827 | 0.1777 | No | ||
| 199 | TMEM59 | 21397 | -1.420 | 0.1311 | No | ||
| 200 | ARFGAP2 | 21459 | -1.458 | 0.1303 | No | ||
| 201 | PCBP2 | 21541 | -1.510 | 0.1287 | No | ||
| 202 | DAP3 | 21675 | -1.597 | 0.1252 | No | ||
| 203 | COG4 | 21678 | -1.600 | 0.1268 | No | ||
| 204 | JTB | 21732 | -1.645 | 0.1264 | No | ||
| 205 | SETD3 | 22838 | -2.755 | 0.0869 | No | ||
| 206 | MDH1 | 23098 | -3.088 | 0.0801 | No | ||
| 207 | SRP9 | 23252 | -3.333 | 0.0776 | No | ||
| 208 | BUD31 | 23285 | -3.391 | 0.0798 | No | ||
| 209 | CTBP1 | 23872 | -4.662 | 0.0621 | No | ||
| 210 | PRKAR1A | 24489 | -6.684 | 0.0453 | No | ||
| 211 | SAP18 | 24907 | -8.930 | 0.0384 | No | ||
| 212 | CNBP | 25497 | -15.354 | 0.0314 | No |